Differential expression
Last updated: 2025-06-09
Checks: 7 0
Knit directory: prs/
This reproducible R Markdown analysis was created with workflowr (version 1.7.1). The Checks tab describes the reproducibility checks that were applied when the results were created. The Past versions tab lists the development history.
Great! Since the R Markdown file has been committed to the Git repository, you know the exact version of the code that produced these results.
Great job! The global environment was empty. Objects defined in the global environment can affect the analysis in your R Markdown file in unknown ways. For reproduciblity it’s best to always run the code in an empty environment.
The command set.seed(20250417) was run prior to running
the code in the R Markdown file. Setting a seed ensures that any results
that rely on randomness, e.g. subsampling or permutations, are
reproducible.
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
Nice! There were no cached chunks for this analysis, so you can be confident that you successfully produced the results during this run.
Great job! Using relative paths to the files within your workflowr project makes it easier to run your code on other machines.
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility.
The results in this page were generated with repository version 66b1a17. See the Past versions tab to see a history of the changes made to the R Markdown and HTML files.
Note that you need to be careful to ensure that all relevant files for
the analysis have been committed to Git prior to generating the results
(you can use wflow_publish or
wflow_git_commit). workflowr only checks the R Markdown
file, but you know if there are other scripts or data files that it
depends on. Below is the status of the Git repository when the results
were generated:
Ignored files:
Ignored: .DS_Store
Ignored: .Rhistory
Ignored: .Rproj.user/
Ignored: analysis/.DS_Store
Ignored: analysis/.Rhistory
Ignored: data/.DS_Store
Untracked files:
Untracked: analysis/asthma_lung/
Untracked: analysis/asthma_wb/
Untracked: analysis/continuous_artery/
Untracked: analysis/continuous_wb/
Untracked: analysis/continuous_wb_m2/
Untracked: analysis/metadata.txt
Untracked: analysis/metadata_artery.txt
Untracked: analysis/metadata_artery_quantile.txt
Untracked: analysis/metadata_lung_asthma.txt
Untracked: analysis/metadata_lung_asthma_quantile.txt
Untracked: analysis/metadata_quantile.txt
Untracked: analysis/metadata_wb_asthma.txt
Untracked: analysis/metadata_wb_asthma_quantile.txt
Untracked: analysis/normalized_counts.rda
Untracked: analysis/prs_asthma.txt
Untracked: analysis/quantile_artery/
Untracked: analysis/quantile_wb/
Untracked: analysis/quantile_wb_m2/
Untracked: analysis/vst norm counts.rda
Untracked: data/Artery_Aorta.v8.covariates.txt
Untracked: data/GTEx_v8.bk
Untracked: data/GTEx_v8.rds
Untracked: data/Lung.v8.covariates.txt
Untracked: data/PGS001787.txt
Untracked: data/PGS001787_hmPOS_GRCh38.txt
Untracked: data/T2D_hmPOS_GRCh38.txt
Untracked: data/Whole_Blood.v8.covariates.txt
Untracked: data/blood_cell/
Untracked: data/endotype_sumstats/
Untracked: data/gene_reads_2017-06-05_v8_artery_aorta.gct
Untracked: data/gene_reads_2017-06-05_v8_lung.gct
Untracked: data/gene_reads_2017-06-05_v8_whole_blood.gct
Untracked: data/immune/
Untracked: data/pca.eigenval
Untracked: data/pca.eigenvec
Untracked: data/protein-coding_gene.txt
Unstaged changes:
Modified: analysis/PRS.Rmd
Deleted: analysis/QC.Rmd
Modified: analysis/asthma_wb.Rmd
Deleted: analysis/normalized_counts.txt
Modified: analysis/prs_blood_cell.txt
Modified: prs.Rproj
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.
These are the previous versions of the repository in which changes were
made to the R Markdown
(analysis/differential_expression.Rmd) and HTML
(docs/differential_expression.html) files. If you’ve
configured a remote Git repository (see ?wflow_git_remote),
click on the hyperlinks in the table below to view the files as they
were in that past version.
| File | Version | Author | Date | Message |
|---|---|---|---|---|
| Rmd | 66b1a17 | ElisaChen | 2025-06-09 | workflowr::wflow_publish(c("analysis/differential_expression.Rmd", |
| html | c49ada7 | ElisaChen | 2025-05-20 | Build site. |
| html | c2d7279 | ElisaChen | 2025-05-20 | Build site. |
| Rmd | e3d43a7 | ElisaChen | 2025-05-20 | workflowr::wflow_publish("analysis/differential_expression.Rmd") |
| html | cf16954 | ElisaChen | 2025-05-20 | Build site. |
| Rmd | 1200397 | ElisaChen | 2025-05-20 | workflowr::wflow_publish("analysis/differential_expression.Rmd") |
| html | 51ffd48 | ElisaChen | 2025-05-20 | Build site. |
| Rmd | 3a1e4dd | ElisaChen | 2025-05-20 | workflowr::wflow_publish("analysis/differential_expression.Rmd") |
| html | e4dd48b | ElisaChen | 2025-05-16 | Build site. |
| Rmd | 4bdd6d2 | ElisaChen | 2025-05-16 | workflowr::wflow_publish("analysis/differential_expression.Rmd") |
| html | d78bc7e | ElisaChen | 2025-05-16 | Build site. |
| Rmd | f9c370a | ElisaChen | 2025-05-16 | workflowr::wflow_publish("analysis/differential_expression.Rmd") |
| html | 1ef6ae9 | ElisaChen | 2025-05-14 | Build site. |
| Rmd | 0591bd8 | ElisaChen | 2025-05-14 | workflowr::wflow_publish(c("analysis/differential_expression.Rmd", |
| html | 3bb86a4 | ElisaChen | 2025-05-13 | Build site. |
| Rmd | a264964 | ElisaChen | 2025-05-13 | workflowr::wflow_publish("analysis/differential_expression.Rmd") |
| html | 2b73cc3 | ElisaChen | 2025-05-13 | Build site. |
| Rmd | c5d5446 | ElisaChen | 2025-05-13 | workflowr::wflow_publish(c("analysis/PRS.Rmd", "analysis/index.Rmd", |
| html | daf5170 | ElisaChen | 2025-05-07 | Build site. |
| Rmd | 060c7ac | ElisaChen | 2025-05-07 | wflow_publish("analysis/differential_expression.Rmd") |
| html | d129b3e | ElisaChen | 2025-05-07 | Build site. |
| Rmd | 7a5b0e4 | ElisaChen | 2025-05-07 | wflow_publish(c("analysis/index.Rmd", "analysis/metadata.Rmd", |
| html | 5e7a42e | ElisaChen | 2025-04-30 | Build site. |
| Rmd | 6a1157a | ElisaChen | 2025-04-30 | workflowr::wflow_publish("analysis/differential_expression.Rmd") |
| html | 959e031 | ElisaChen | 2025-04-30 | Build site. |
| Rmd | 9017040 | ElisaChen | 2025-04-30 | workflowr::wflow_publish("analysis/differential_expression.Rmd") |
| html | ae6b2a5 | ElisaChen | 2025-04-30 | Build site. |
| Rmd | 6488595 | ElisaChen | 2025-04-30 | workflowr::wflow_publish("analysis/differential_expression.Rmd") |
| html | c4ece77 | ElisaChen | 2025-04-30 | Build site. |
| Rmd | 5b80e27 | ElisaChen | 2025-04-30 | workflowr::wflow_publish(c("analysis/differential_expression.Rmd", |
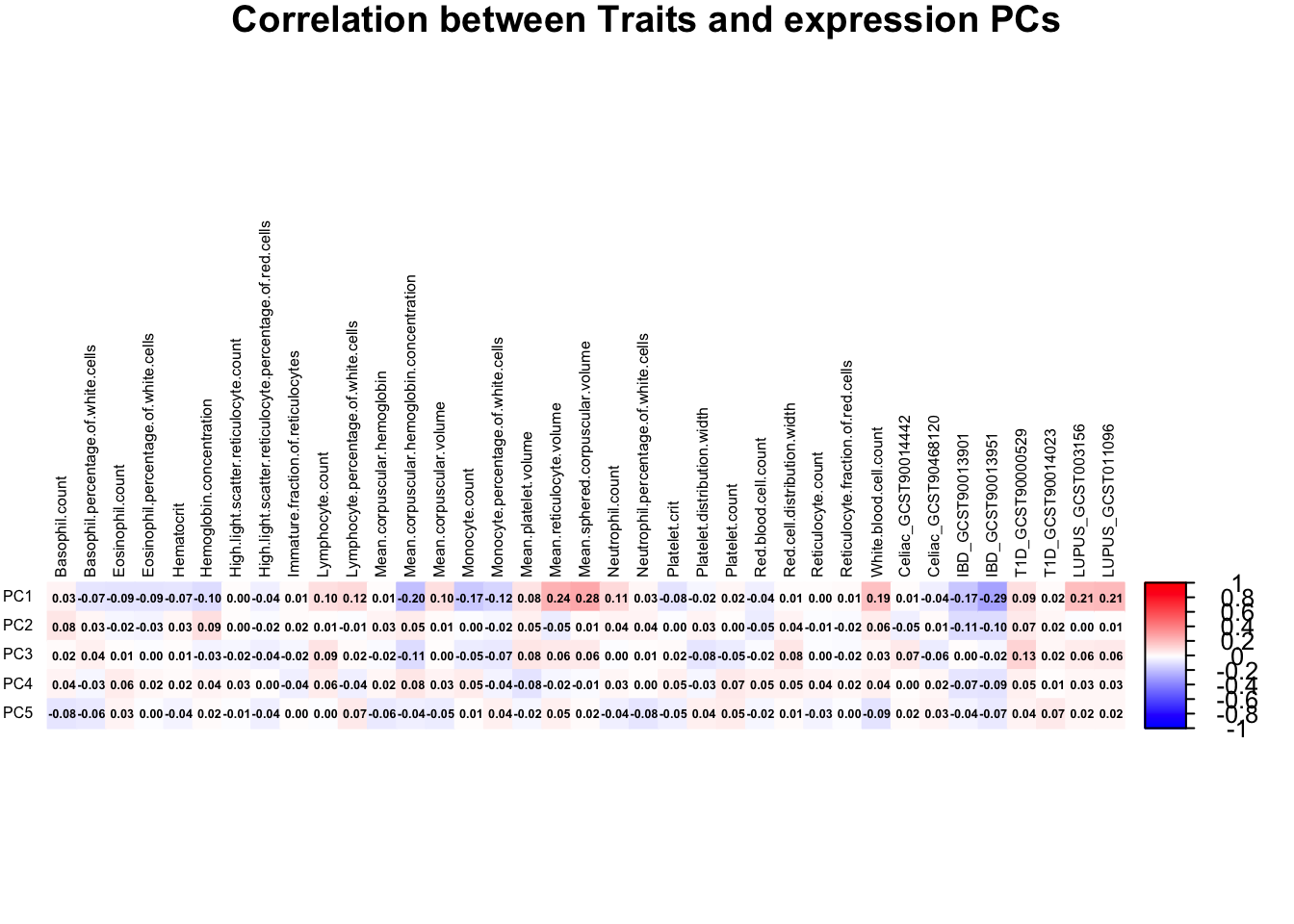
Correlation between PRS & expression PCs (whole blood)
# load prs & pcs
metadata_file <- "analysis/metadata.txt"
metadata <- read.csv(metadata_file, header = T, sep = "\t", stringsAsFactors = T)
metadata$sex <- as.factor(metadata$sex)
traits <- metadata[, 7:43]
pc <- metadata[, c(1:5)]
# Calculate the correlation between each trait and each PC
correlation_matrix <- cor(traits, pc)
range(correlation_matrix)[1] -0.2850667 0.2787222correlation_matrix <- t(correlation_matrix)
# Create the heatmap using corrplot
corrplot(correlation_matrix, method = "color",
col = colorRampPalette(c("blue", "white", "red"))(200), # color palette
addCoef.col = "black", # Add correlation coefficients to the plot
number.cex = 0.4, # Adjust the font size of the numbers
tl.col = "black", # text label color
tl.srt = 90, # rotate text labels
tl.cex = 0.5,
title = "Correlation between Traits and expression PCs",
mar = c(0, 0, 1, 0)
)
Perform DESeq2 differential expression analysis for each trait
Model 1: expression ~ PRS + sex + expression PCs
Continuous PRS:
# Load the gene expression data
gene_expr_file <- "data/gene_reads_2017-06-05_v8_whole_blood.gct"
raw_count_df <- fread(gene_expr_file, header = TRUE, sep = "\t", drop = "id")
# load protein_coding list
protein_coding <- fread("data/protein-coding_gene.txt", sep = "\t")
protein_coding <- protein_coding[, c("symbol", "ensembl_gene_id")]
# keep only protein-coding genes
raw_count_df <- raw_count_df[raw_count_df$Description %in% protein_coding$symbol, ]
id <- raw_count_df$Name
raw_count <- raw_count_df[, -c(1:2)]
# modify GTEx sample names matching names used in PRS data
colnames(raw_count) <- sub("^(GTEX-[^-.]+).*", "\\1", colnames(raw_count))
matching_samples <- intersect(rownames(metadata), colnames(raw_count))
final_count <- raw_count[ , ..matching_samples]
# prefilter: keep only rows that have a count of at least 10
keep_genes <- rowSums(final_count >= 10) > 0
final_count <- final_count[keep_genes, ]
id <- id[keep_genes]
dim(final_count)
# Loop through each trait and run DESeq2
for (trait in colnames(traits)) {
# Standardize PRS for the current trait
prs_trait <- traits[,trait]
prs_trait <- scale(prs_trait) # Standardize PRS to mean = 0, sd = 1
# Add the standardized PRS to the metadata for continuous trait
metadata[,trait] <- prs_trait
# Create the DESeqDataSet for the current trait
dds <- DESeqDataSetFromMatrix(
countData = as.matrix(final_count), # Raw counts
colData = metadata[, c(1:6, which(colnames(metadata) == trait))],
design = as.formula(paste("~ PC1 + PC2 + PC3 + PC4 + PC5 + sex +", trait))
)
rownames(dds) <- id
# Run DESeq2 analysis
dds <- DESeq(dds, parallel = TRUE, BPPARAM = MulticoreParam(4))
# Get the results for the current trait
res <- results(dds)
# Save the results to a file
write.csv(res, paste0("differential_expression_", trait, "_results.csv"))
# print a summary of the results
print(paste("Results for trait:", trait))
print(summary(res))
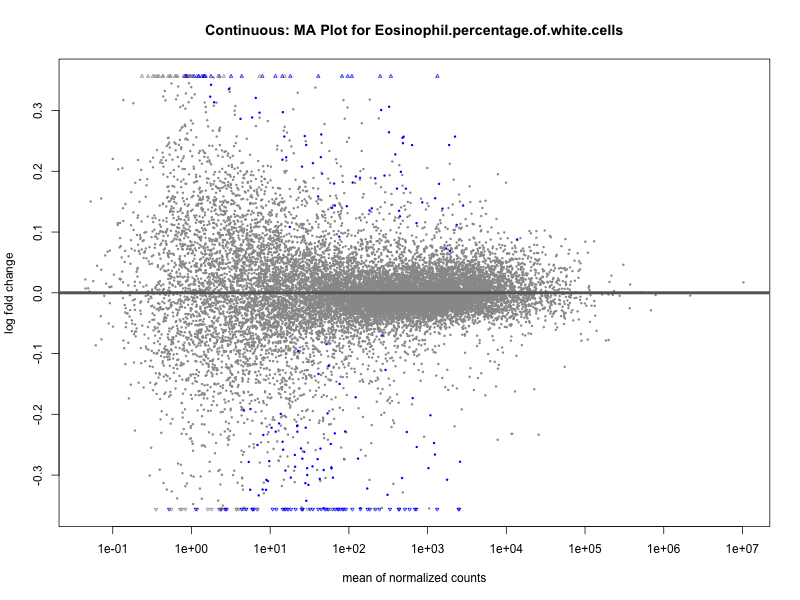
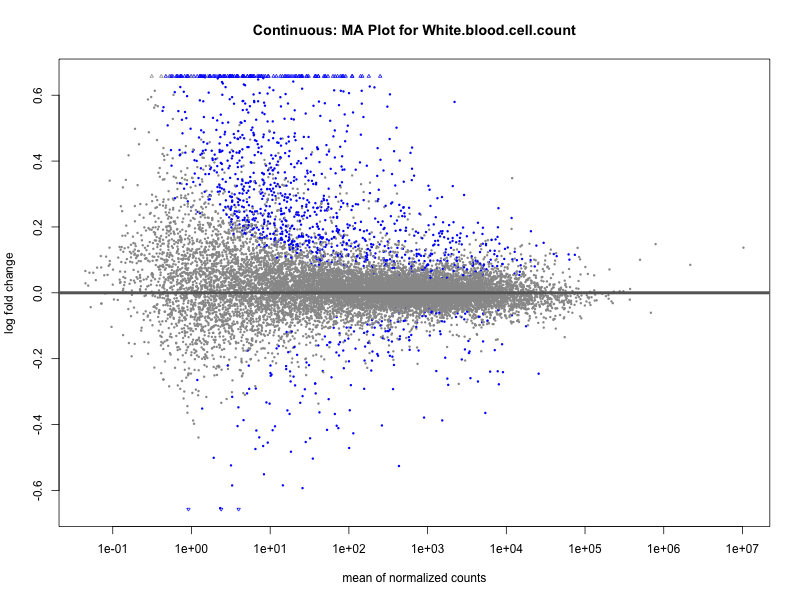
# plot the MA-plot for the current trait
png(paste0("ma_plot_", trait, ".png"), width = 800, height = 600)
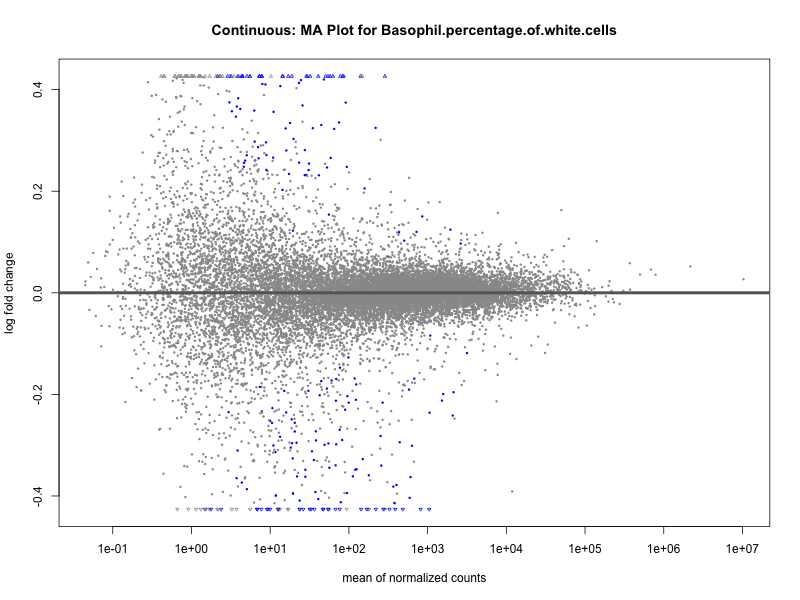
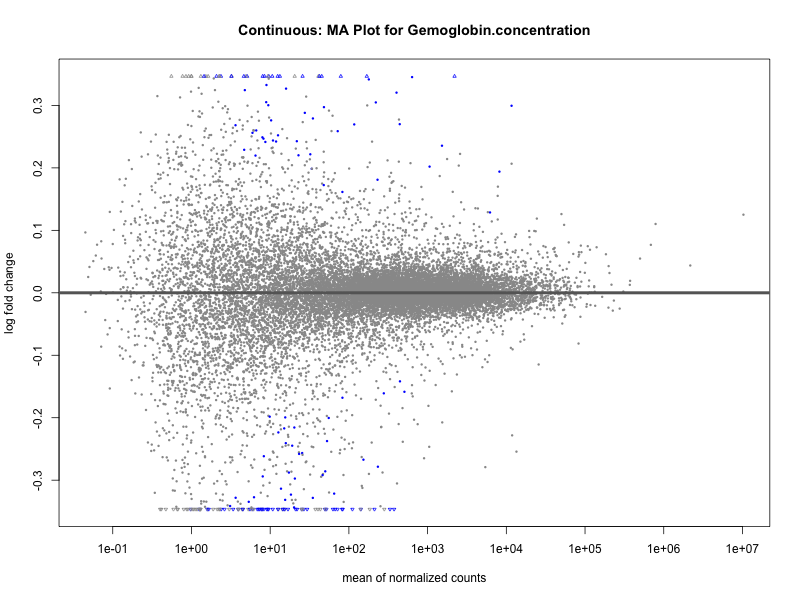
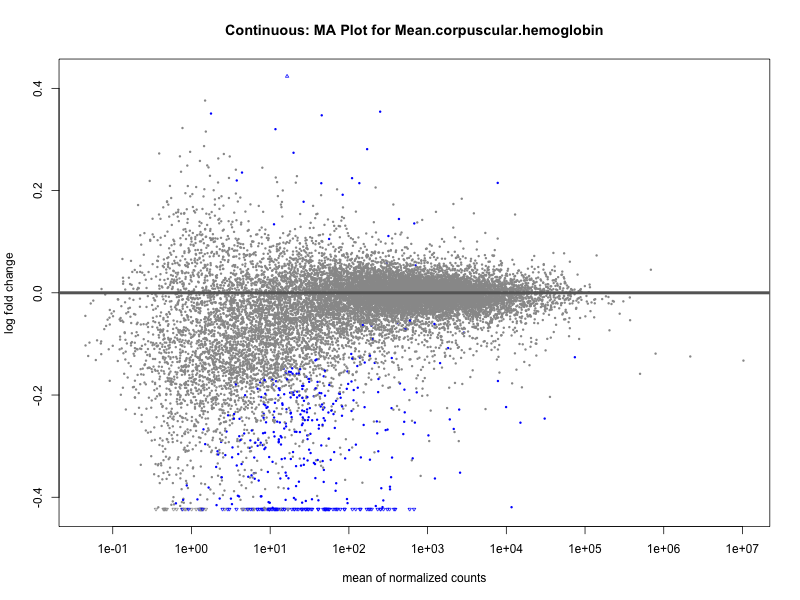
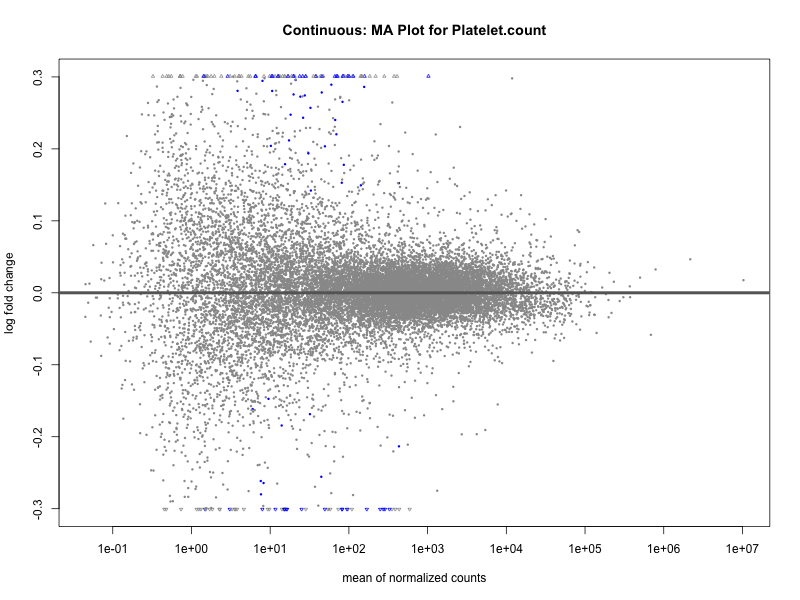
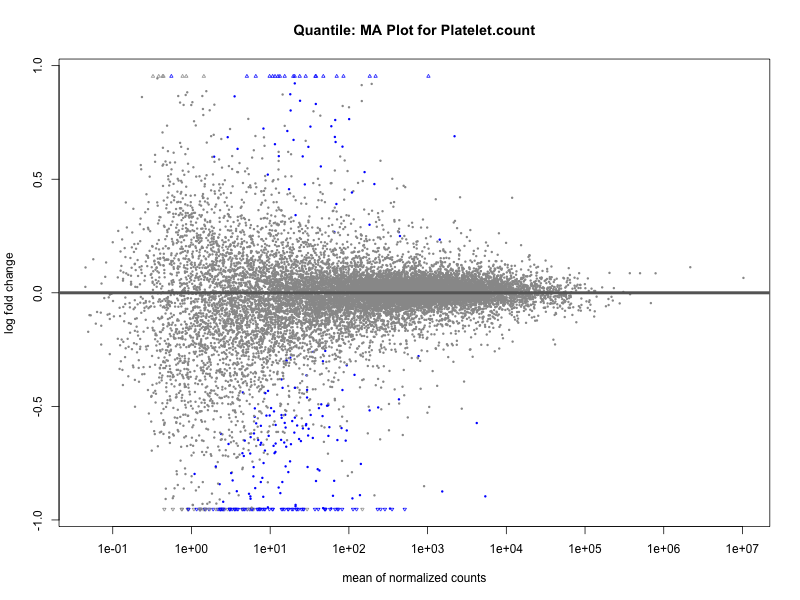
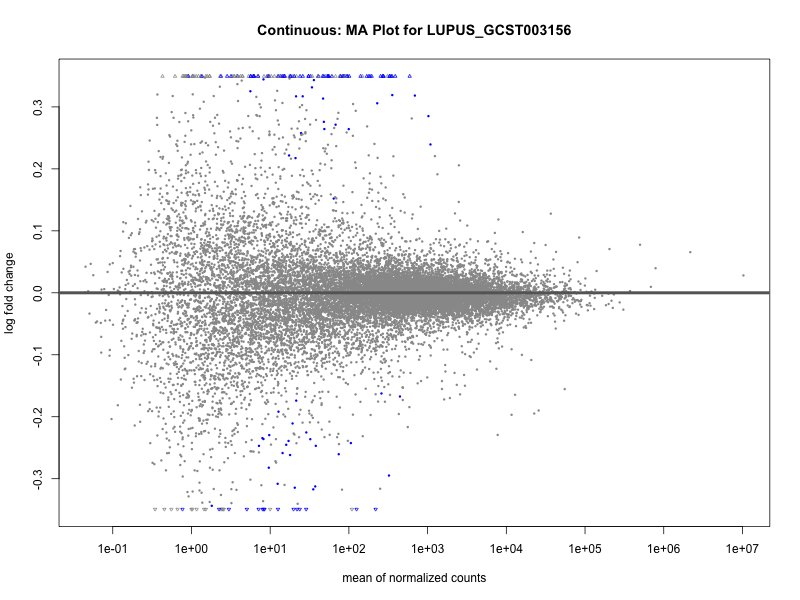
plotMA(res, main = paste("Continuous: MA Plot for", trait))
dev.off()
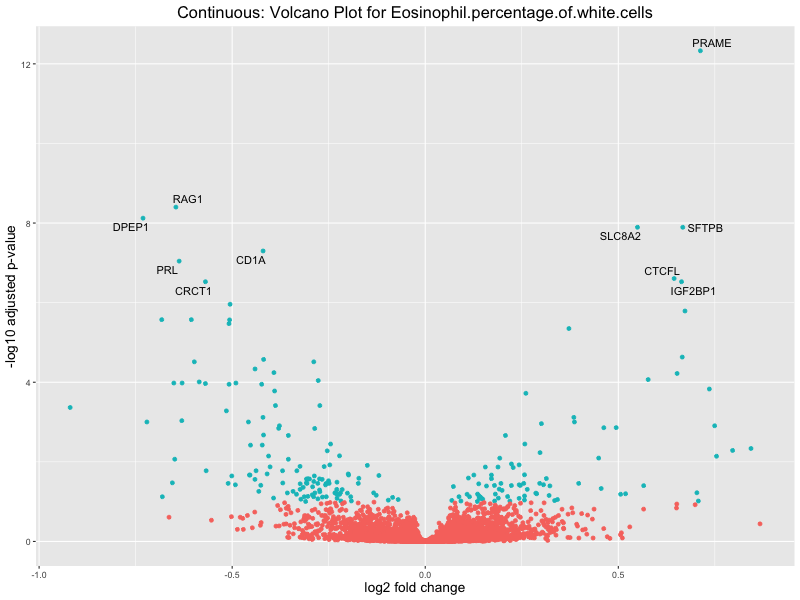
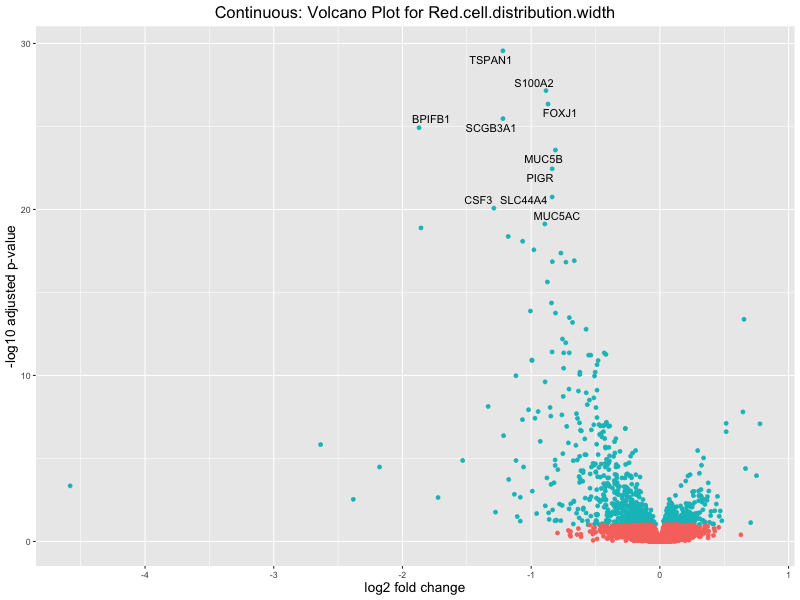
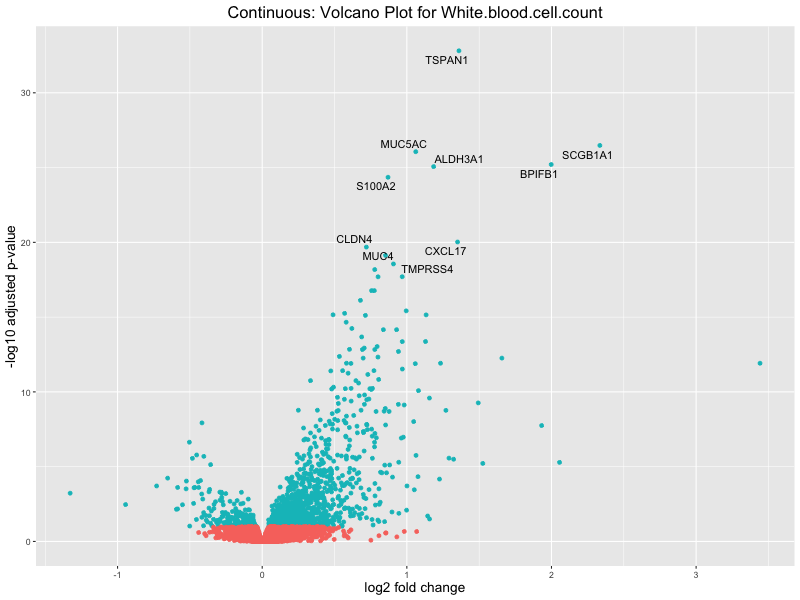
# volcano plot
res_tableOE <- as.data.frame(res)
res_tableOE$gene_name <- raw_count_df$Description[keep_genes]
res_tableOE <- mutate(res_tableOE, threshold_OE = padj < 0.1)
res_tableOE <- res_tableOE %>% arrange(padj) %>% mutate(genelabels = "")
res_tableOE$genelabels[1:10] <- res_tableOE$gene_name[1:10]
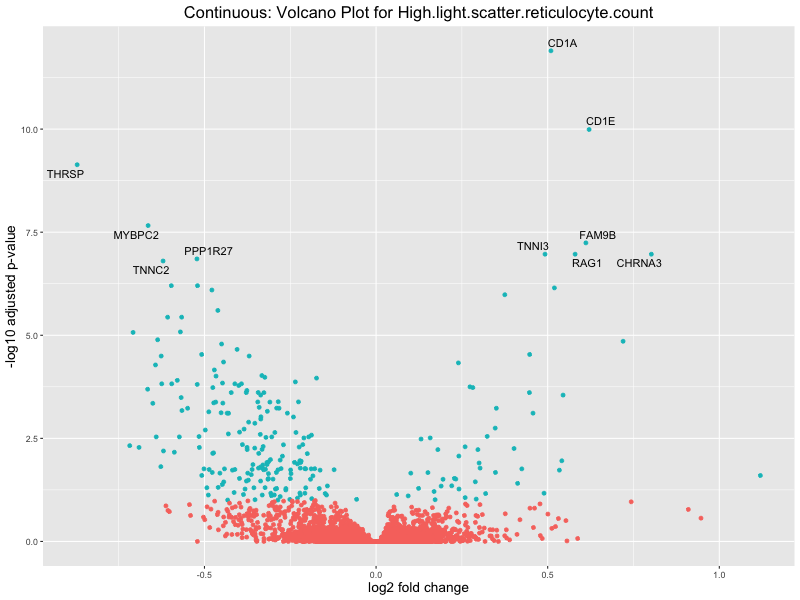
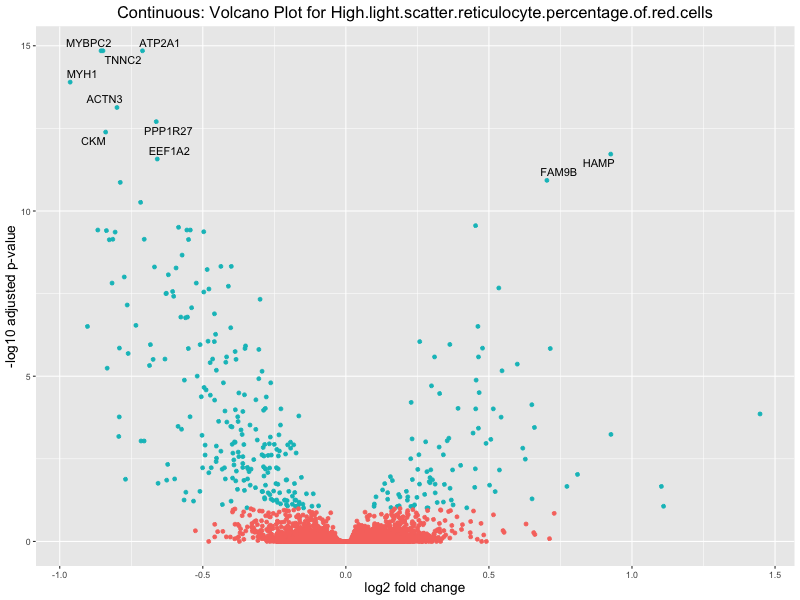
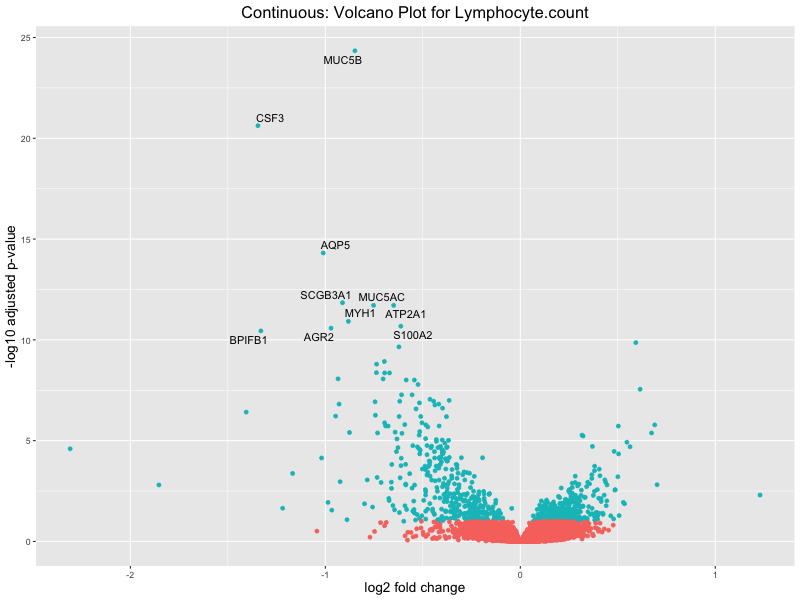
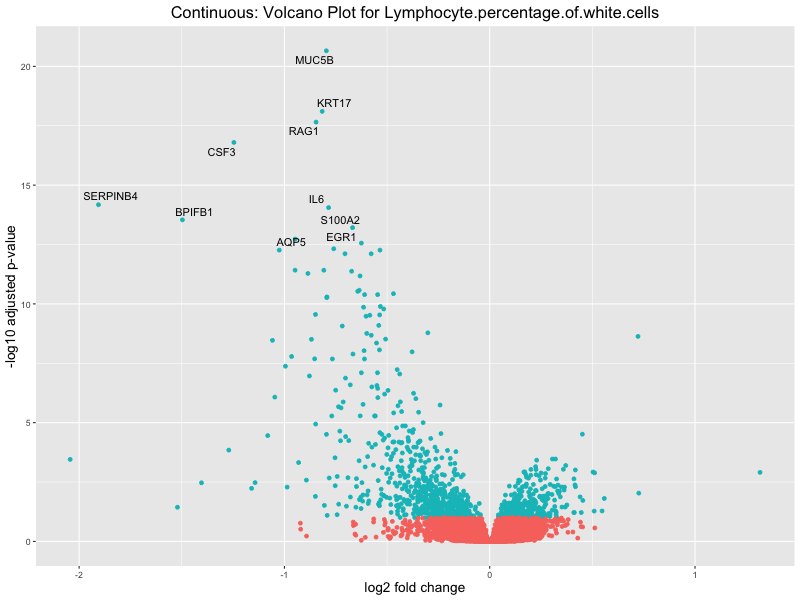
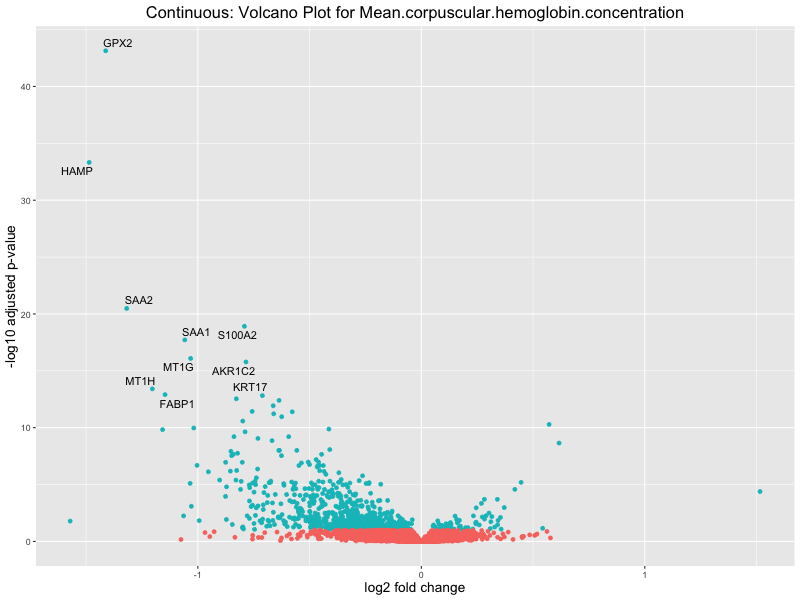
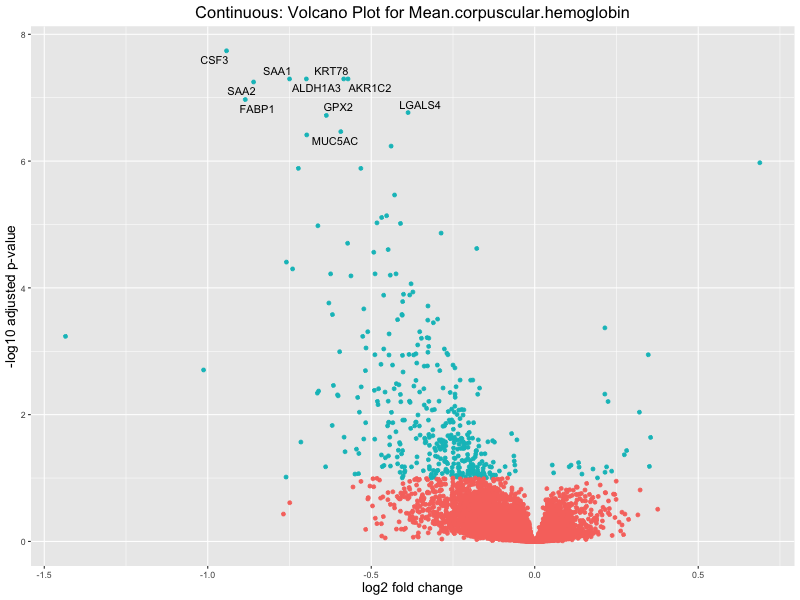
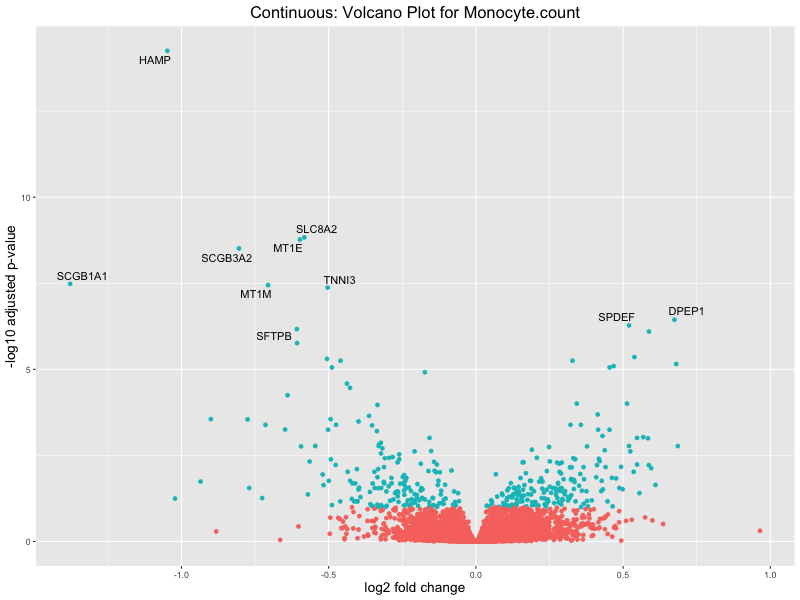
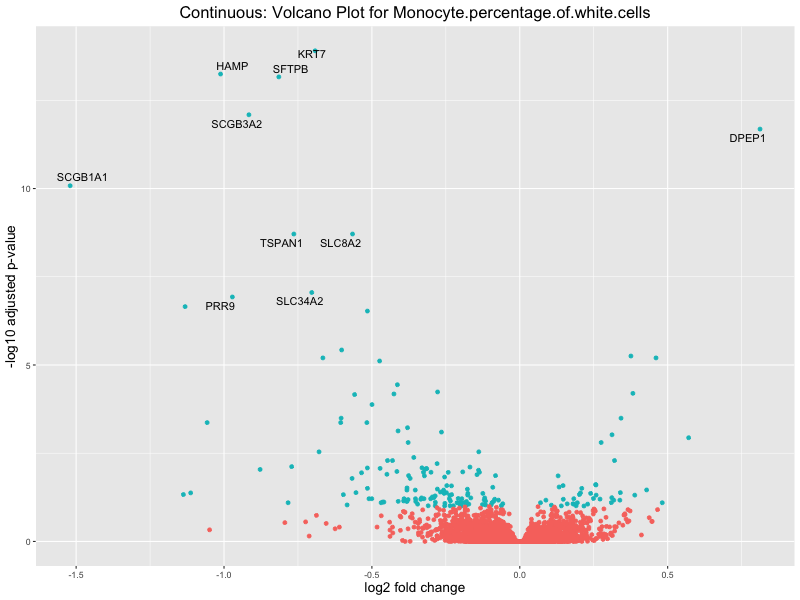
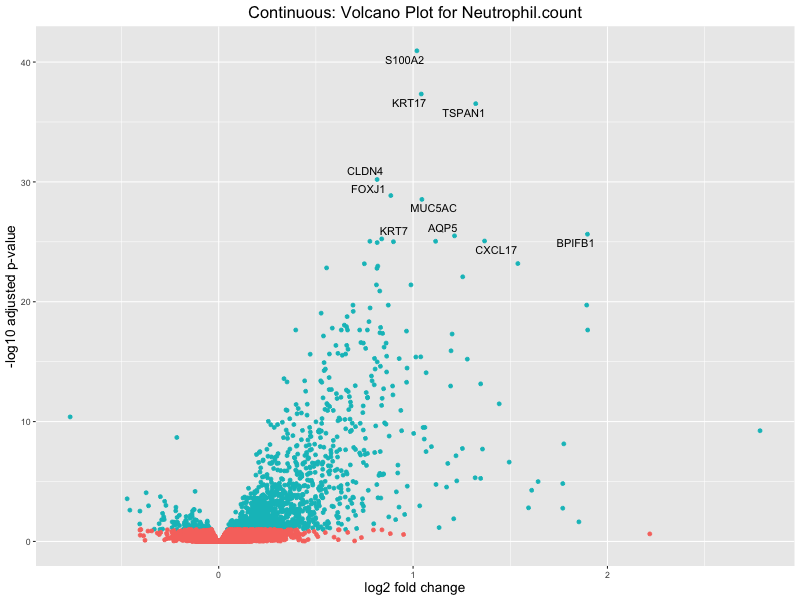
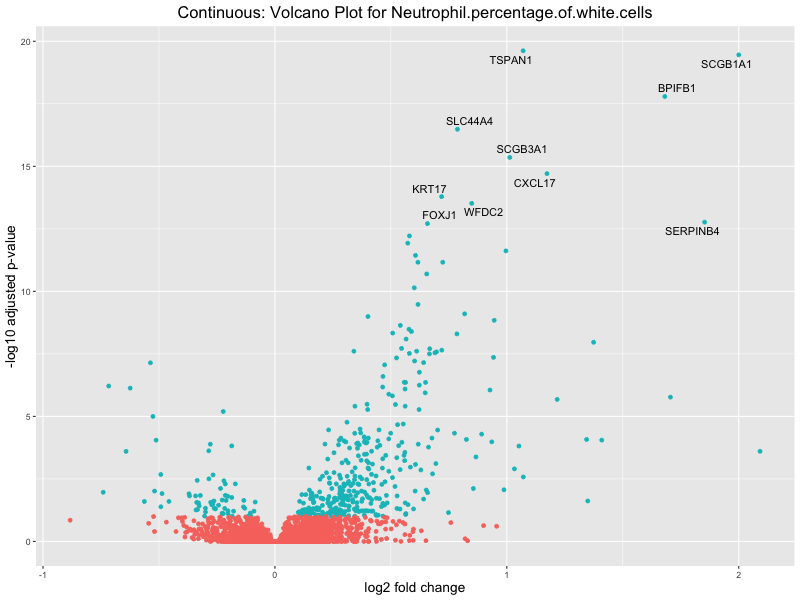
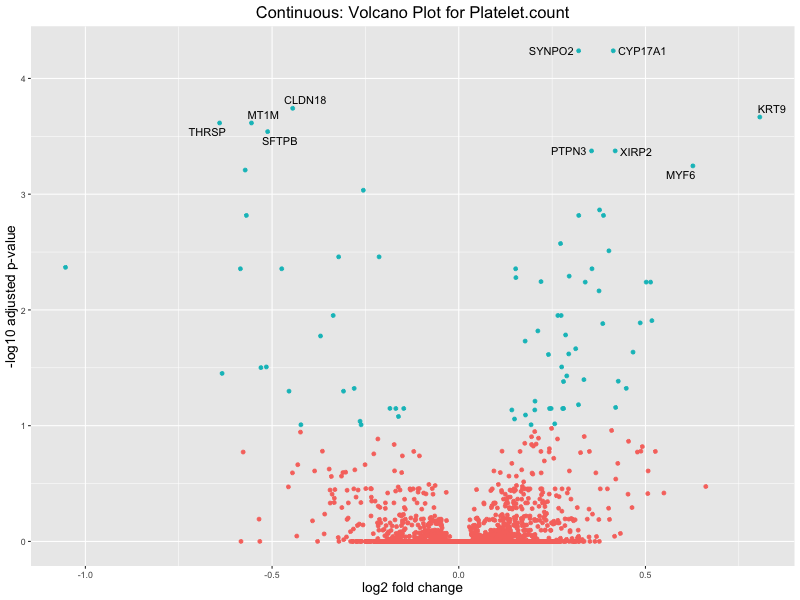
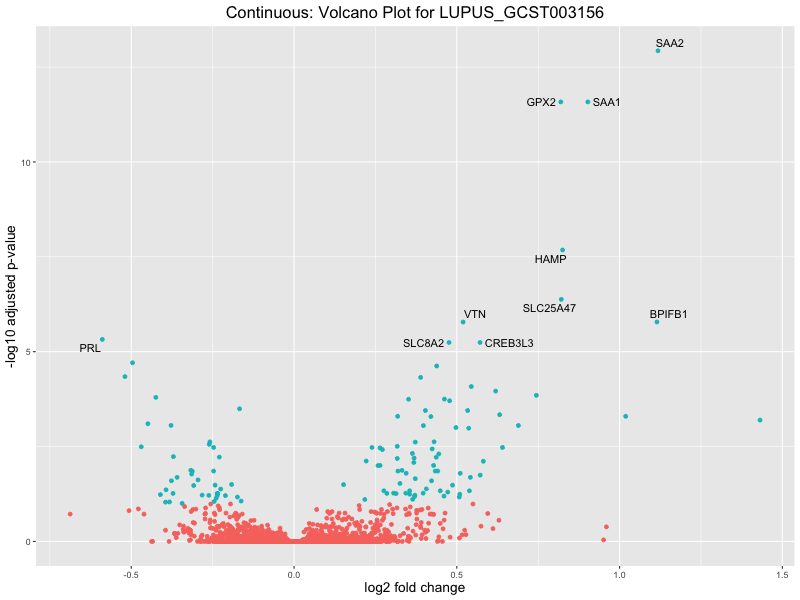
volcano_plot <- ggplot(res_tableOE, aes(x = log2FoldChange, y = -log10(padj))) +
geom_point(aes(colour = threshold_OE)) +
geom_text_repel(aes(label = genelabels)) +
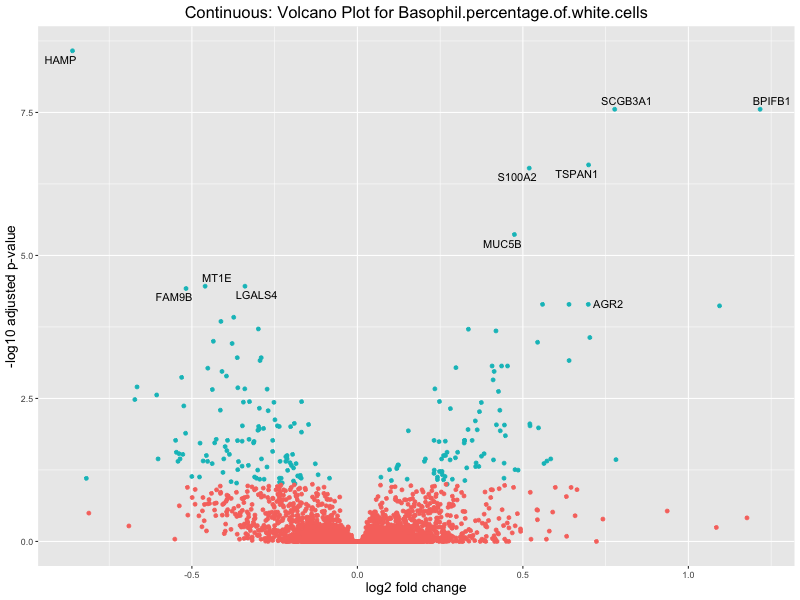
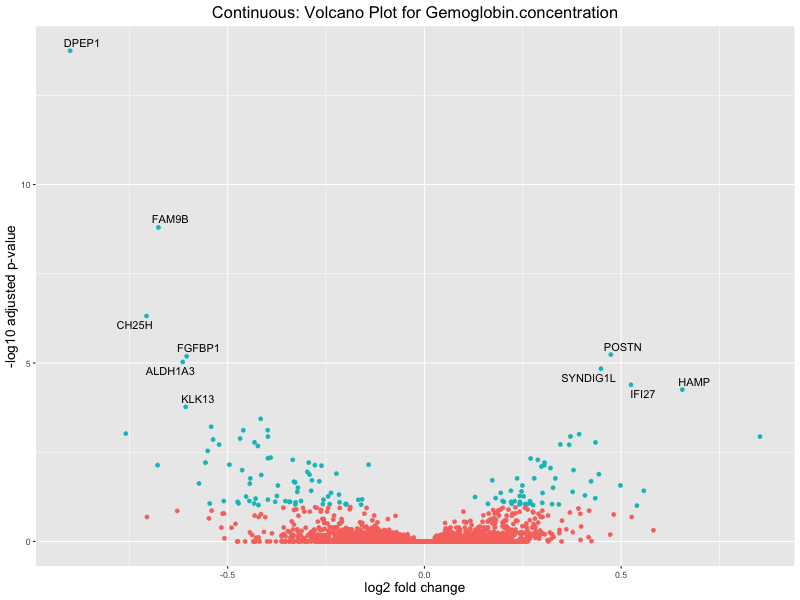
ggtitle(paste("Continuous: Volcano Plot for", trait)) +
xlab("log2 fold change") +
ylab("-log10 adjusted p-value") +
theme(legend.position = "none",
plot.title = element_text(size = rel(1.5), hjust = 0.5),
axis.title = element_text(size = rel(1.25)))
# Save the volcano plot
png(paste0("volcano_plot_", trait, ".png"), width = 800, height = 600)
print(volcano_plot)
dev.off()
}Quantile PRS:
metadata_file <- "analysis/metadata_quantile.txt"
metadata <- read.csv(metadata_file, header = T, sep = "\t", stringsAsFactors = T)
metadata$sex <- as.factor(metadata$sex)
traits <- metadata[, 7:43]
# Loop through each trait and run DESeq2
for (trait in colnames(traits)) {
# Create the DESeqDataSet for the current trait
dds <- DESeqDataSetFromMatrix(
countData = as.matrix(final_count), # Raw counts
colData = metadata[, c(1:6, which(colnames(metadata) == trait))],
design = as.formula(paste("~ PC1 + PC2 + PC3 + PC4 + PC5 + sex +", trait))
)
rownames(dds) <- id
# Run DESeq2 analysis
dds <- DESeq(dds, parallel = TRUE, BPPARAM = MulticoreParam(4))
# Get the results for the current trait
res <- results(dds)
# Save the results to a file
write.csv(res, paste0("differential_expression_", trait, "_quantile_results.csv"))
# print a summary of the results
print(paste("Results for trait:", trait))
print(summary(res))
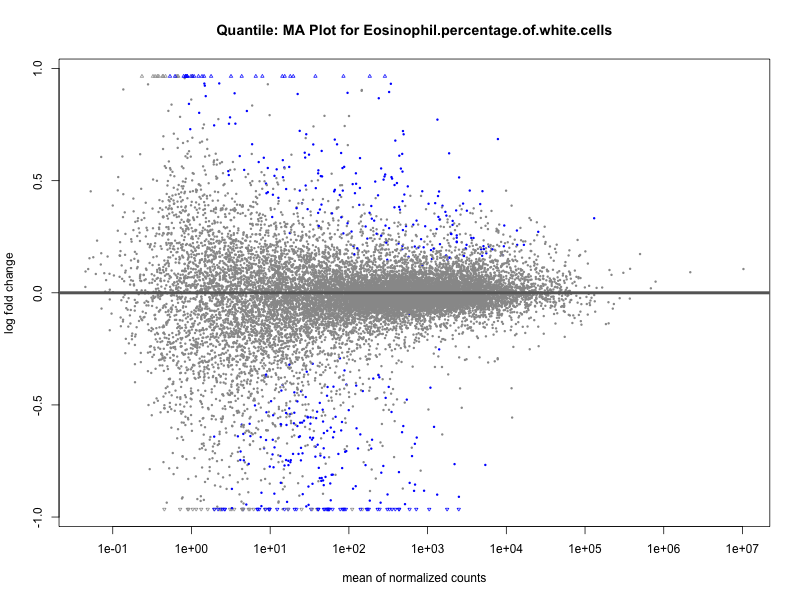
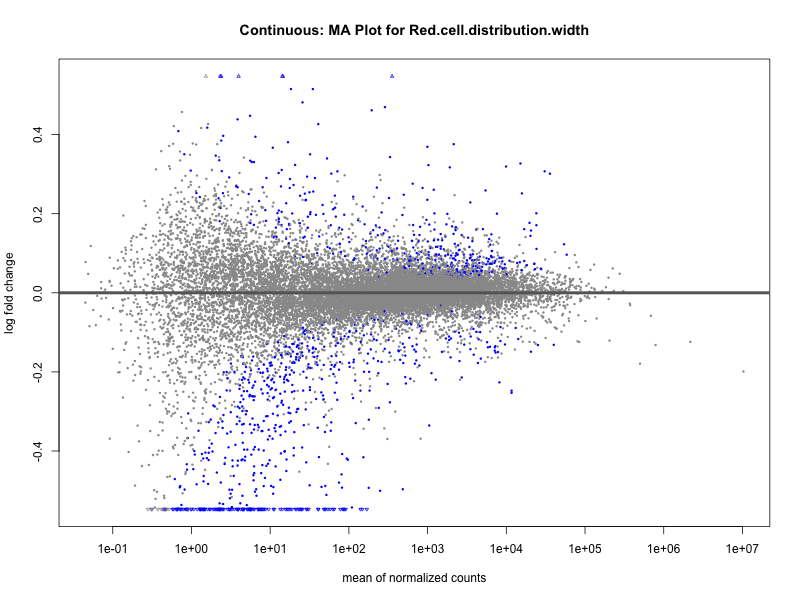
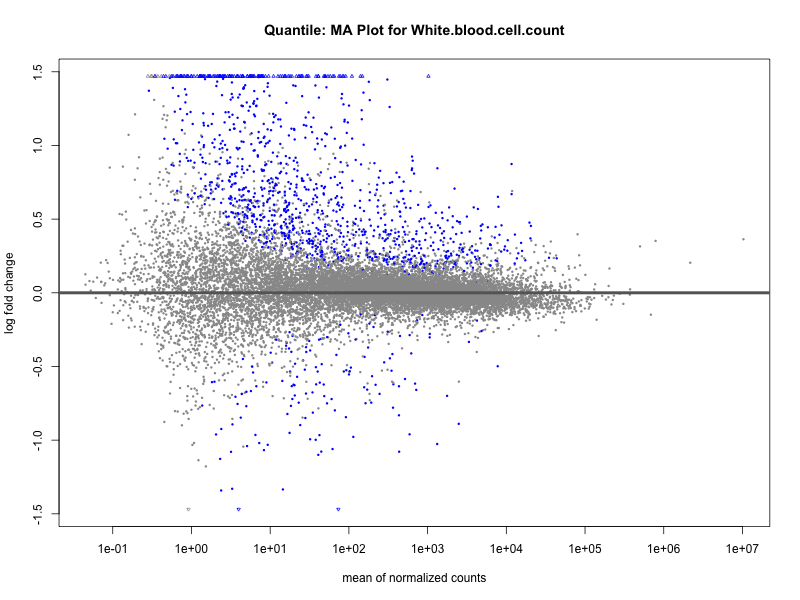
# plot the MA-plot for the current trait
png(paste0("ma_plot_quantile_", trait, ".png"), width = 800, height = 600)
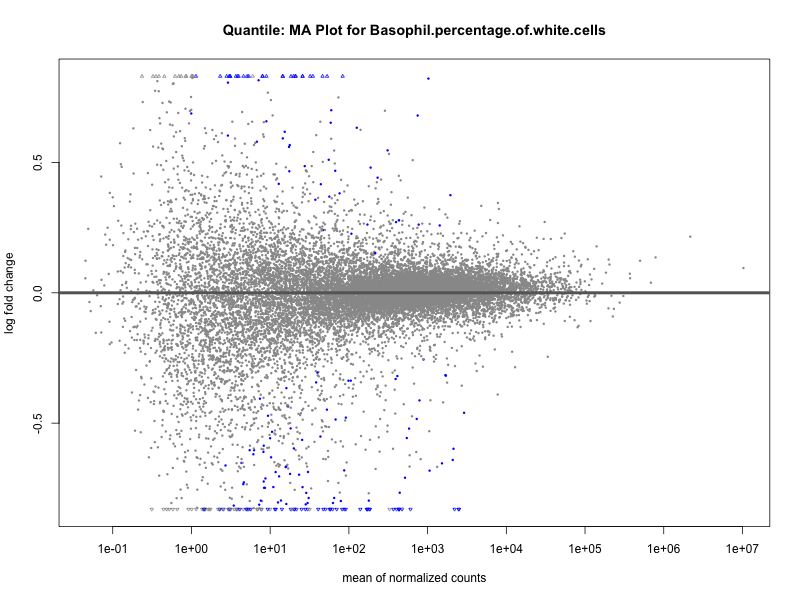
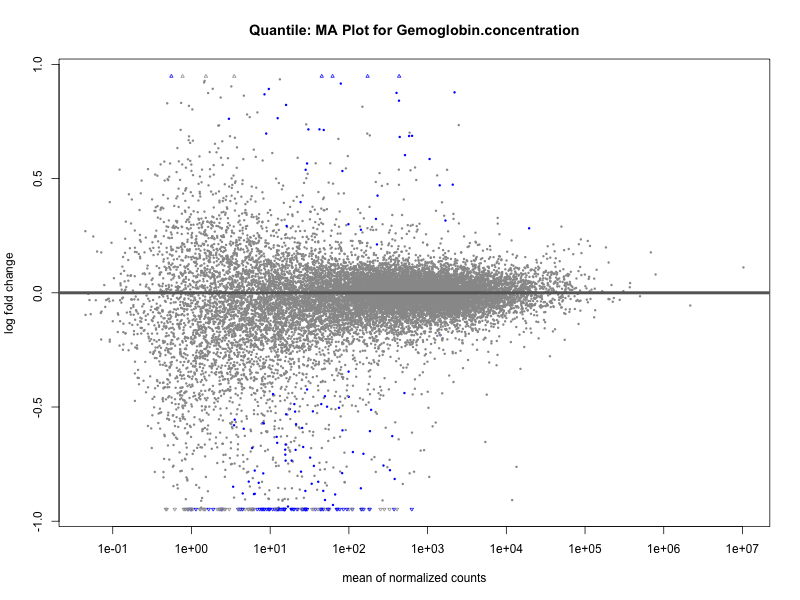
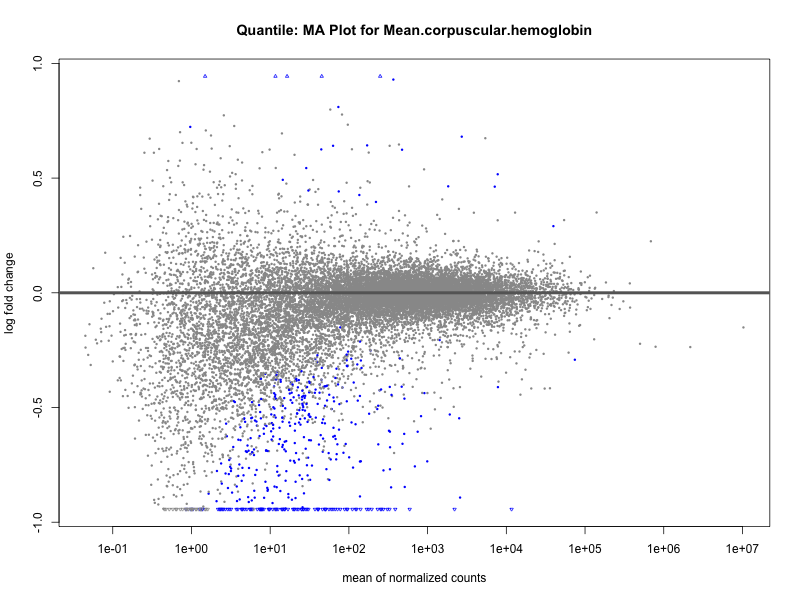
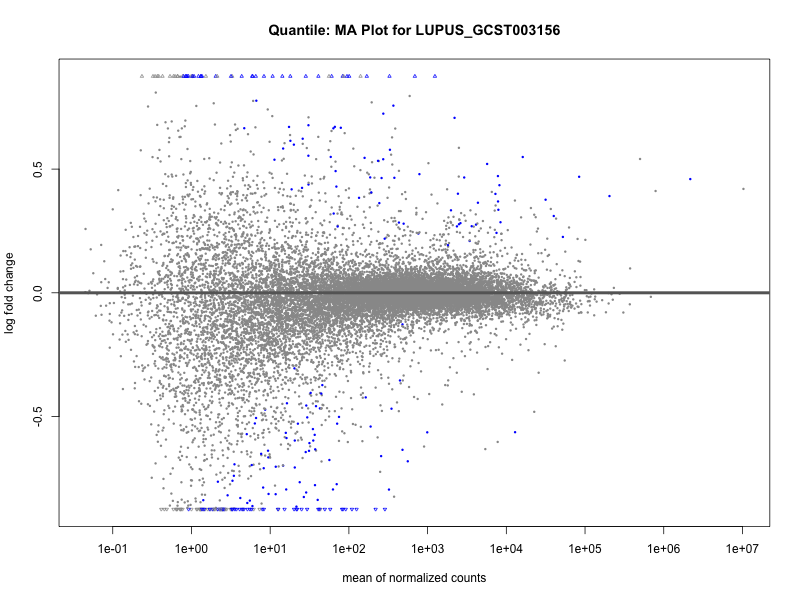
plotMA(res, main = paste("Quantile: MA Plot for", trait))
dev.off()
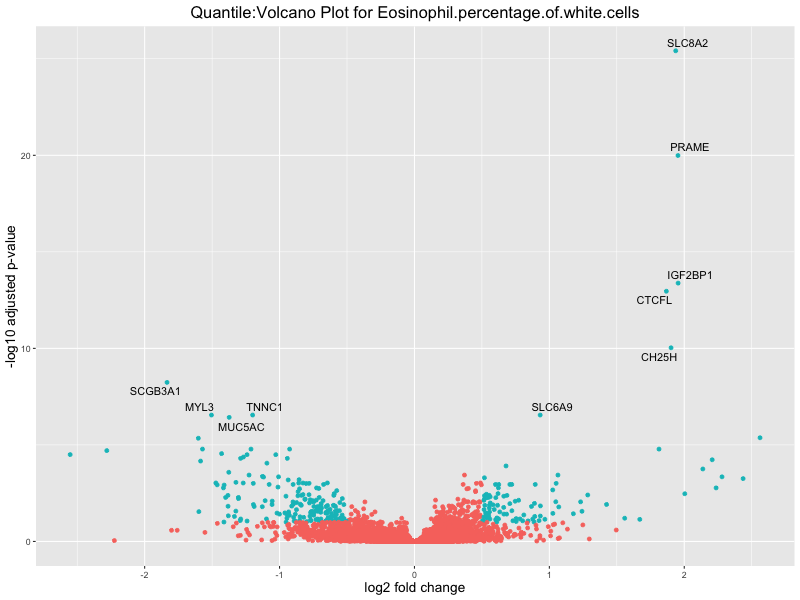
# volcano plot
res_tableOE <- as.data.frame(res)
res_tableOE$gene_name <- raw_count_df$Description[keep_genes]
res_tableOE <- mutate(res_tableOE, threshold_OE = padj < 0.1 &
abs(log2FoldChange) >= 0.5)
res_tableOE <- res_tableOE %>% arrange(padj) %>% mutate(genelabels = "")
res_tableOE$genelabels[1:10] <- res_tableOE$gene_name[1:10]
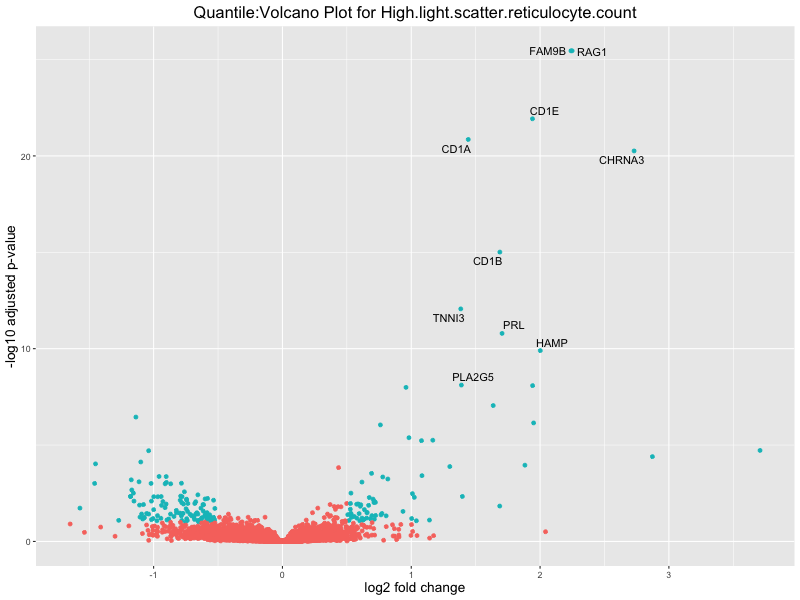
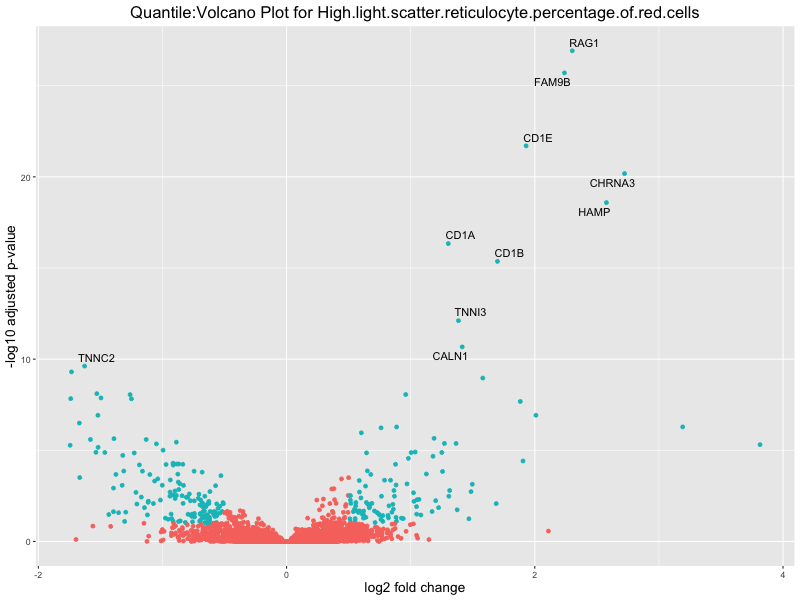
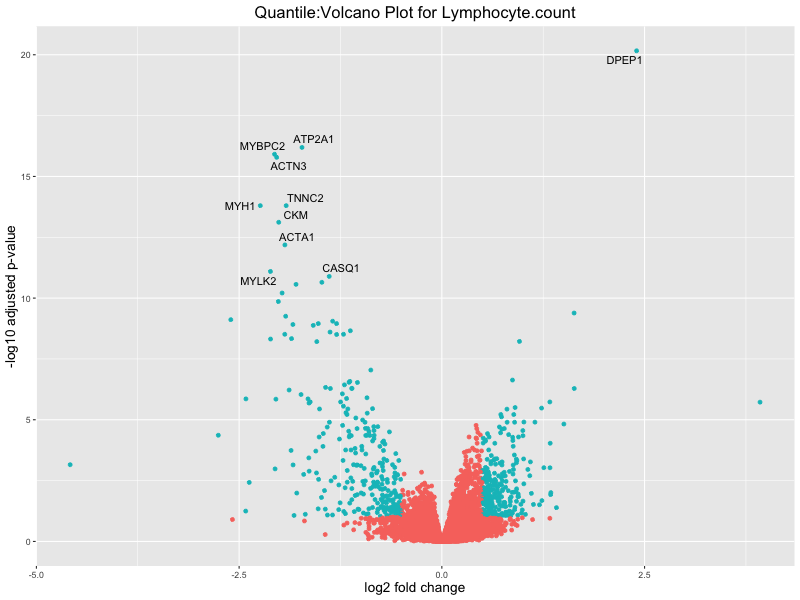
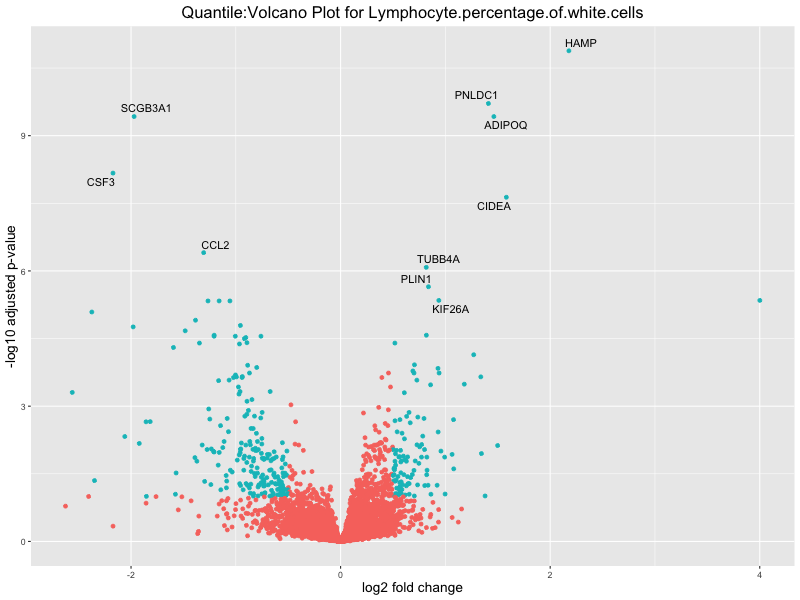
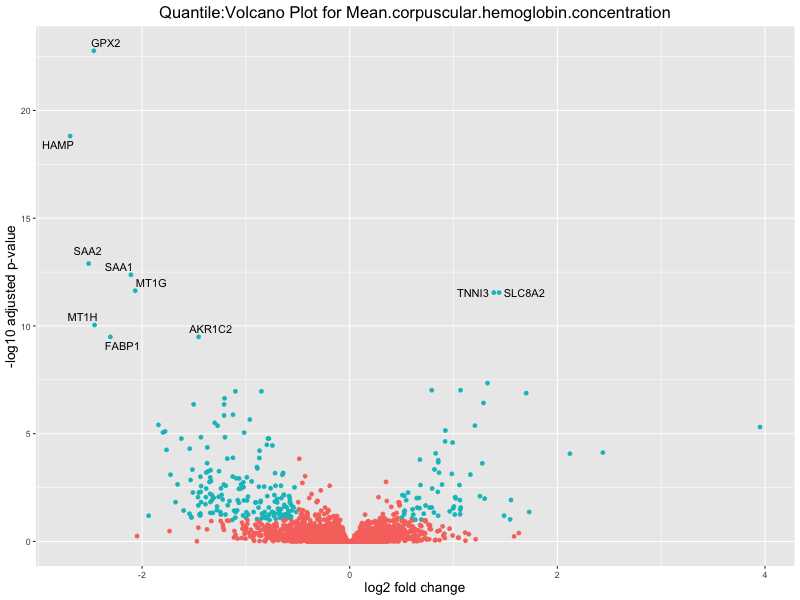
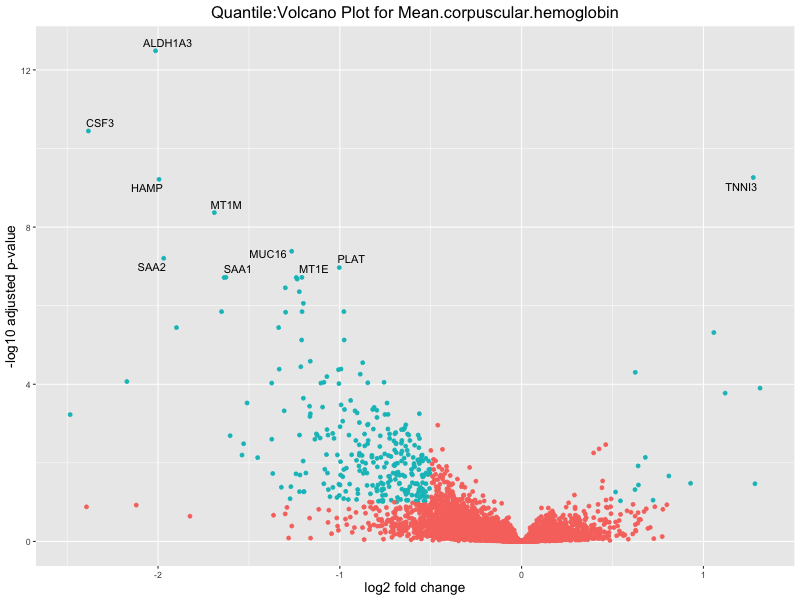
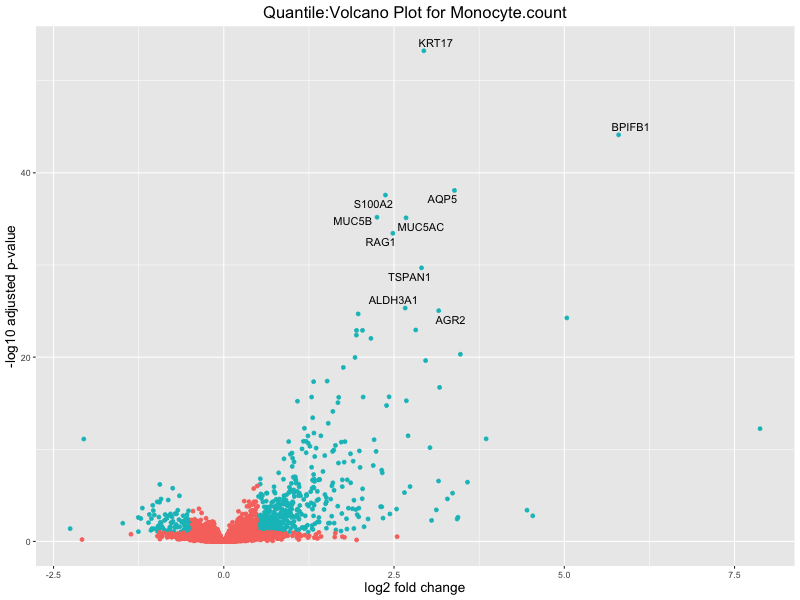
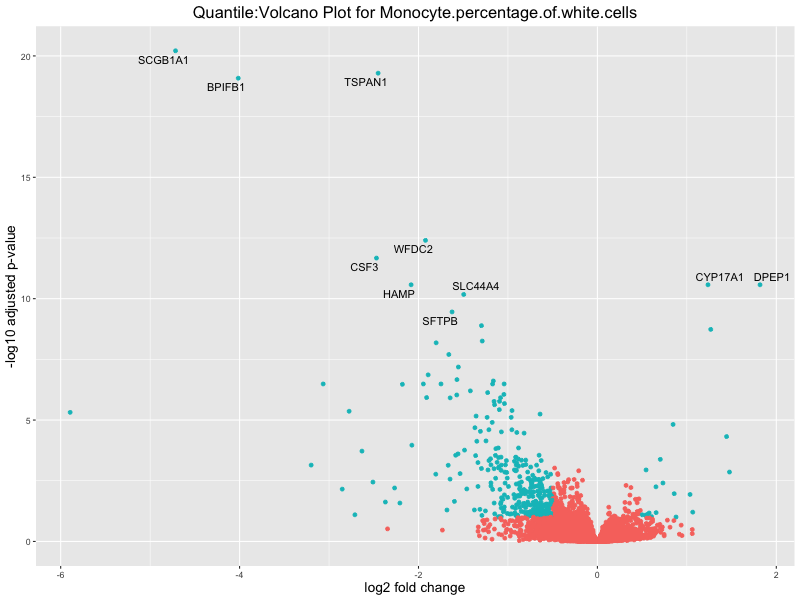
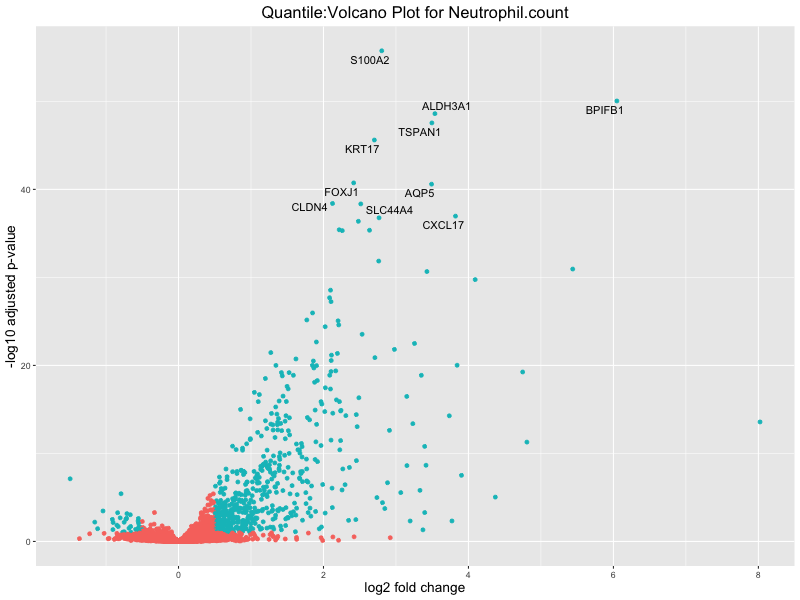
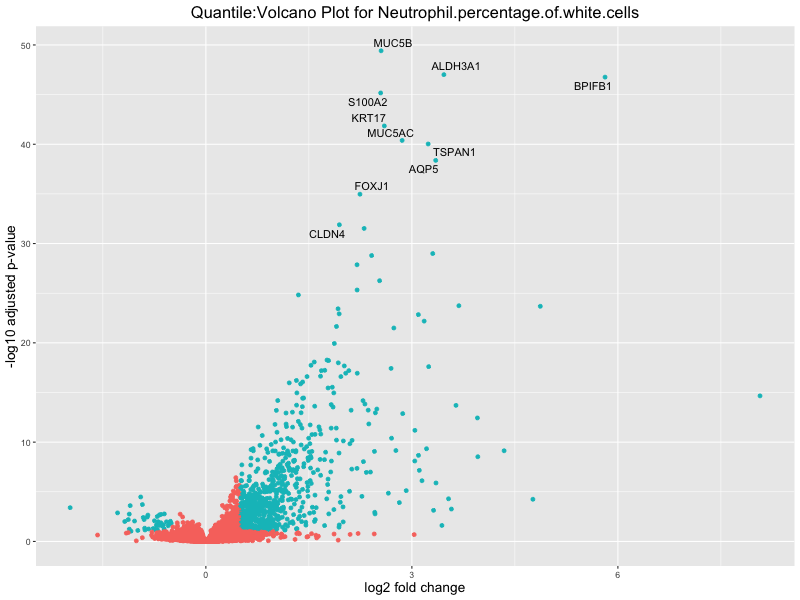
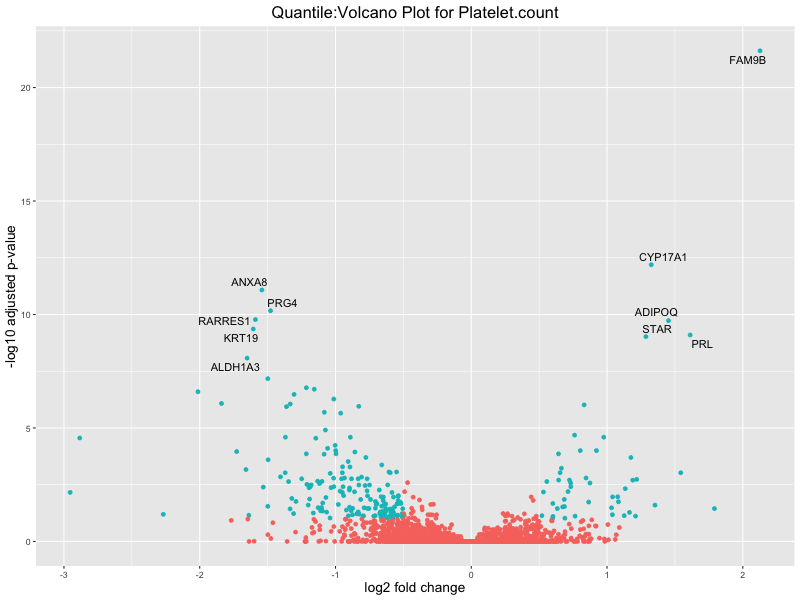
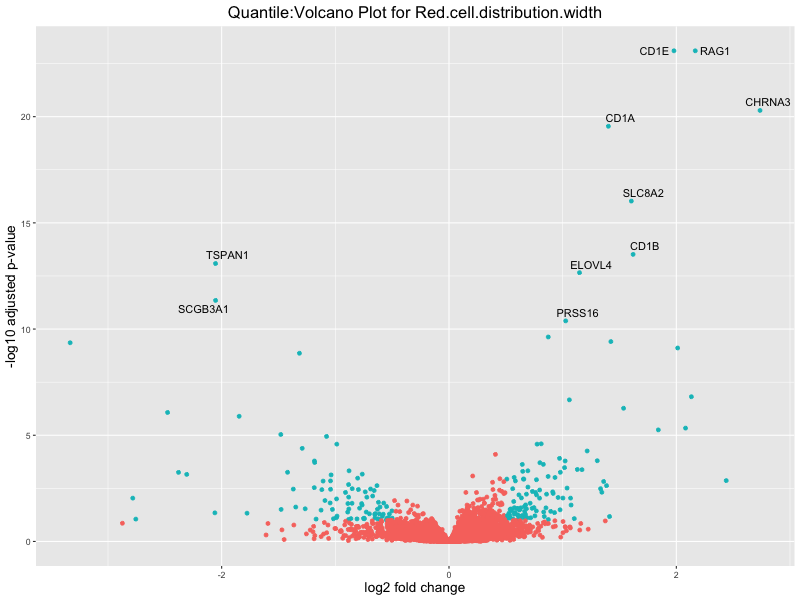
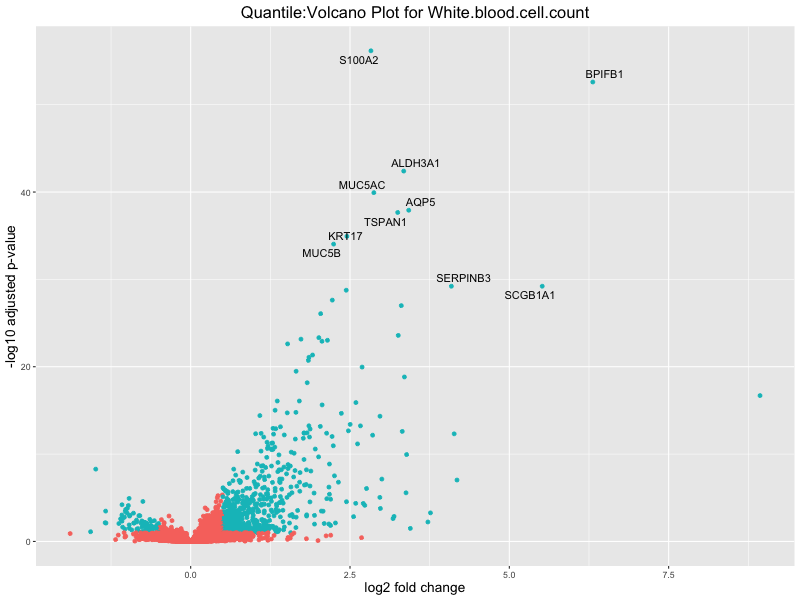
volcano_plot <- ggplot(res_tableOE, aes(x = log2FoldChange, y = -log10(padj))) +
geom_point(aes(colour = threshold_OE)) +
geom_text_repel(aes(label = genelabels)) +
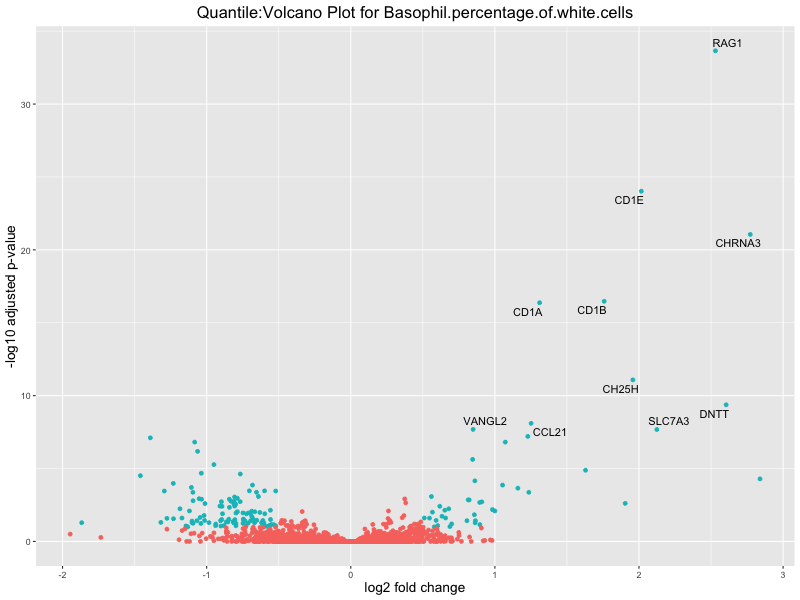
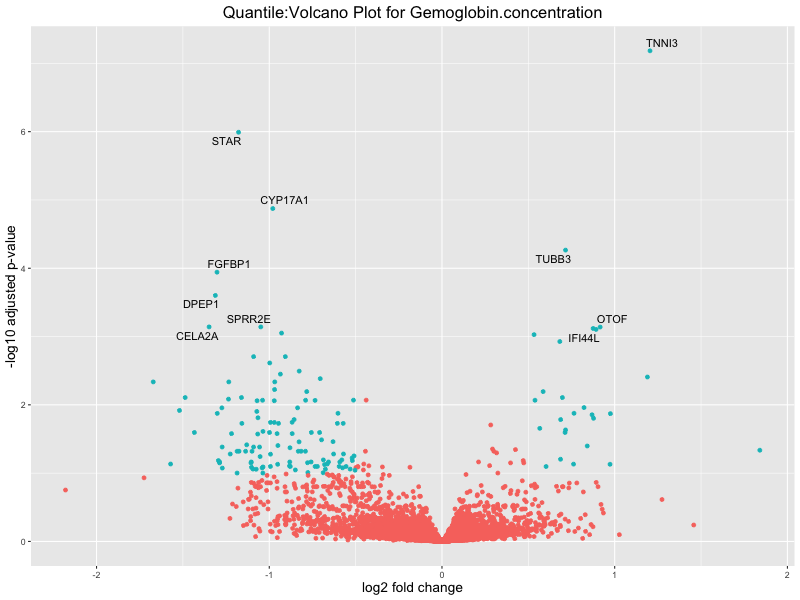
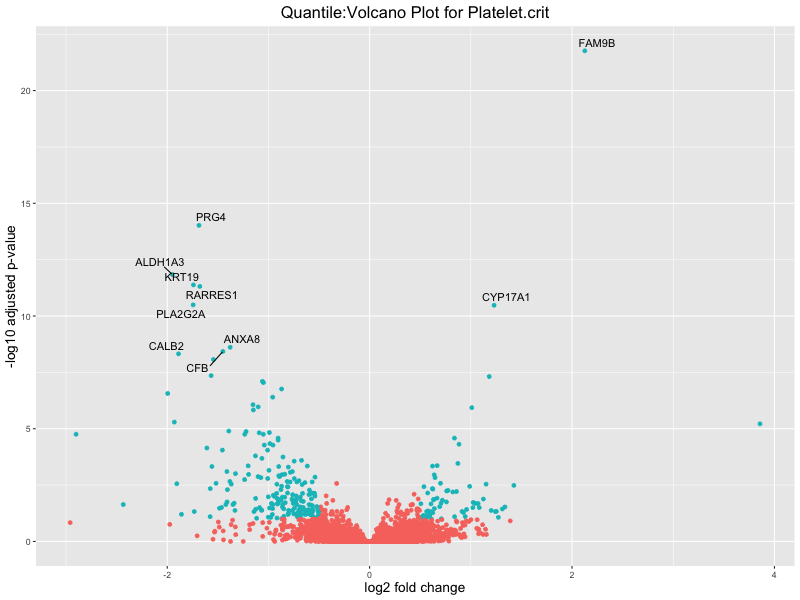
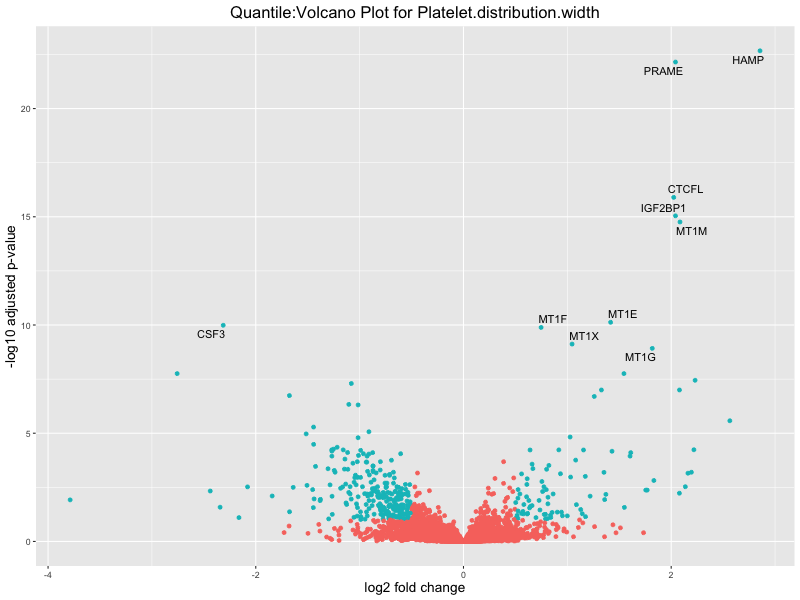
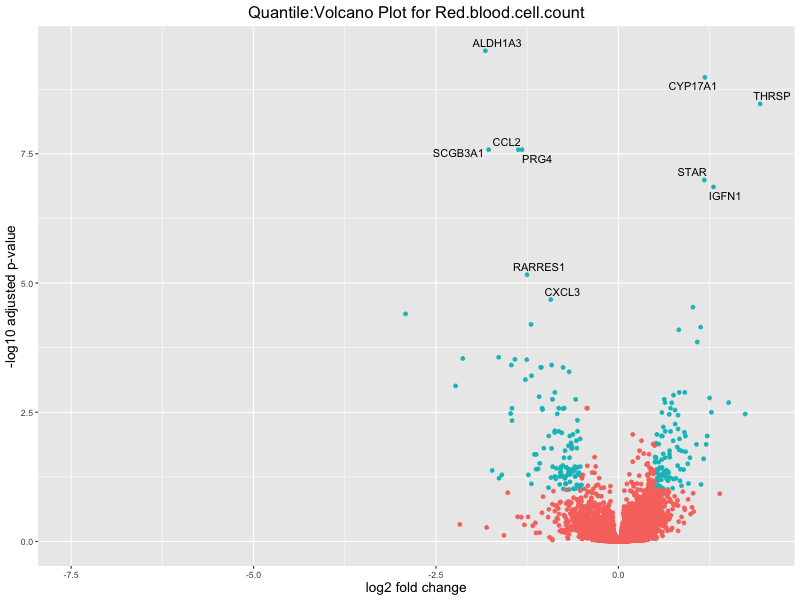
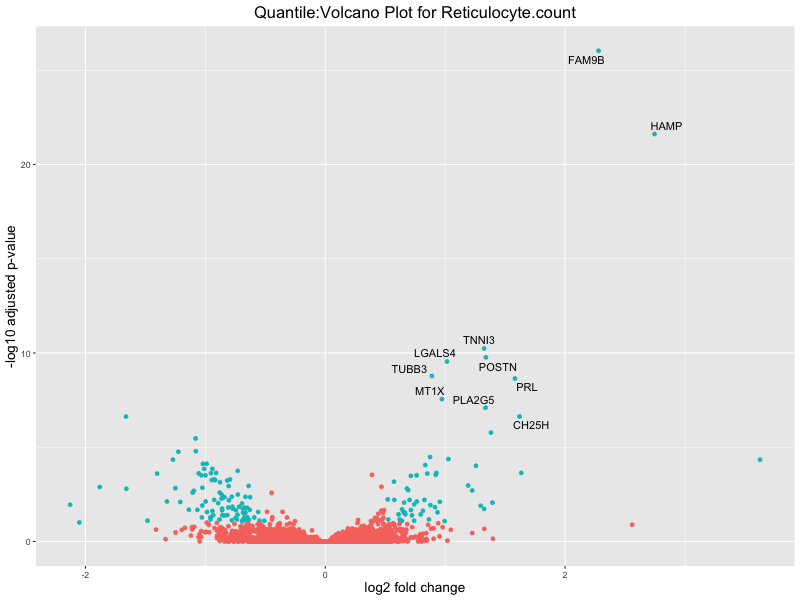
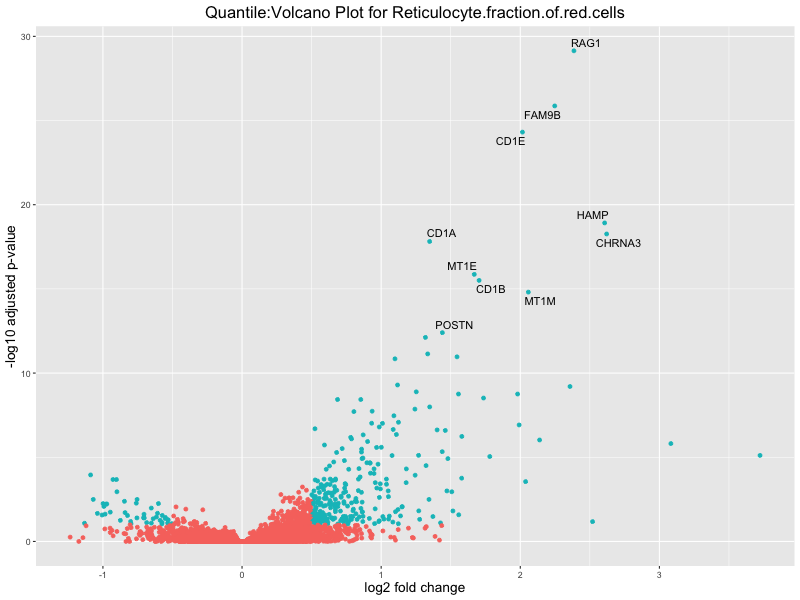
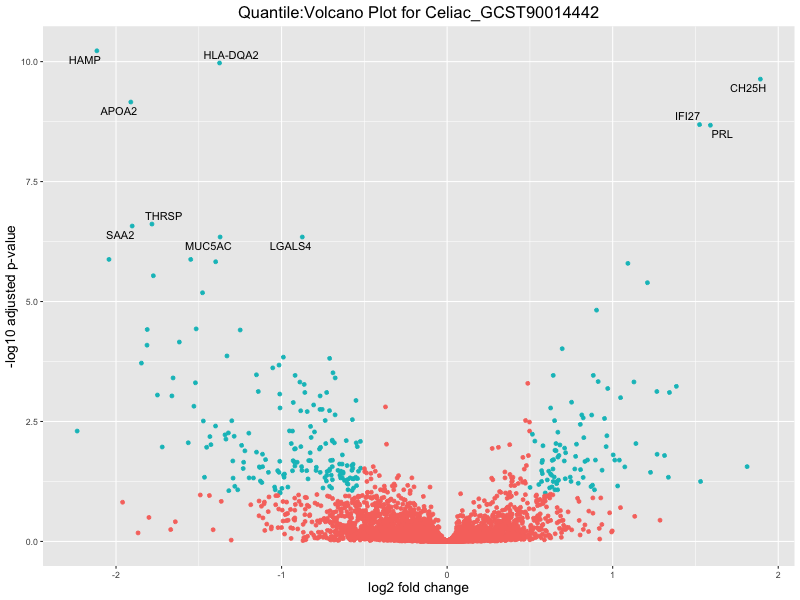
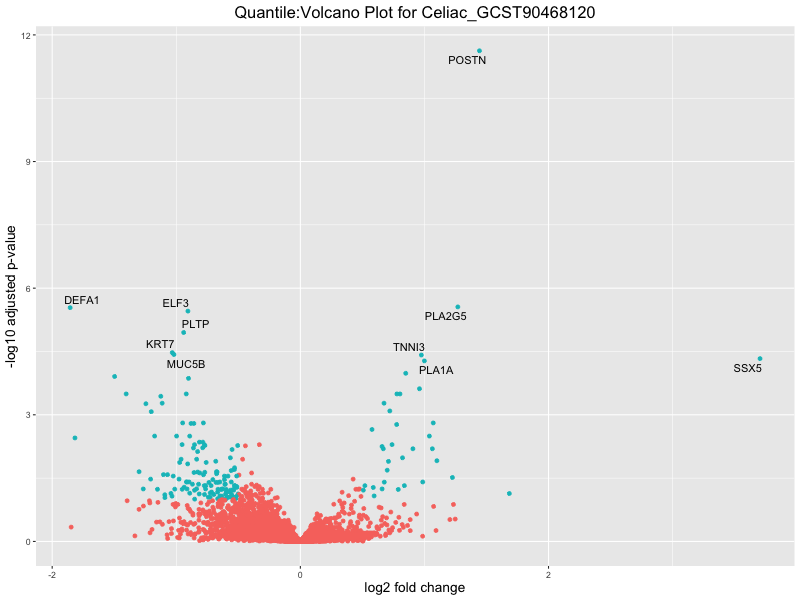
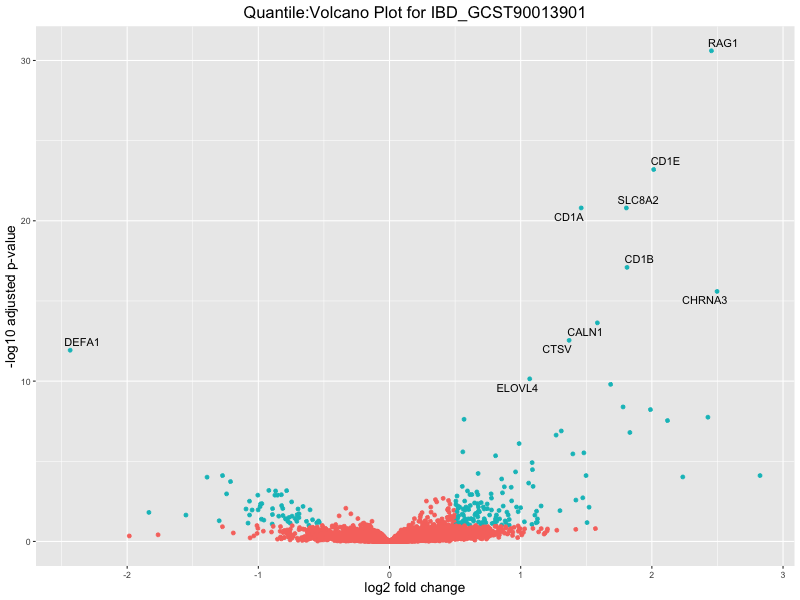
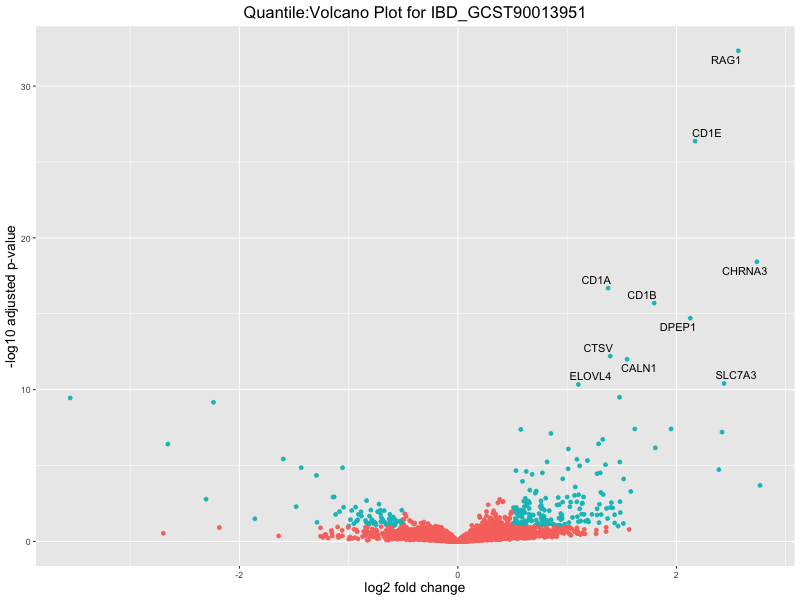
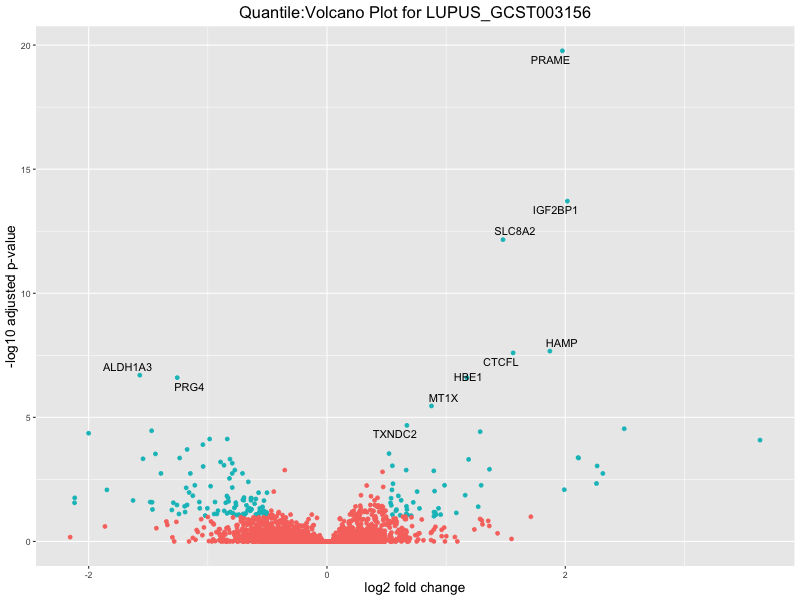
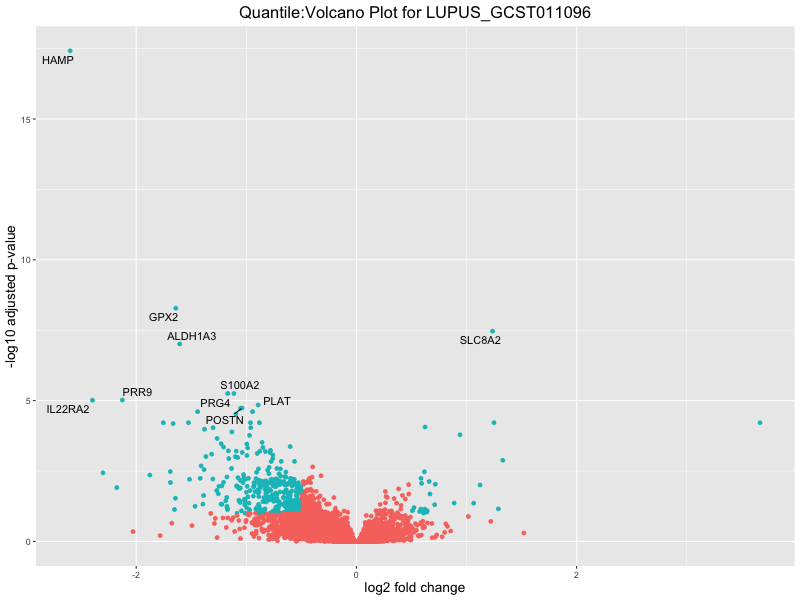
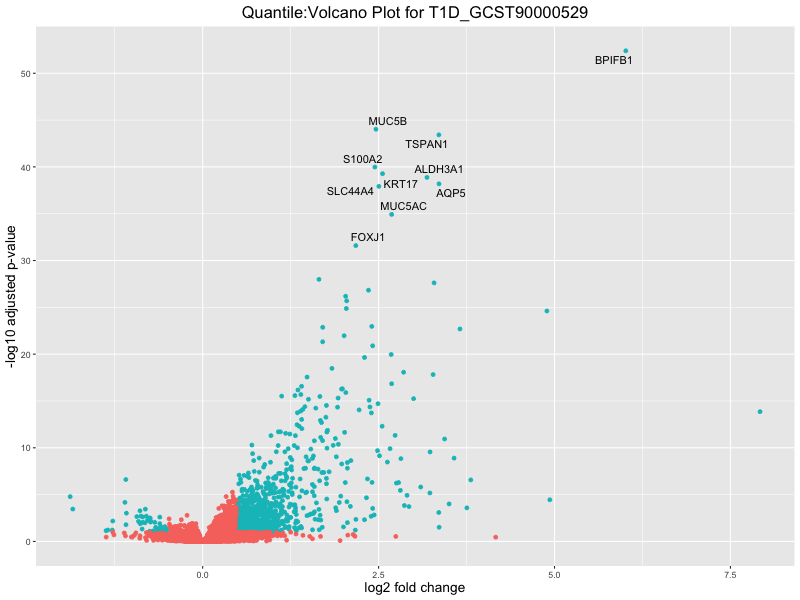
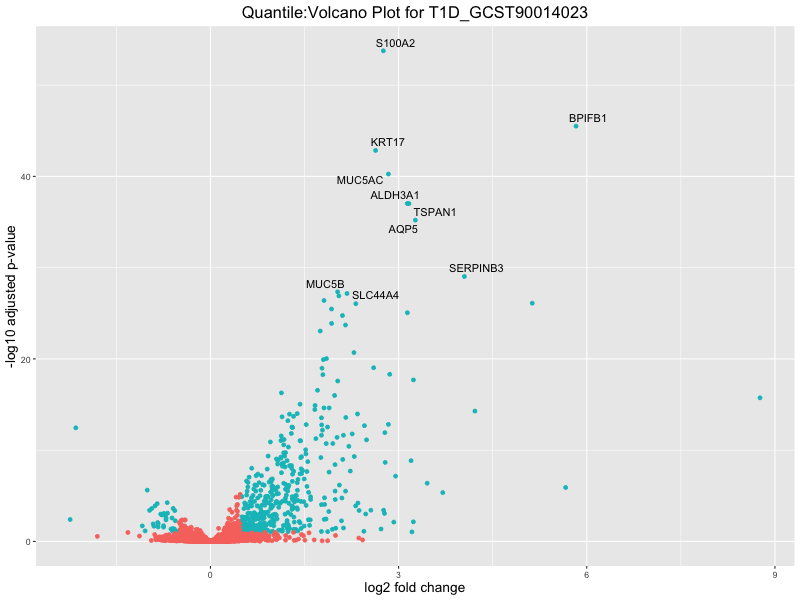
ggtitle(paste("Quantile:Volcano Plot for", trait)) +
xlab("log2 fold change") +
ylab("-log10 adjusted p-value") +
theme(legend.position = "none",
plot.title = element_text(size = rel(1.5), hjust = 0.5),
axis.title = element_text(size = rel(1.25)))
# Save the volcano plot
png(paste0("volcano_plot_quantile_", trait, ".png"), width = 800, height = 600)
print(volcano_plot)
dev.off()
}Model 2: expression ~ PRS + sex + expression PCs + 2 genotype PCs
Continuous PRS:
# Load the gene expression data
gene_expr_file <- "data/gene_reads_2017-06-05_v8_whole_blood.gct"
raw_count_df <- fread(gene_expr_file, header = TRUE, sep = "\t", drop = "id")
# load protein_coding list
protein_coding <- fread("data/protein-coding_gene.txt",
sep = "\t")
protein_coding <- protein_coding[, c("symbol", "ensembl_gene_id")]
# keep only protein-coding genes
raw_count_df <- raw_count_df[raw_count_df$Description %in% protein_coding$symbol, ]
id <- raw_count_df$Name
raw_count <- raw_count_df[, -c(1:2)]
# modify GTEx sample names matching names used in PRS data
colnames(raw_count) <- sub("^(GTEX-[^-.]+).*", "\\1", colnames(raw_count))
matching_samples <- intersect(rownames(metadata), colnames(raw_count))
final_count <- raw_count[ , ..matching_samples]
# prefilter: keep only rows that have a count of at least 10
keep_genes <- rowSums(final_count >= 10) > 0
final_count <- final_count[keep_genes, ]
id <- id[keep_genes]
dim(final_count)[1] 16893 670# obtain genotype PCs
geno_pc <- read.table("data/pca.eigenvec")
names(geno_pc) = c("FID","IID",paste0("geno_PC", c(1:(ncol(geno_pc)-2))))
eigenval <- scan("data/pca.eigenval")
pve <- data.frame(PC = 1:20, pve = eigenval/sum(eigenval)*100)
ggplot(geno_pc) + geom_point(aes(x = geno_PC1, y = geno_PC2)) +
labs(x = paste0("PC1 (", signif(pve$pve[1], 3), "%)"),
y = paste0("PC2 (", signif(pve$pve[2], 3), "%)")) + theme_classic()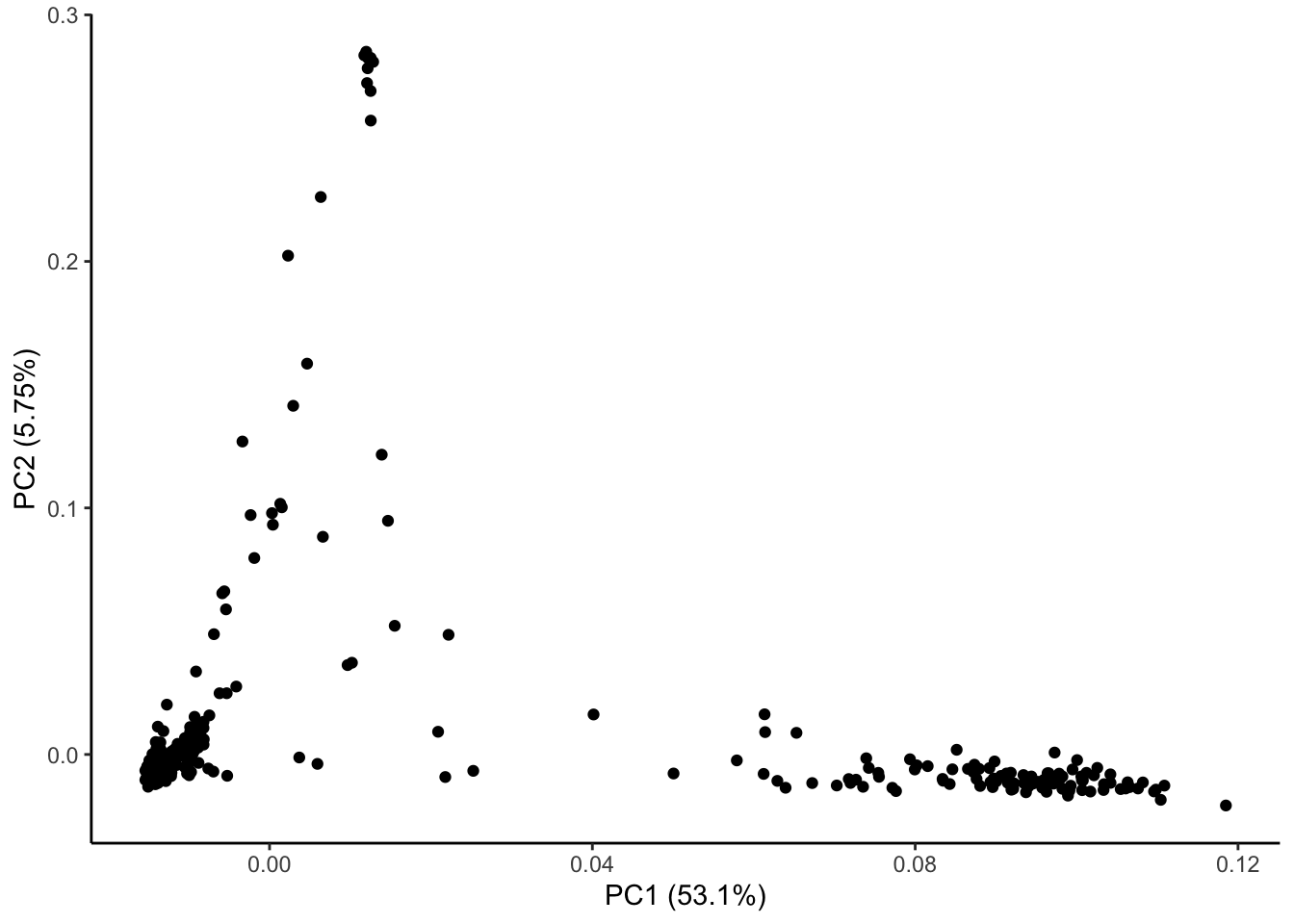
| Version | Author | Date |
|---|---|---|
| 51ffd48 | ElisaChen | 2025-05-20 |
ggplot(geno_pc) + geom_point(aes(x = geno_PC1, y = geno_PC3)) +
labs(x = paste0("PC1 (", signif(pve$pve[1], 3), "%)"),
y = paste0("PC3 (", signif(pve$pve[3], 3), "%)")) + theme_classic()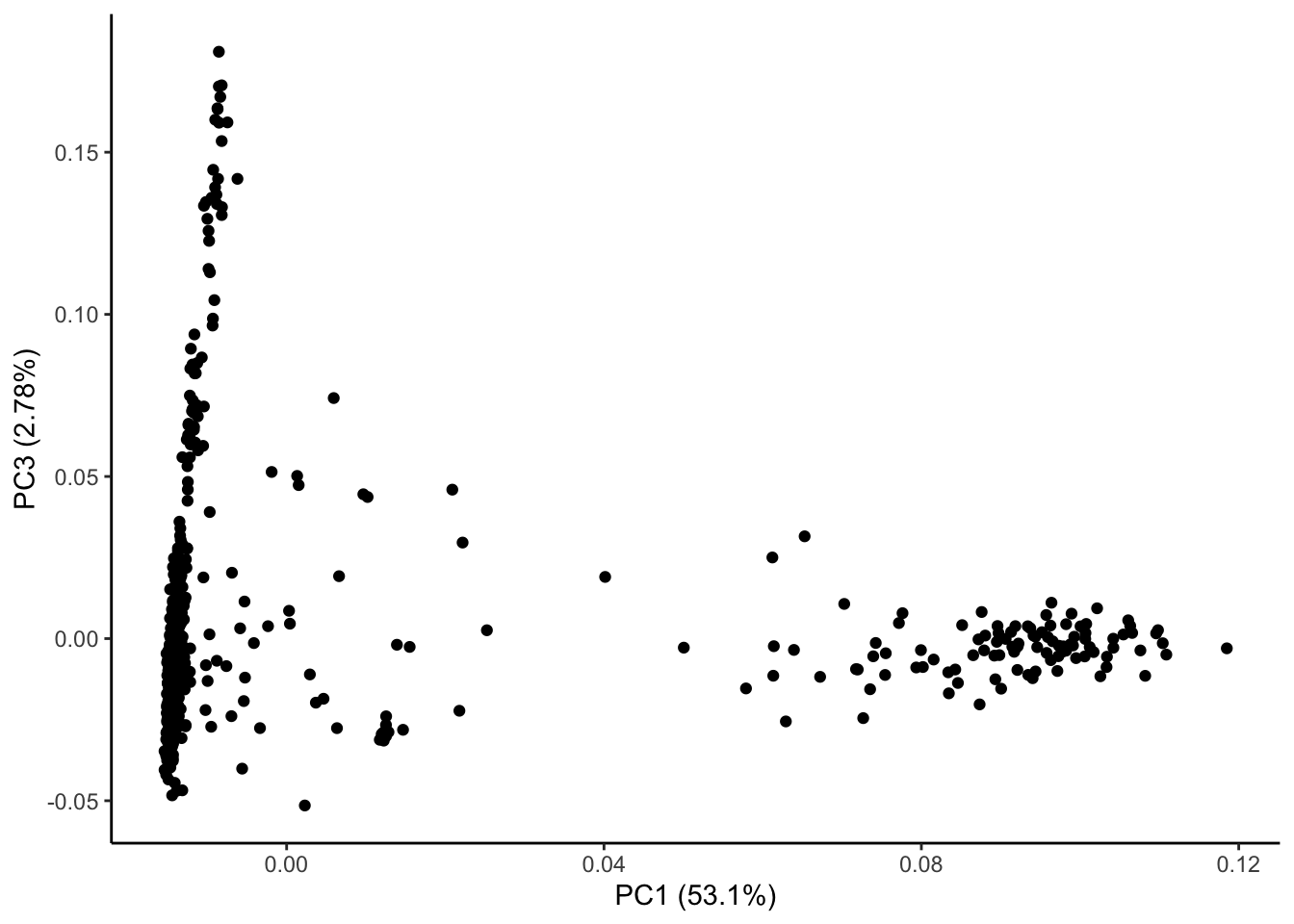
| Version | Author | Date |
|---|---|---|
| 51ffd48 | ElisaChen | 2025-05-20 |
ggplot(pve, aes(PC, pve)) + geom_point() + geom_line() +
labs(x = "Genotype PC", y = "Percentage variance explained") + theme_classic()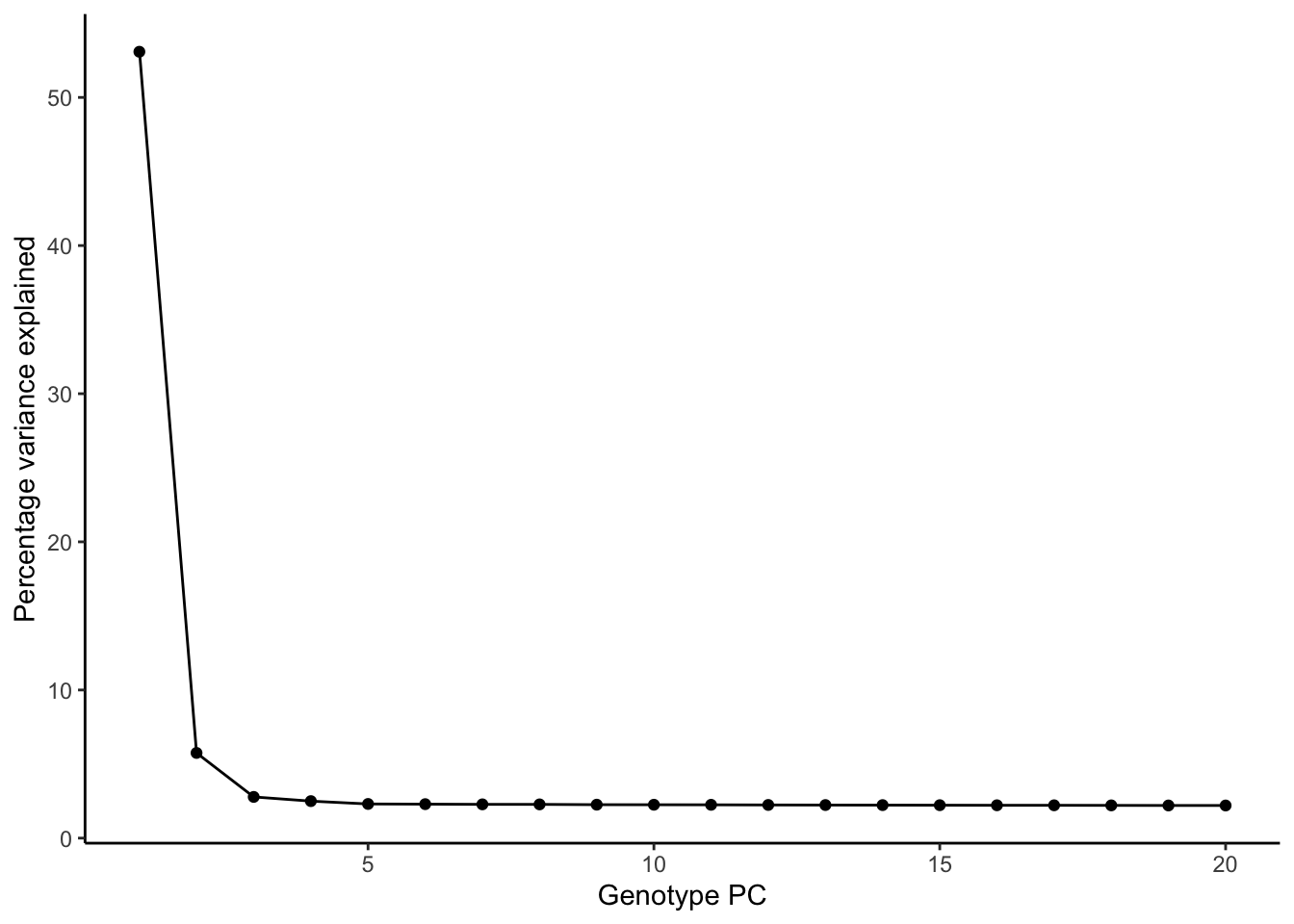
| Version | Author | Date |
|---|---|---|
| 51ffd48 | ElisaChen | 2025-05-20 |
metadata <- cbind(metadata, geno_pc[,3:ncol(geno_pc)])
# Loop through each trait and run DESeq2
for (trait in colnames(traits)) {
# Standardize PRS for the current trait
prs_trait <- traits[,trait]
prs_trait <- scale(prs_trait) # Standardize PRS to mean = 0, sd = 1
# Add the standardized PRS to the metadata for continuous trait
metadata[,trait] <- prs_trait
# Create the DESeqDataSet for the current trait
dds <- DESeqDataSetFromMatrix(
countData = as.matrix(final_count), # Raw counts
colData = metadata[, c(1:6, 44:45, which(colnames(metadata) == trait))],
design = as.formula(paste("~ PC1 + PC2 + PC3 + PC4 + PC5 + sex + geno_PC1 + geno_PC2 +", trait))
)
rownames(dds) <- id
# Run DESeq2 analysis
dds <- DESeq(dds, parallel = TRUE, BPPARAM = MulticoreParam(4))
# Get the results for the current trait
res <- results(dds)
# Save the results to a file
write.csv(res, paste0("differential_expression_", trait, "_results_M2.csv"))
# print a summary of the results
print(paste("Results for trait:", trait))
print(summary(res))
# plot the MA-plot for the current trait
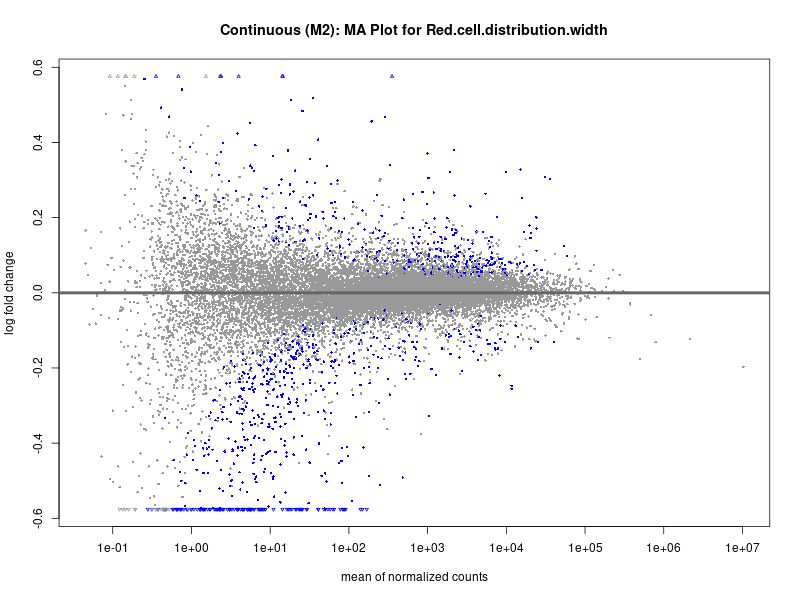
png(paste0("ma_plot_", trait, "_M2.png"), width = 800, height = 600)
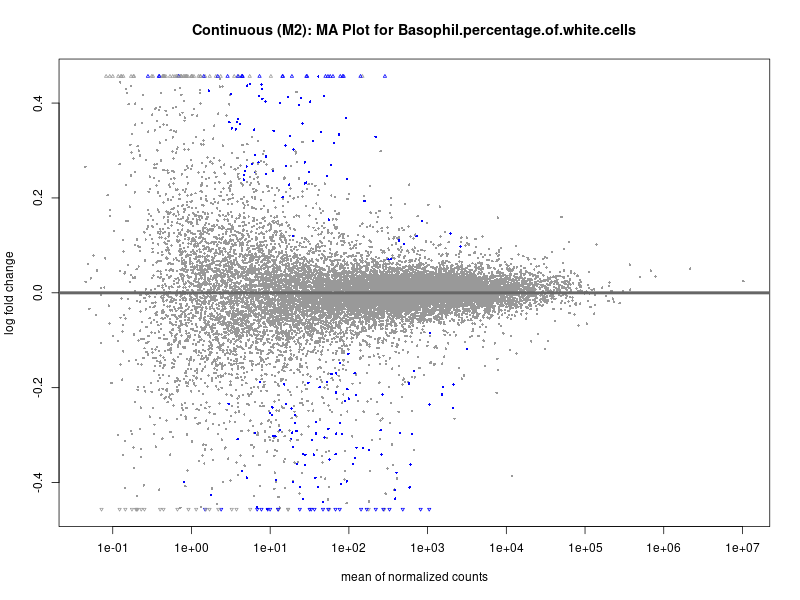
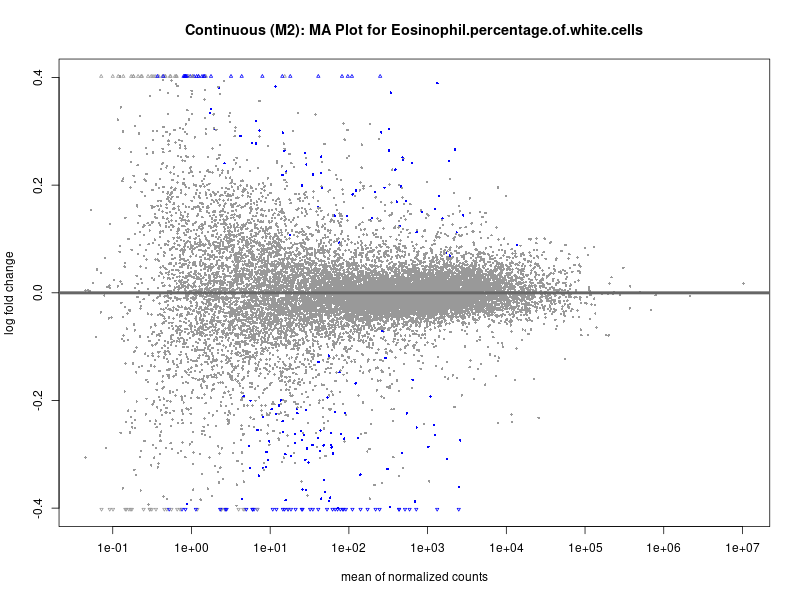
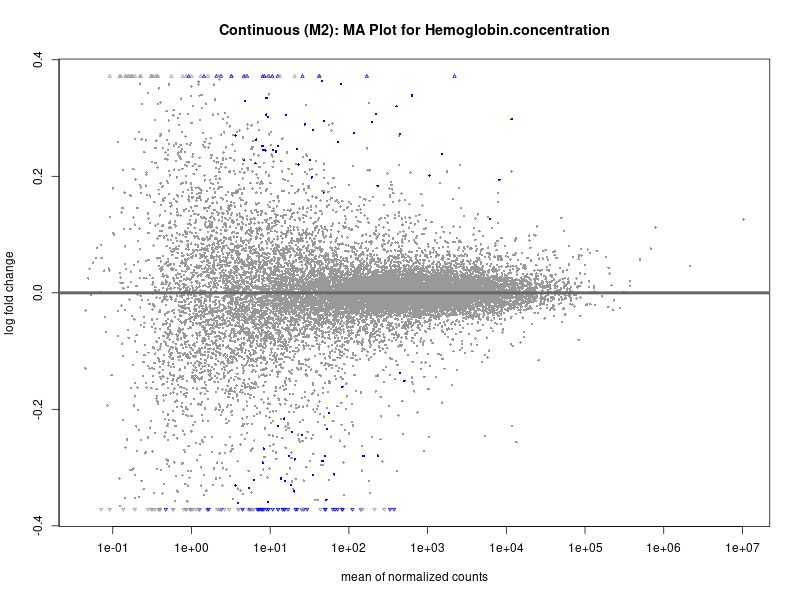
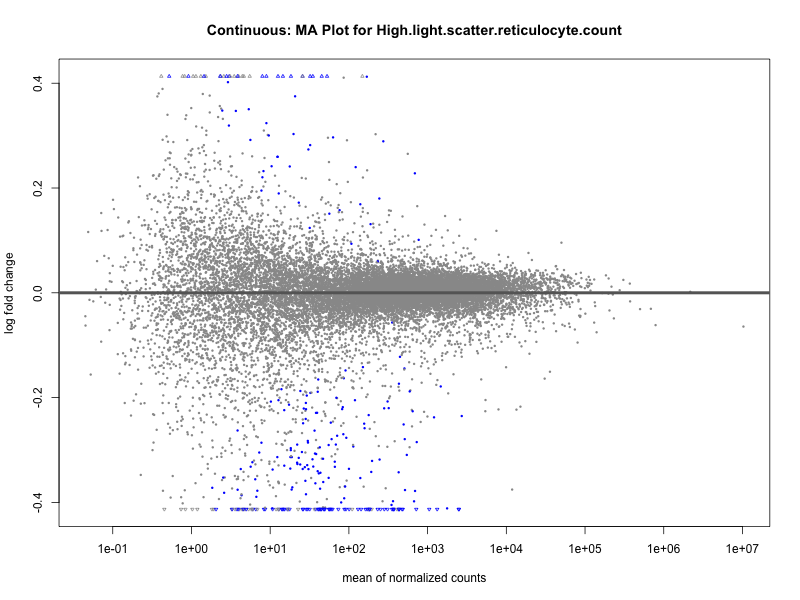
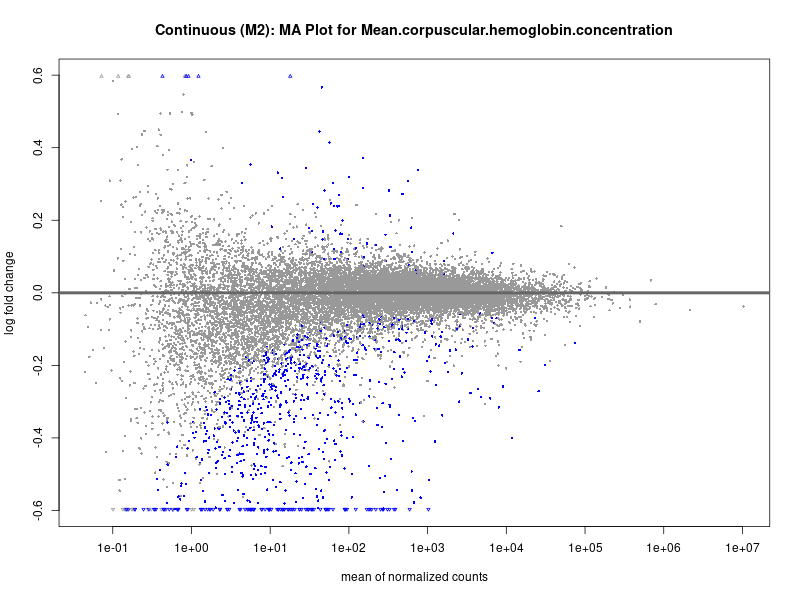
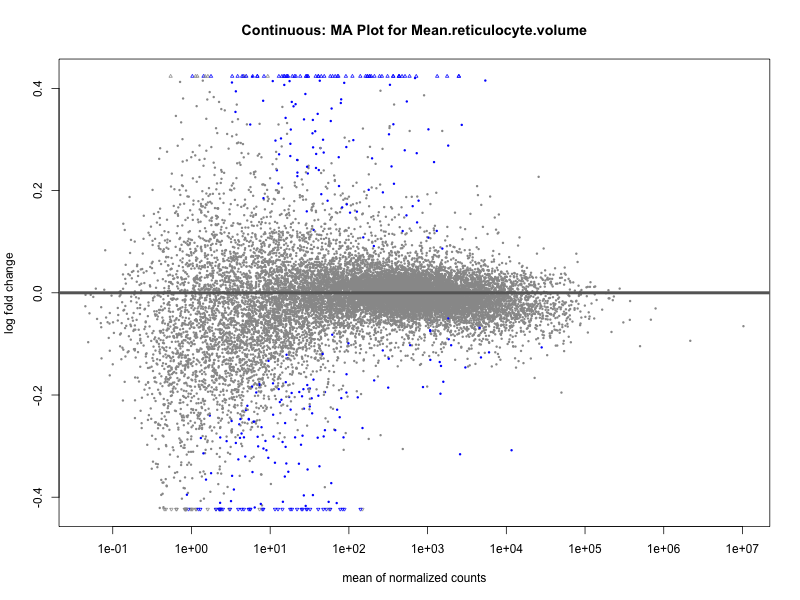
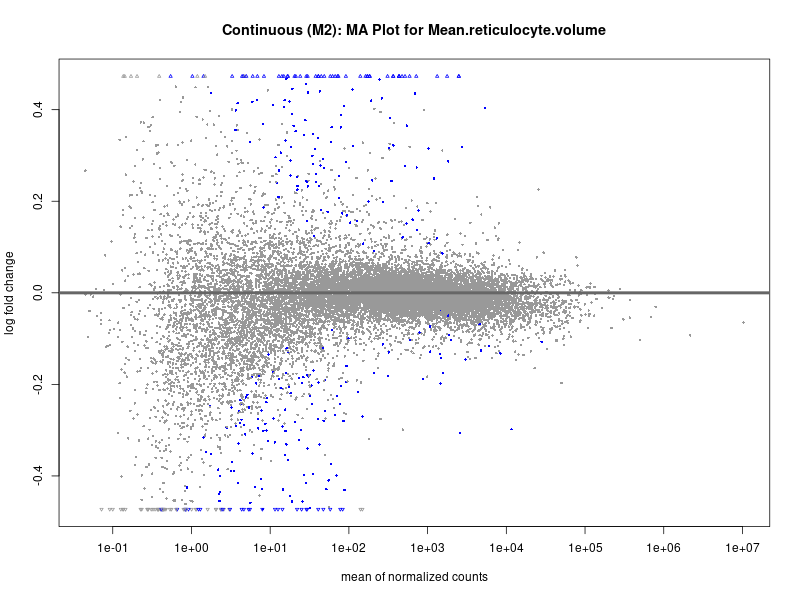
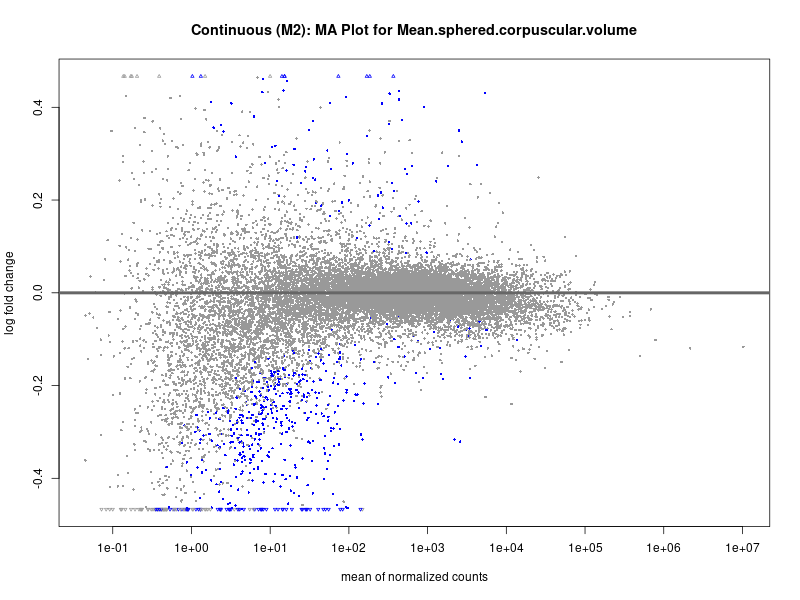
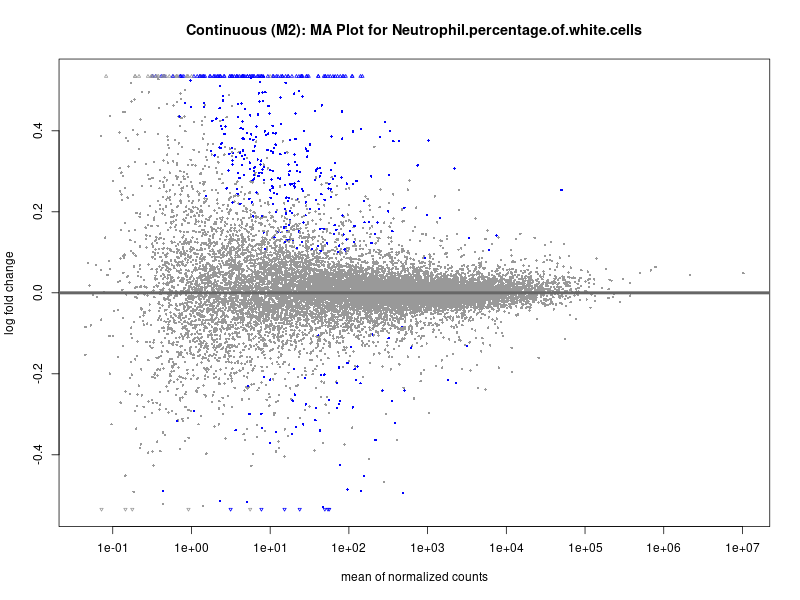
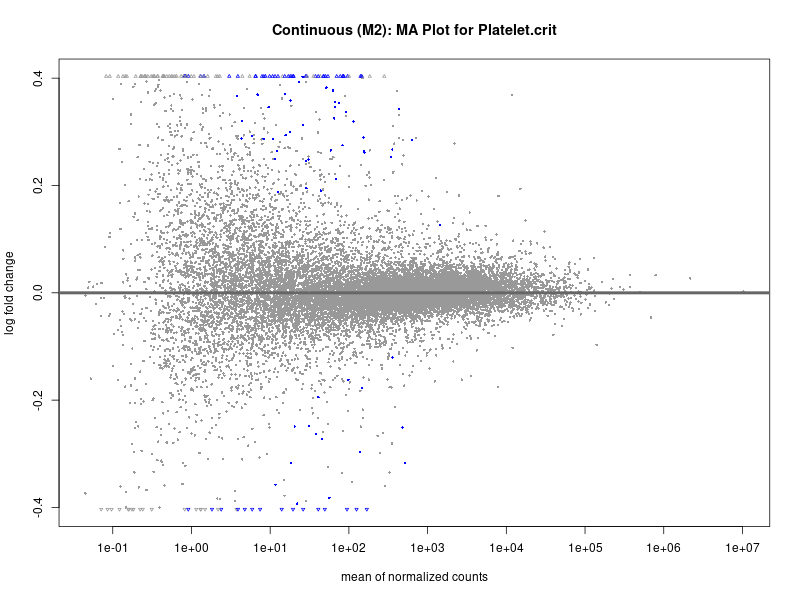
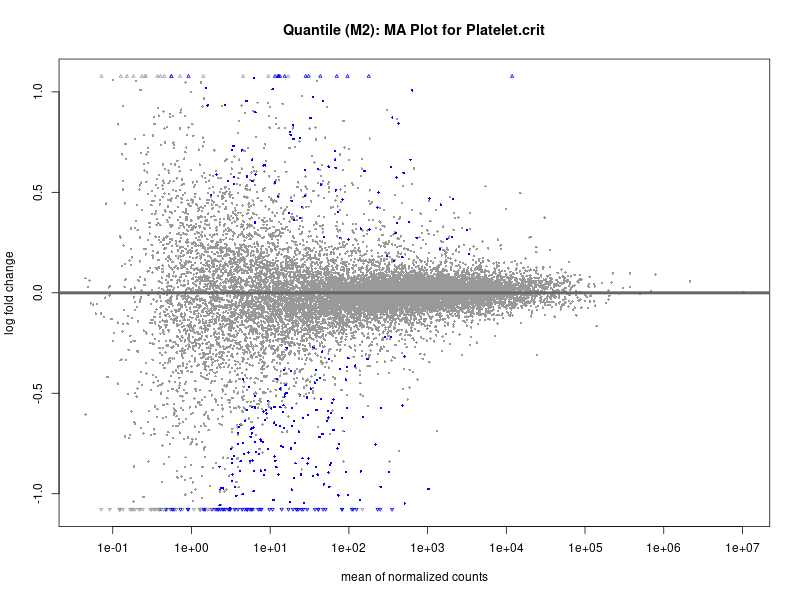
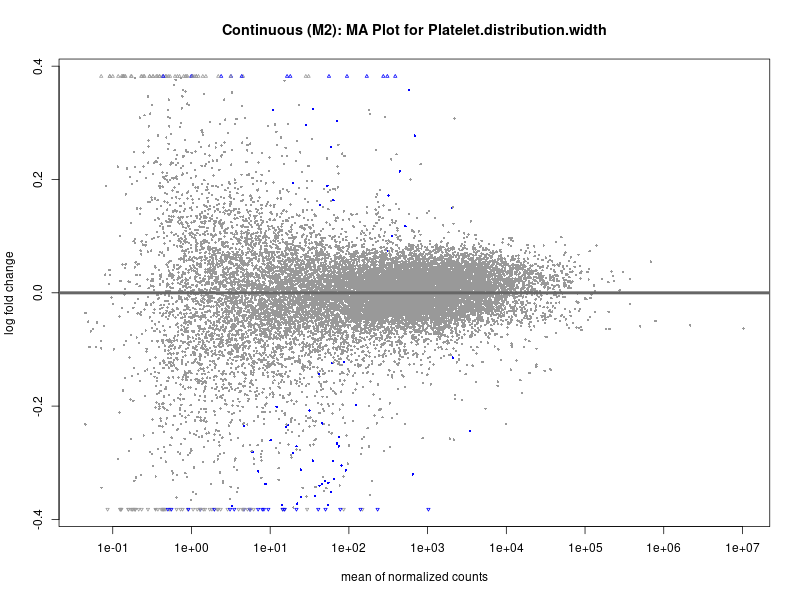
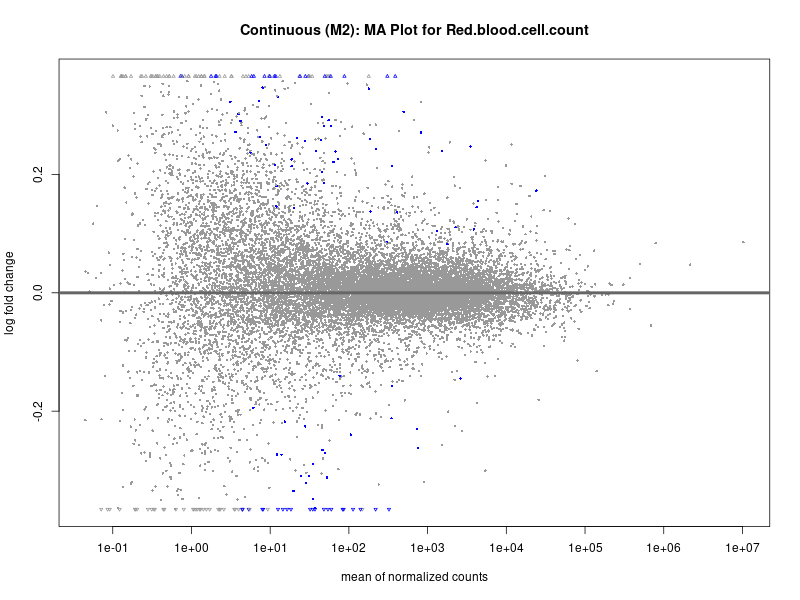
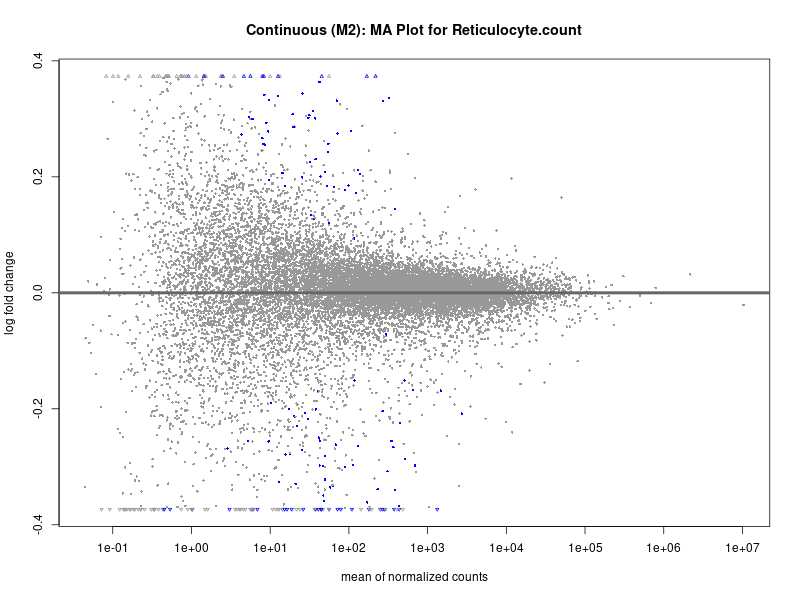
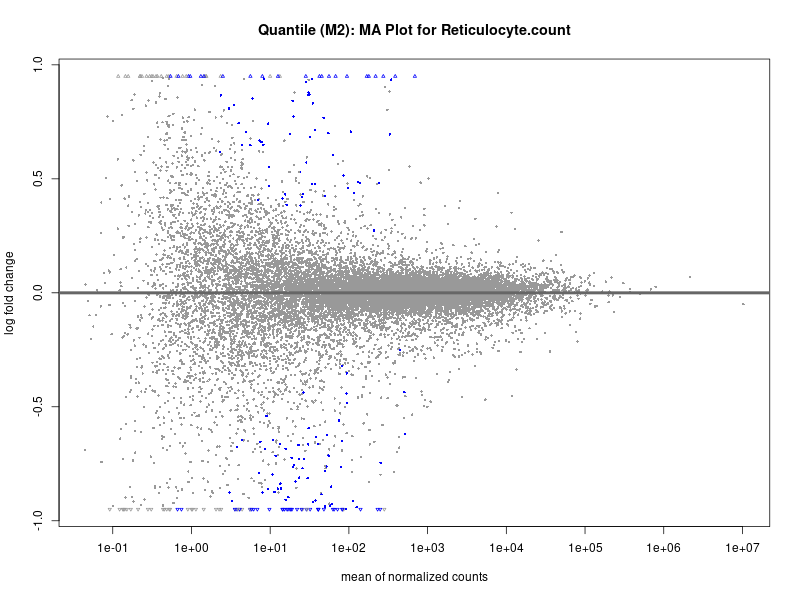
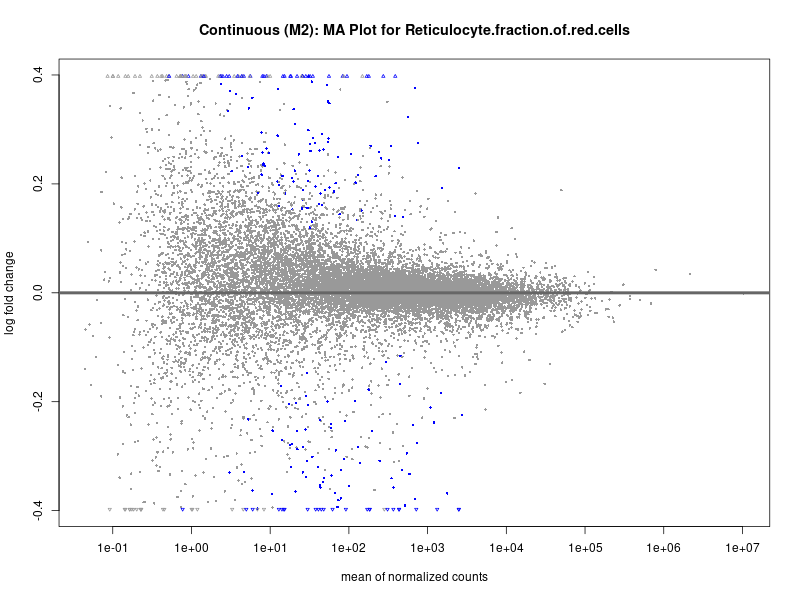
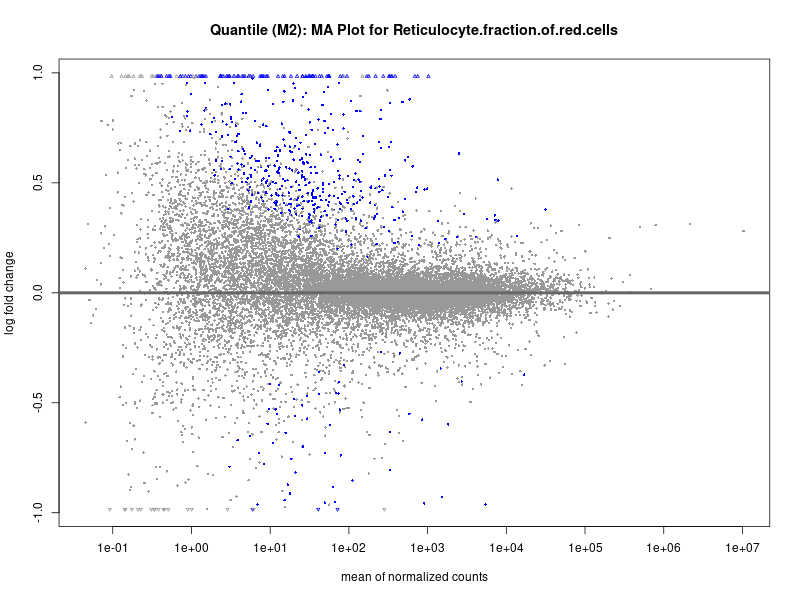
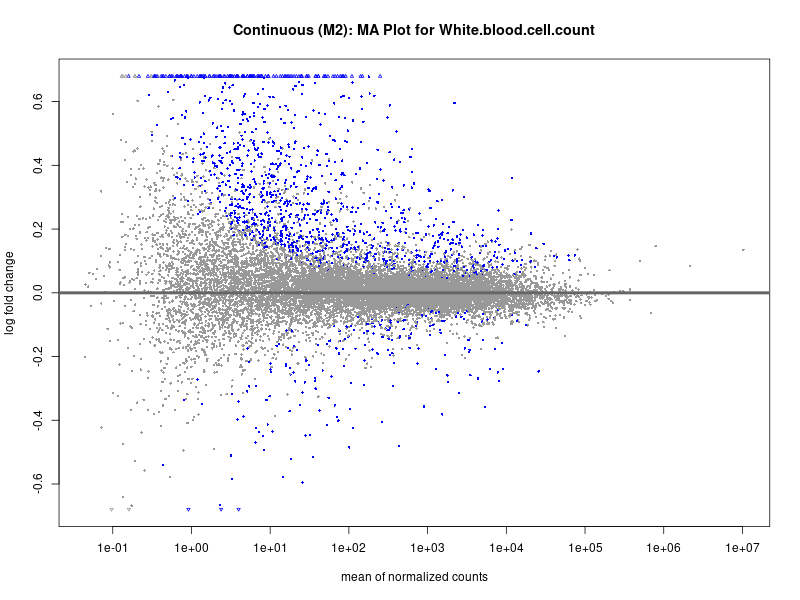
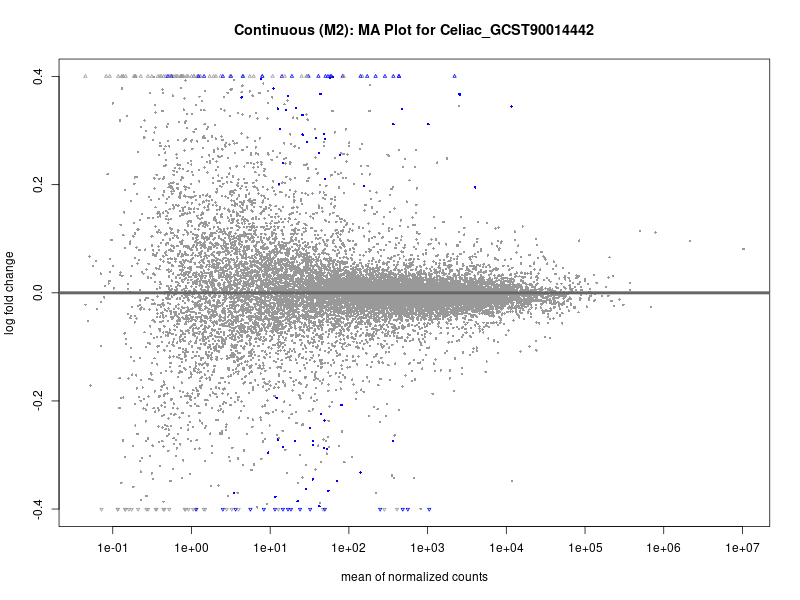
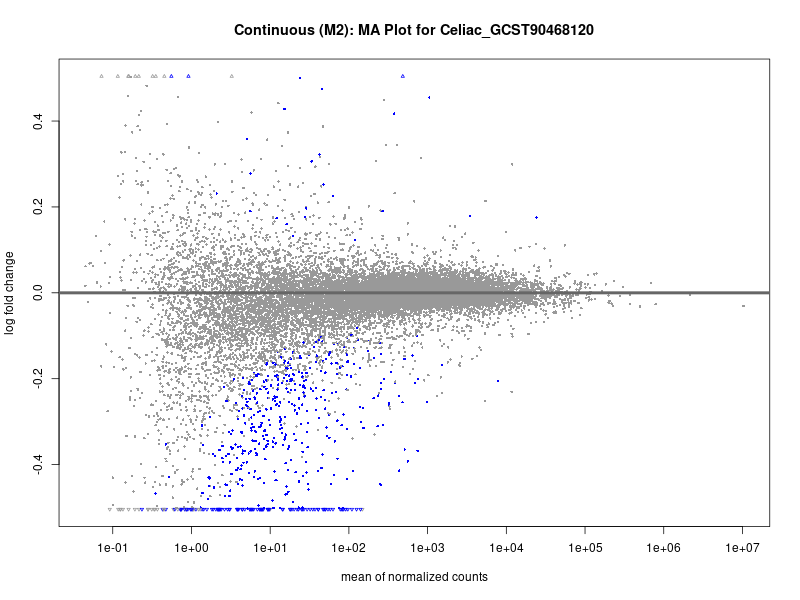
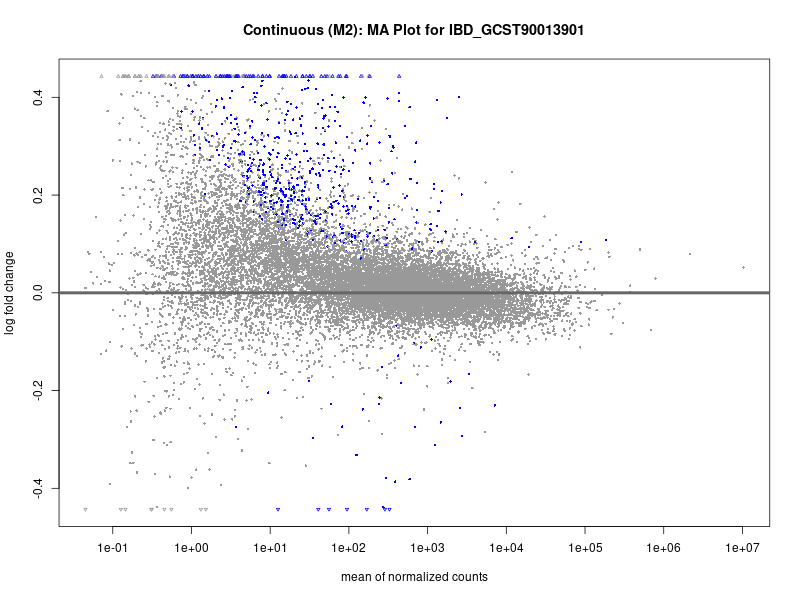
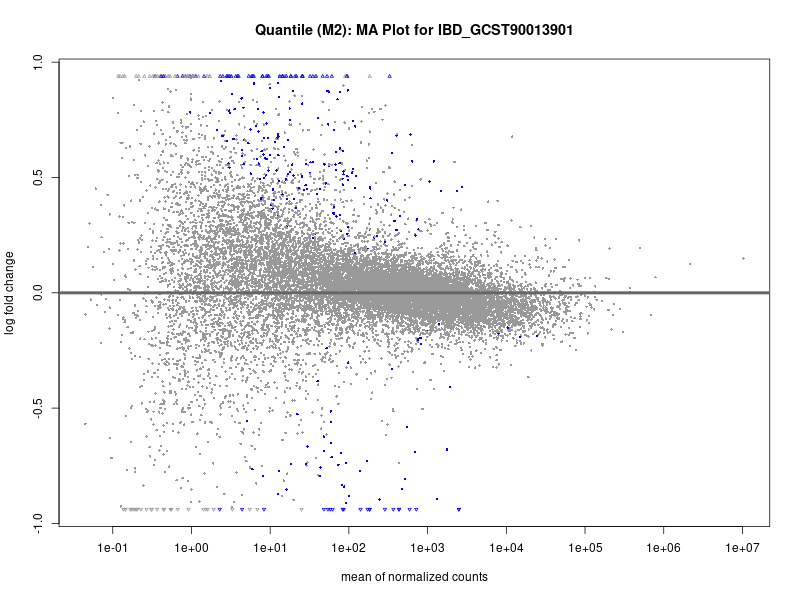
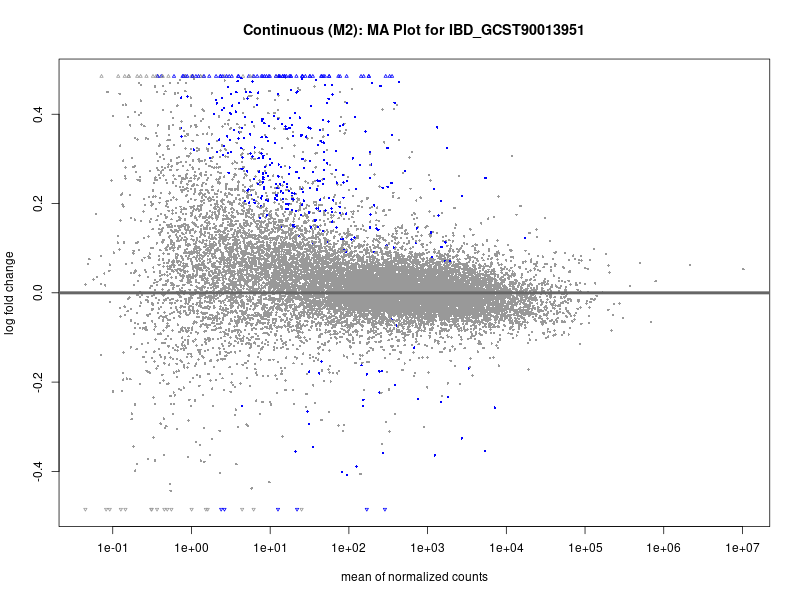
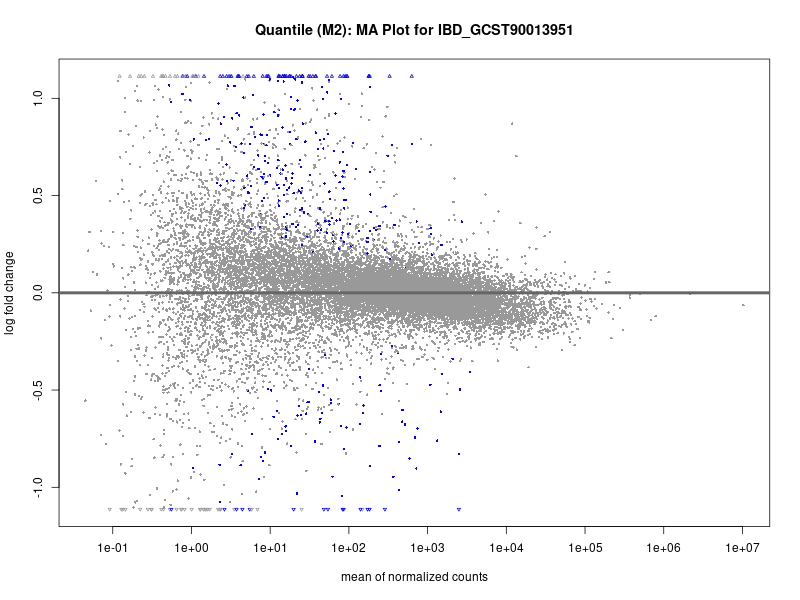
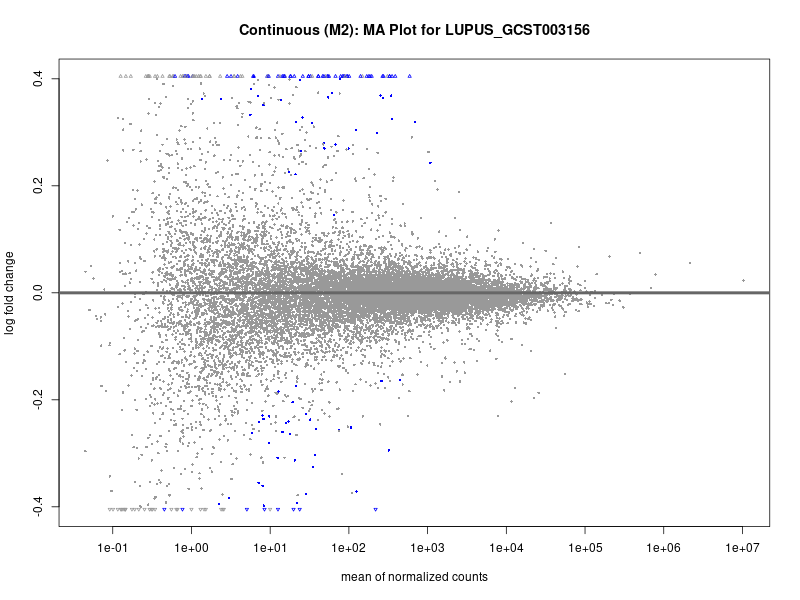
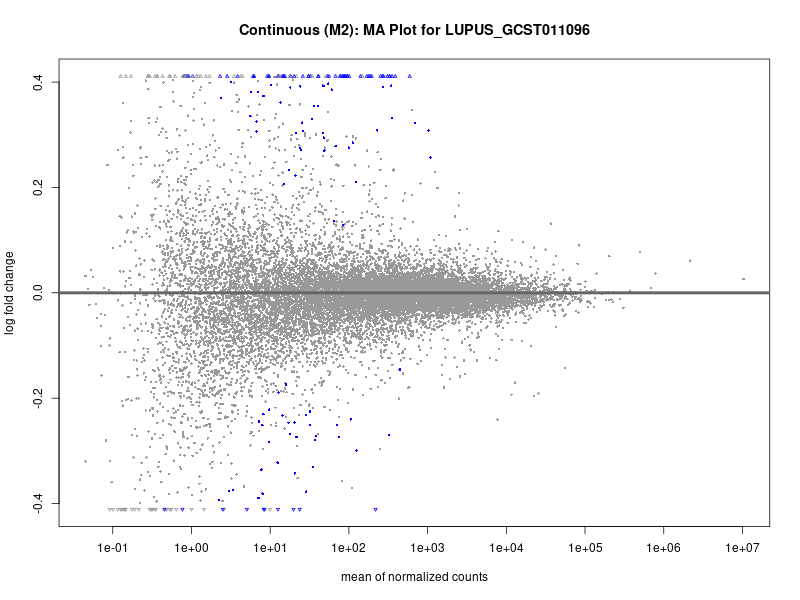
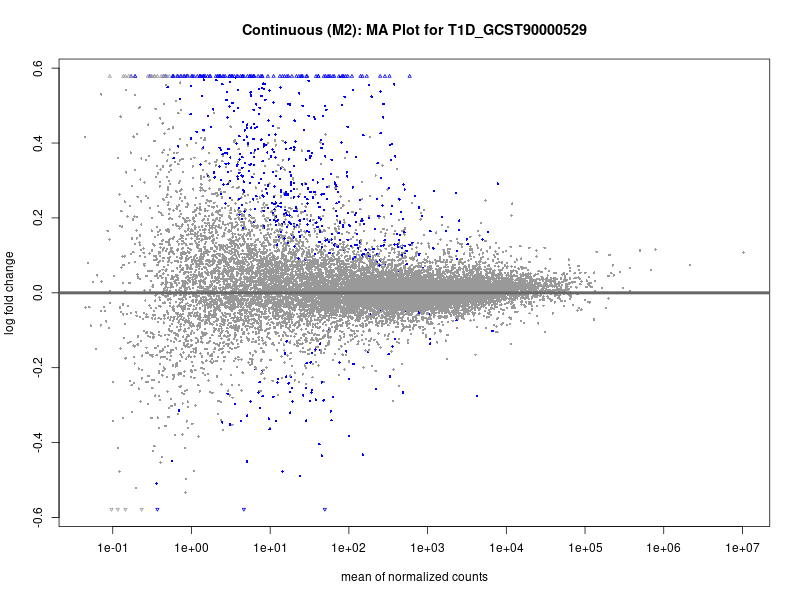
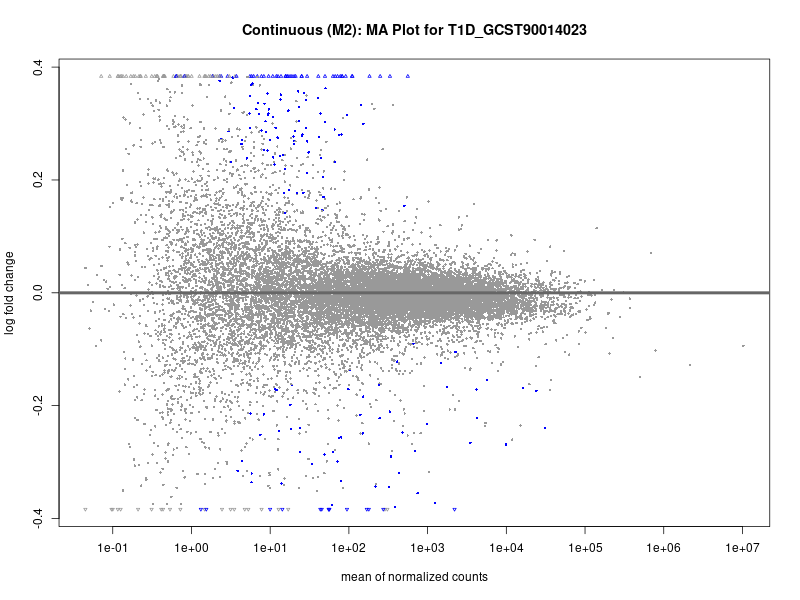
plotMA(res, main = paste("Continuous (M2): MA Plot for", trait))
dev.off()
# volcano plot
res_tableOE <- as.data.frame(res)
res_tableOE$gene_name <- raw_count_df$Description[keep_genes]
res_tableOE <- mutate(res_tableOE, threshold_OE = padj < 0.1)
res_tableOE <- res_tableOE %>% arrange(padj) %>% mutate(genelabels = "")
res_tableOE$genelabels[1:10] <- res_tableOE$gene_name[1:10]
volcano_plot <- ggplot(res_tableOE, aes(x = log2FoldChange, y = -log10(padj))) +
geom_point(aes(colour = threshold_OE)) +
geom_text_repel(aes(label = genelabels)) +
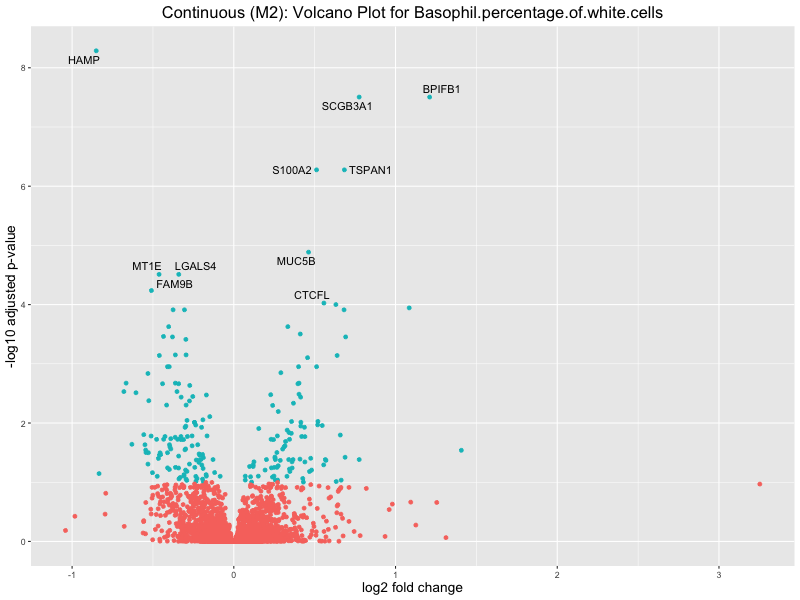
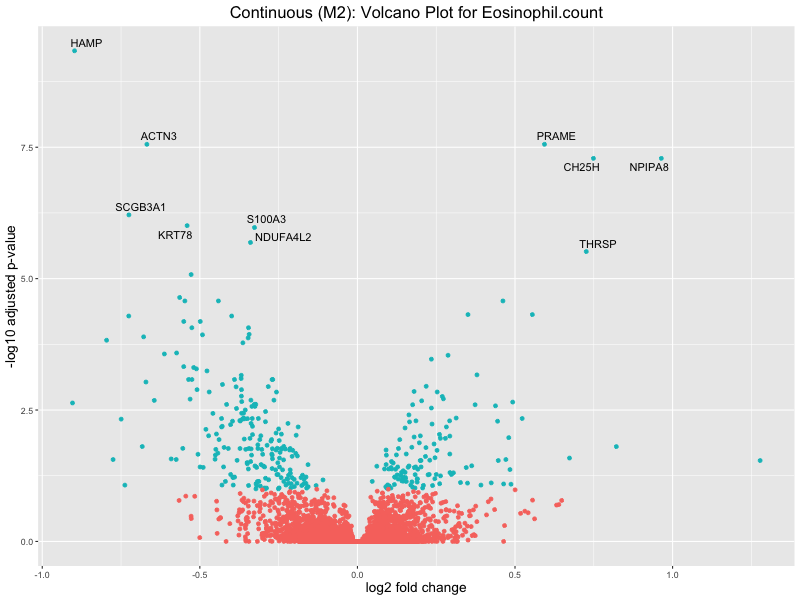
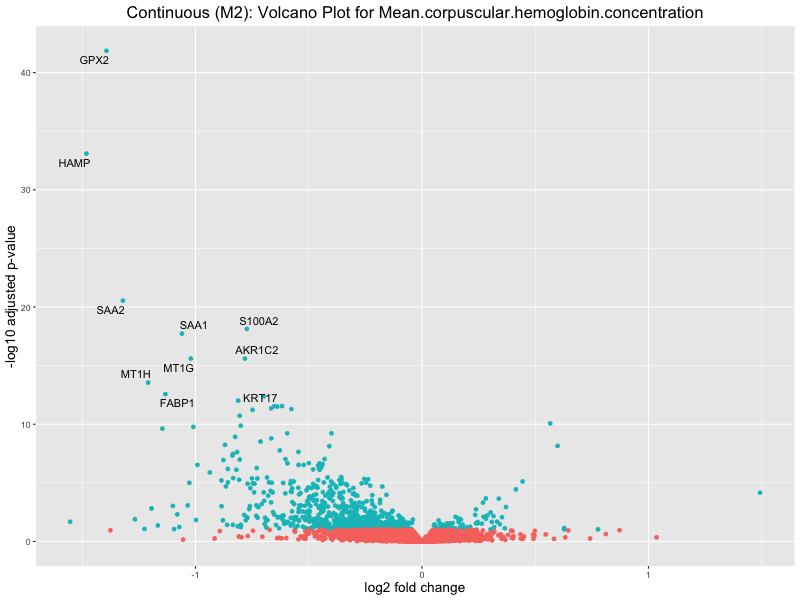
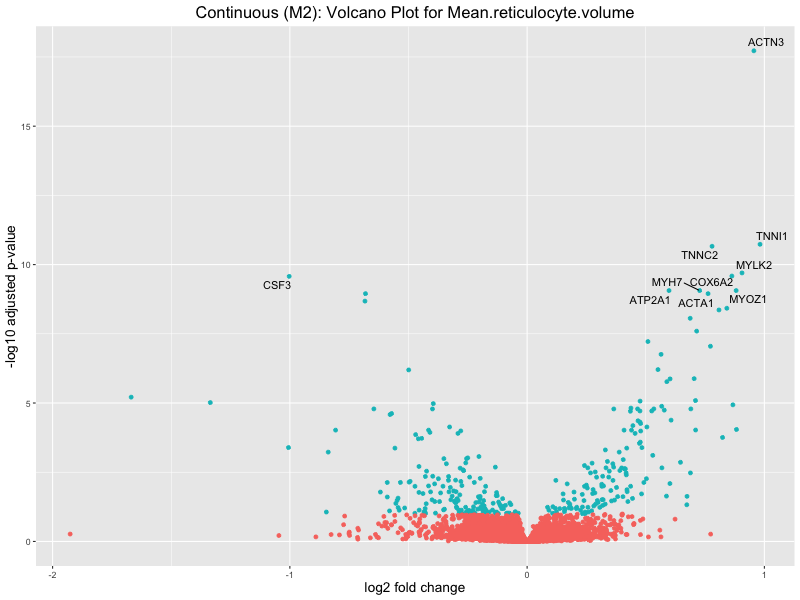
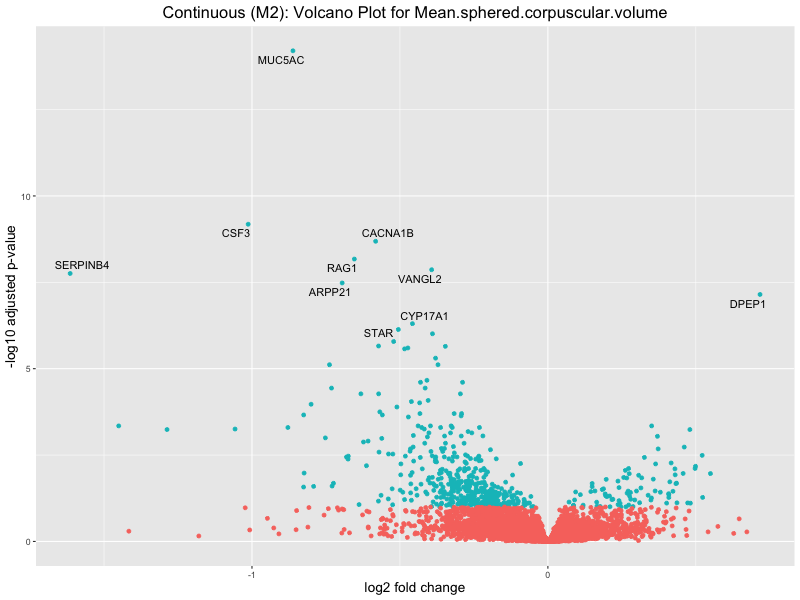
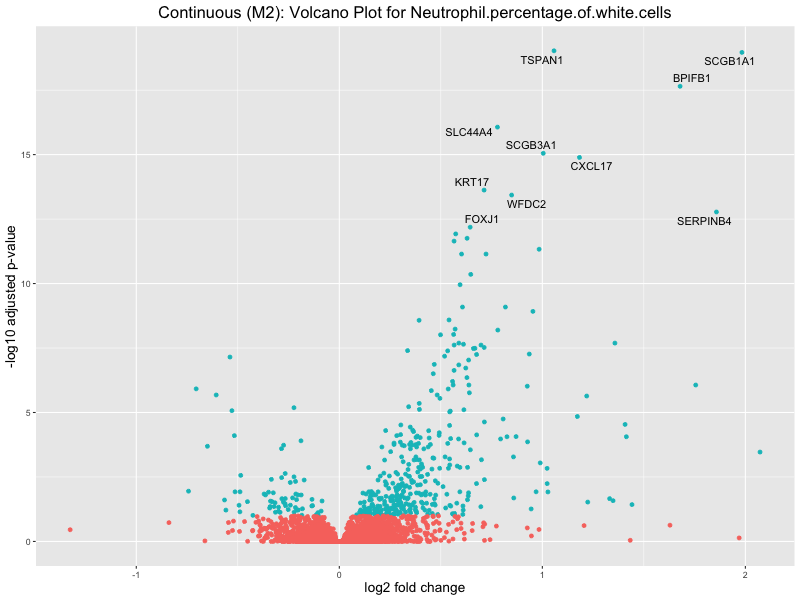
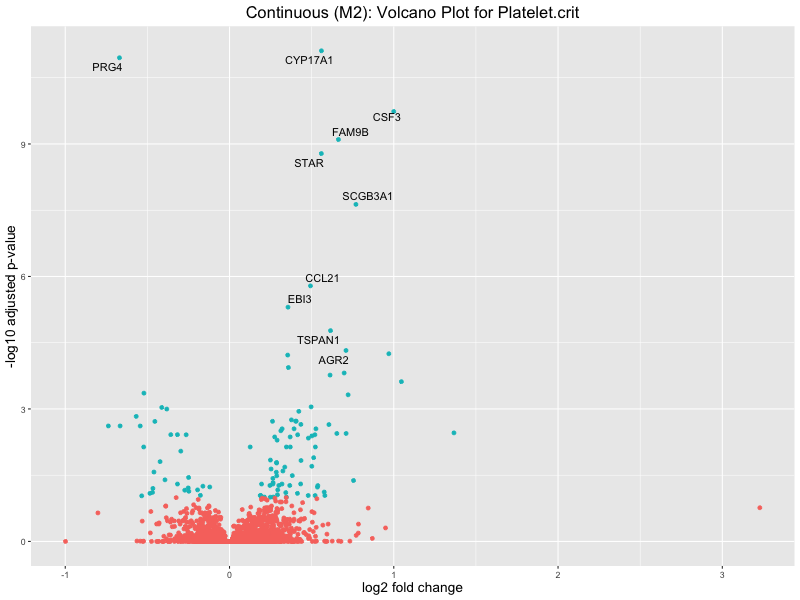
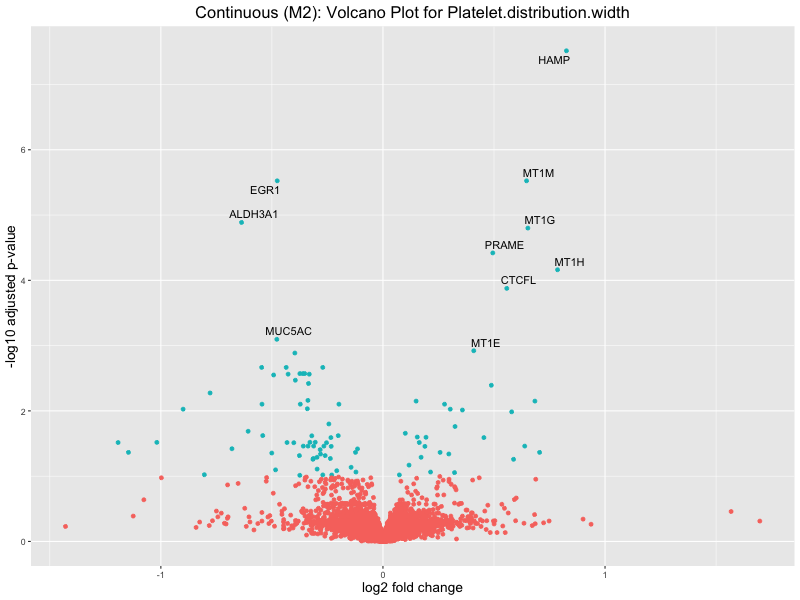
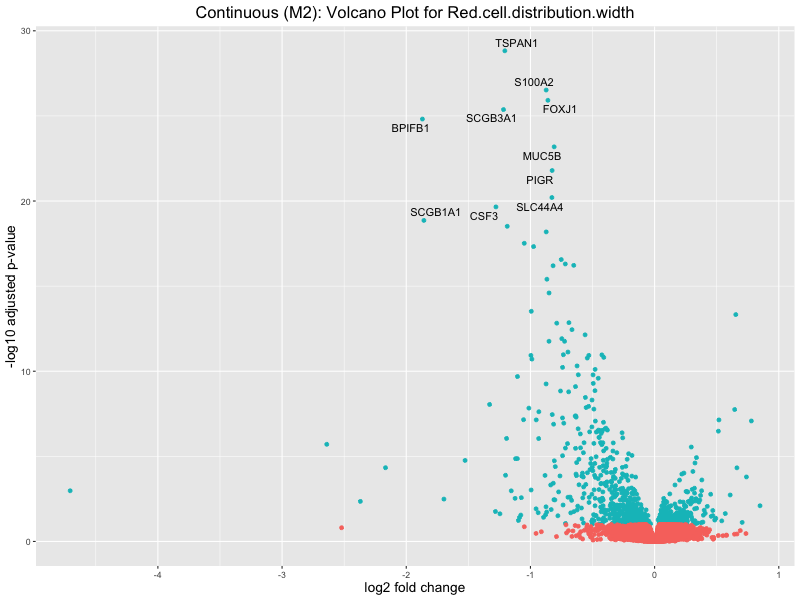
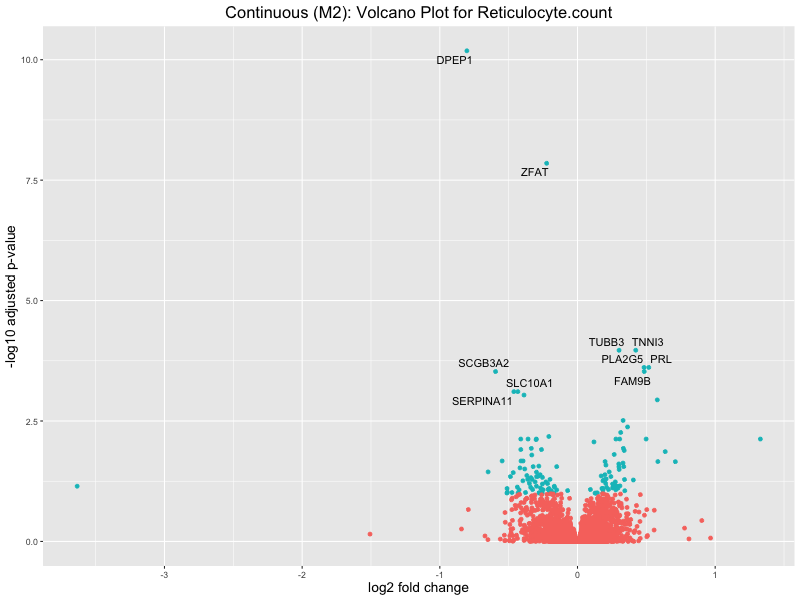
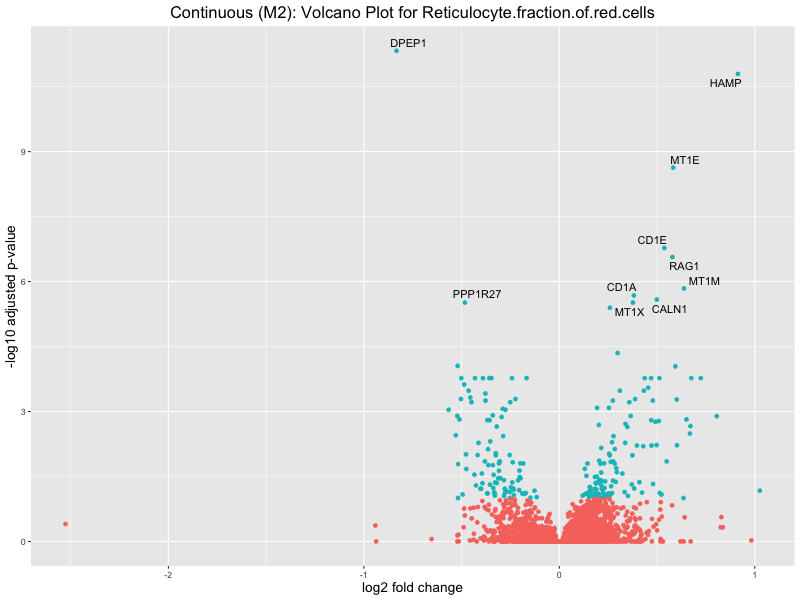
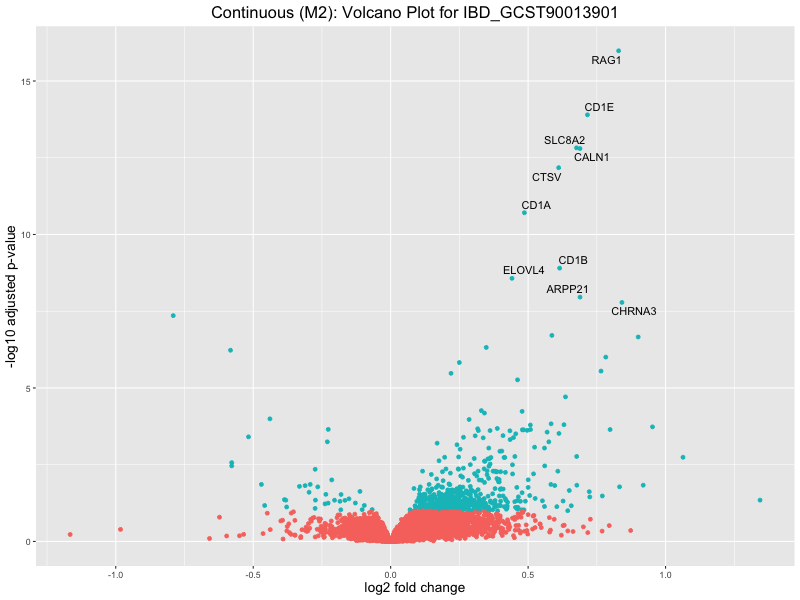
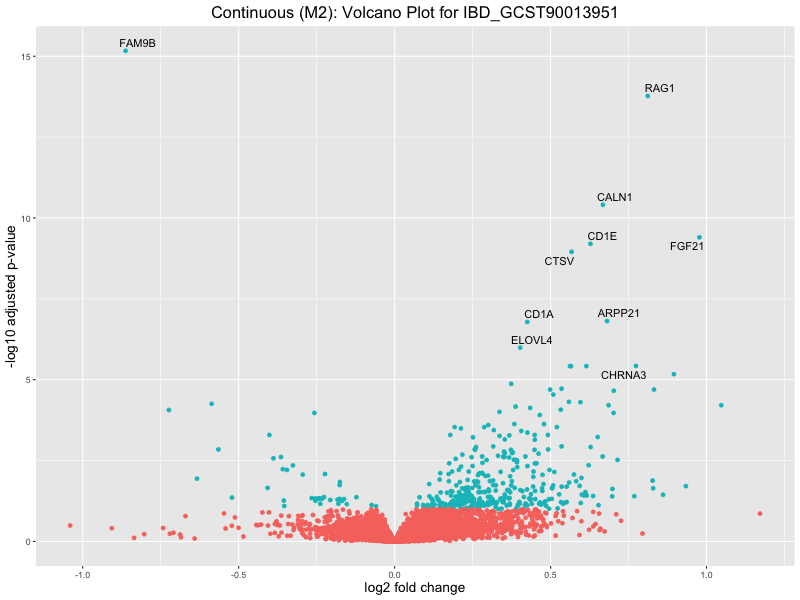
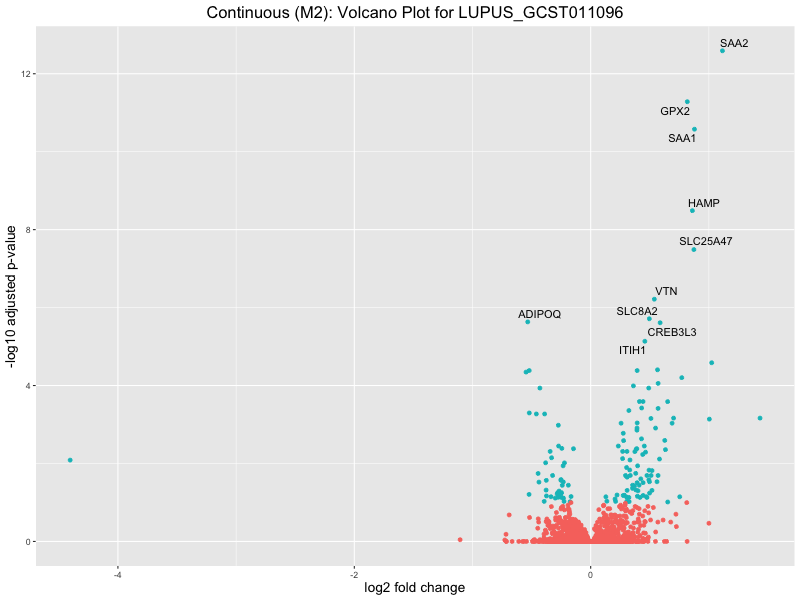
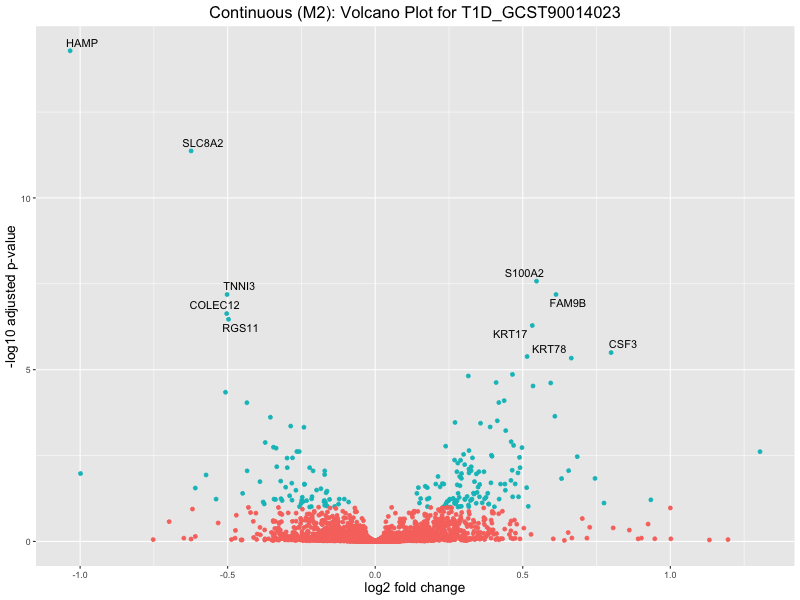
ggtitle(paste("Continuous (M2): Volcano Plot for", trait)) +
xlab("log2 fold change") +
ylab("-log10 adjusted p-value") +
theme(legend.position = "none",
plot.title = element_text(size = rel(1.5), hjust = 0.5),
axis.title = element_text(size = rel(1.25)))
# Save the volcano plot
png(paste0("volcano_plot_", trait, "_M2.png"), width = 800, height = 600)
print(volcano_plot)
dev.off()
}Quantile PRS:
metadata_file <- "analysis/metadata_quantile.txt"
metadata <- read.csv(metadata_file, header = T, sep = "\t", stringsAsFactors = T)
metadata$sex <- as.factor(metadata$sex)
traits <- metadata[, 7:43]
metadata <- cbind(metadata, geno_pc[,3:ncol(geno_pc)])
# Loop through each trait and run DESeq2
for (trait in colnames(traits)) {
# Create the DESeqDataSet for the current trait
dds <- DESeqDataSetFromMatrix(
countData = as.matrix(final_count), # Raw counts
colData = metadata[, c(1:6, which(colnames(metadata) == trait))],
design = as.formula(paste("~ PC1 + PC2 + PC3 + PC4 + PC5 + sex + geno_PC1 + geno_PC2 +", trait))
)
rownames(dds) <- id
# Run DESeq2 analysis
dds <- DESeq(dds, parallel = TRUE, BPPARAM = MulticoreParam(4))
# Get the results for the current trait
res <- results(dds)
# Save the results to a file
write.csv(res, paste0("differential_expression_", trait, "_quantile_results_M2.csv"))
# print a summary of the results
print(paste("Results for trait:", trait))
print(summary(res))
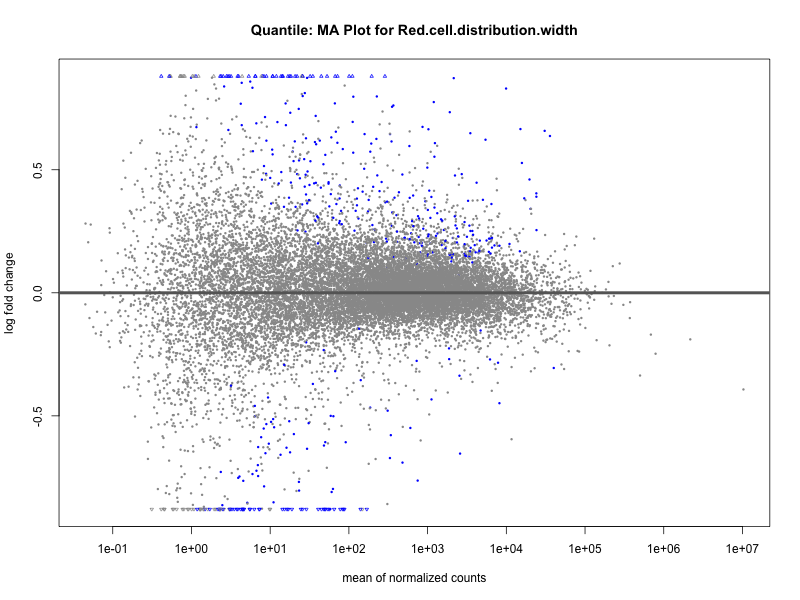
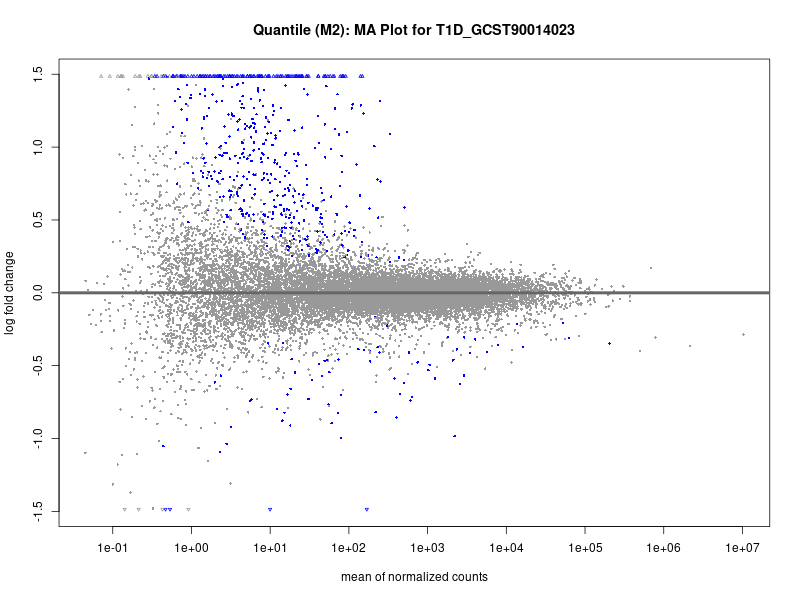
# plot the MA-plot for the current trait
png(paste0("ma_plot_quantile_", trait, "_M2.png"), width = 800, height = 600)
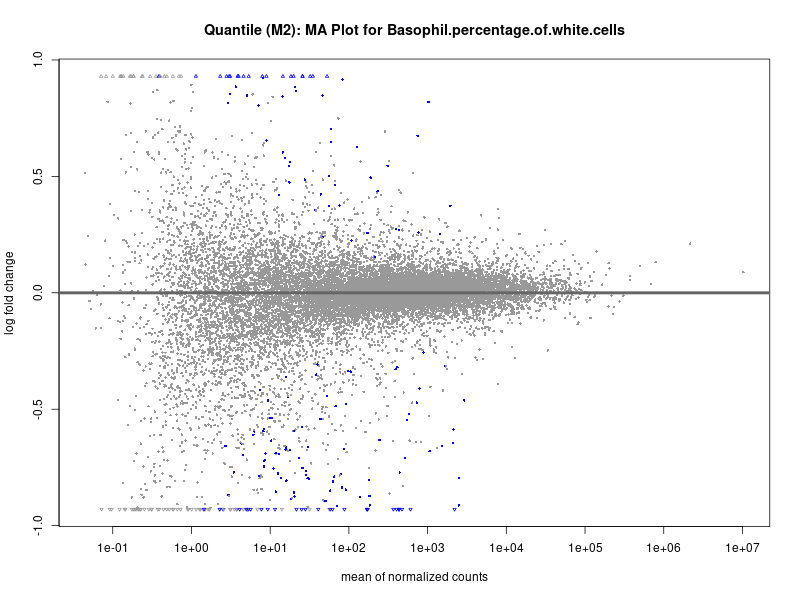
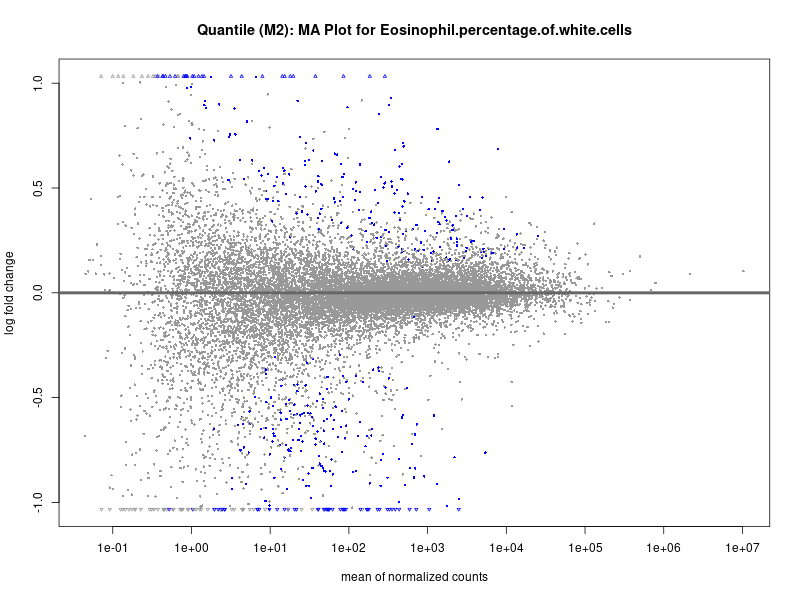
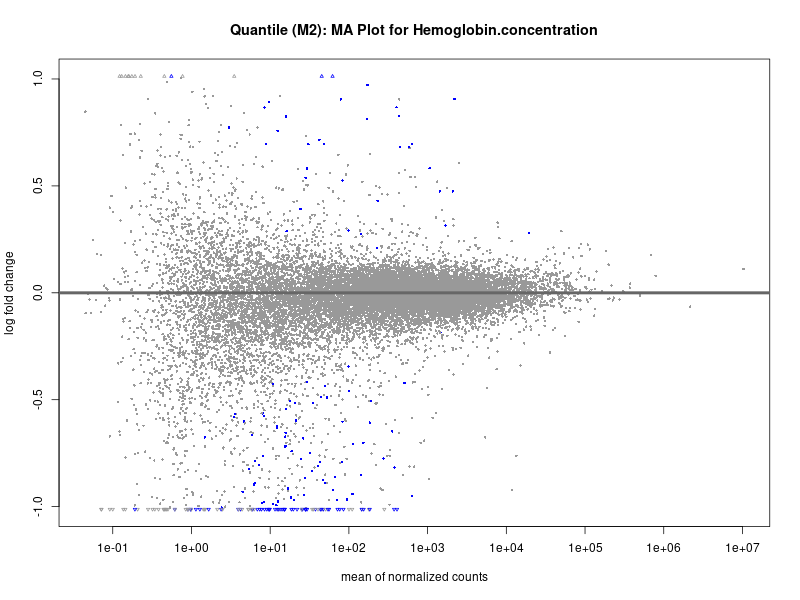
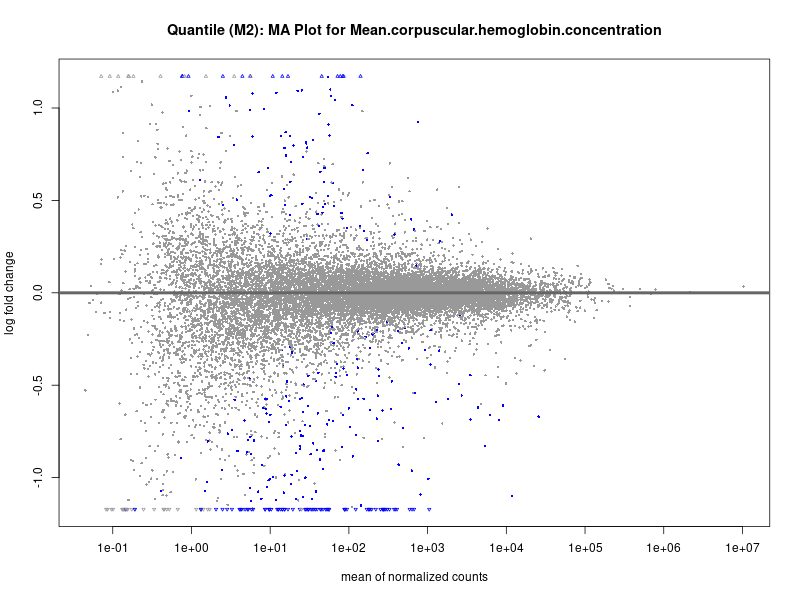
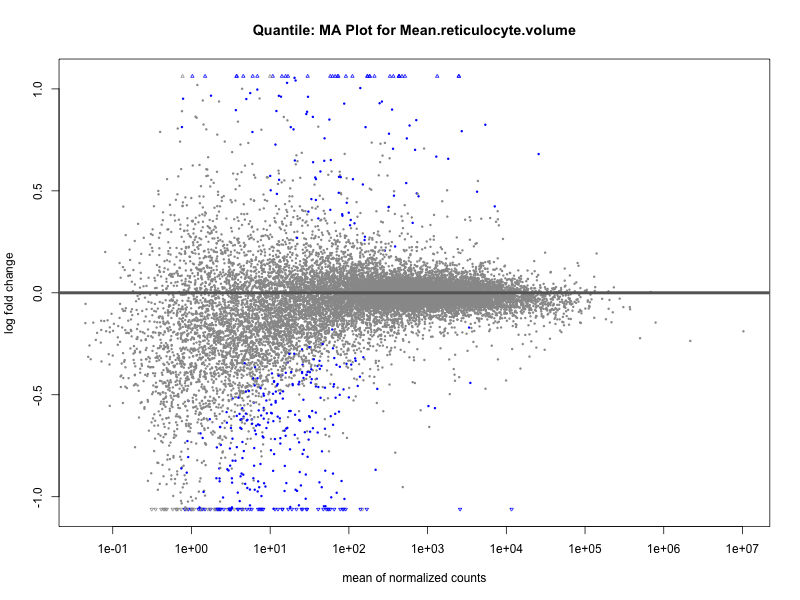
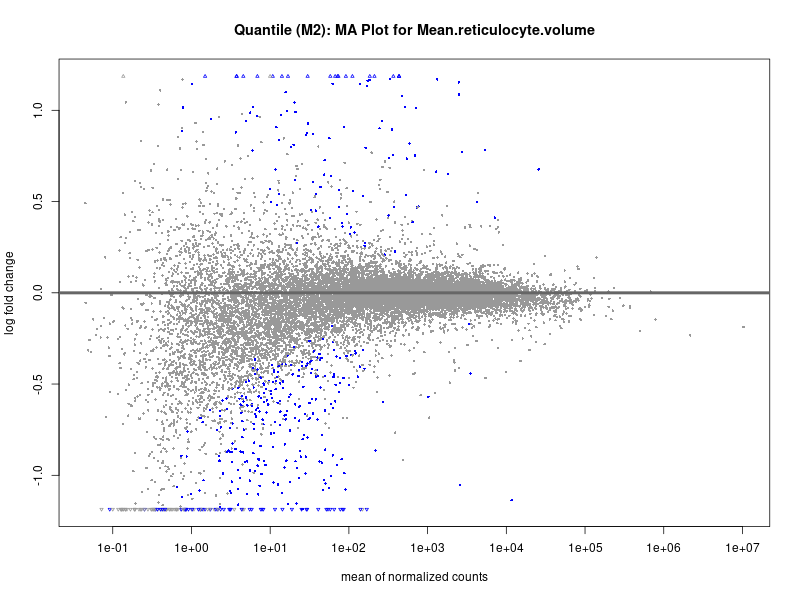
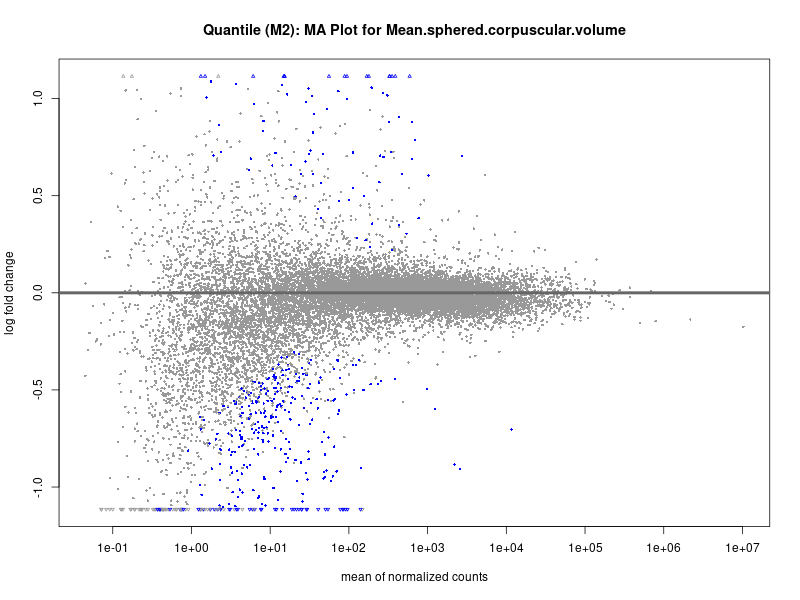
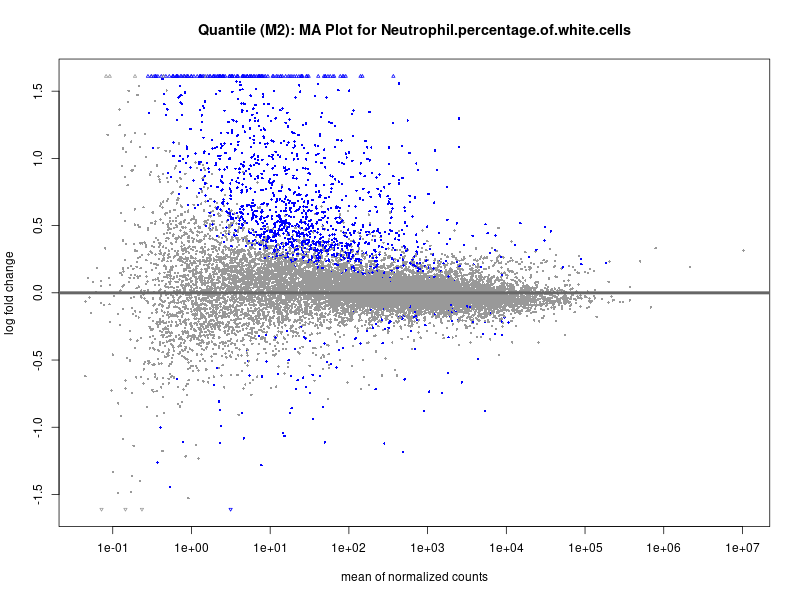
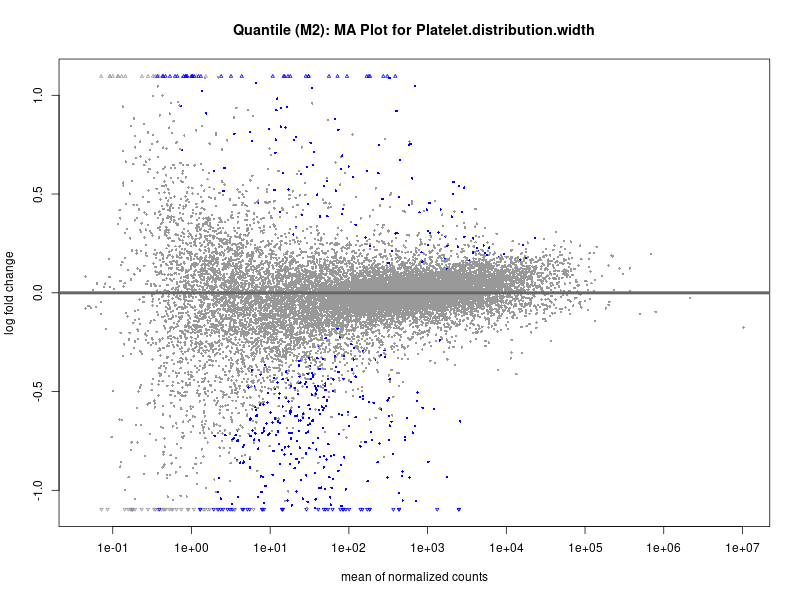
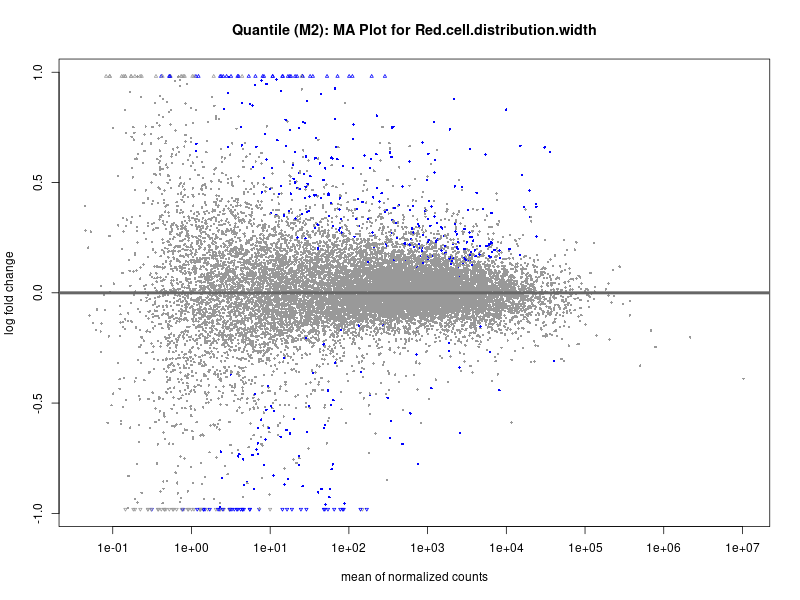
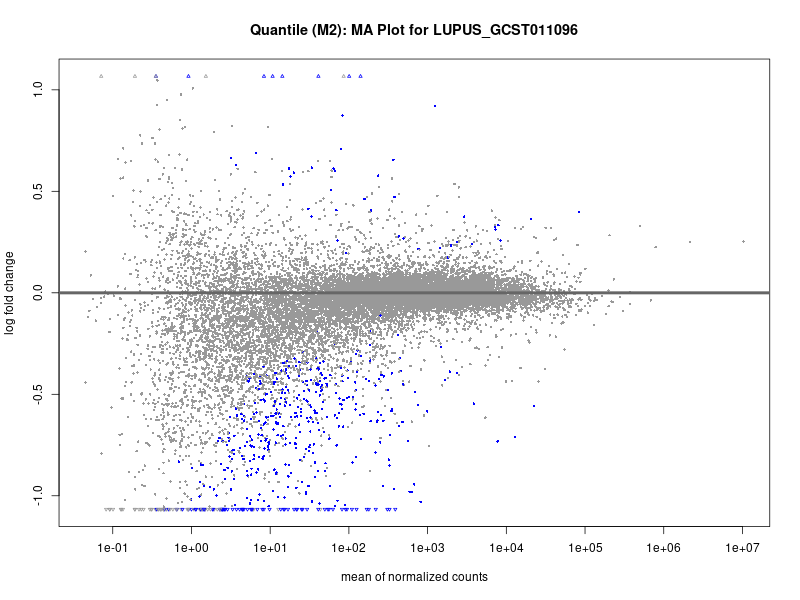
plotMA(res, main = paste("Quantile (M2): MA Plot for", trait))
dev.off()
# volcano plot
res_tableOE <- as.data.frame(res)
res_tableOE$gene_name <- raw_count_df$Description[keep_genes]
res_tableOE <- mutate(res_tableOE, threshold_OE = padj < 0.1 &
abs(log2FoldChange) >= 0.5)
res_tableOE <- res_tableOE %>% arrange(padj) %>% mutate(genelabels = "")
res_tableOE$genelabels[1:10] <- res_tableOE$gene_name[1:10]
volcano_plot <- ggplot(res_tableOE, aes(x = log2FoldChange, y = -log10(padj))) +
geom_point(aes(colour = threshold_OE)) +
geom_text_repel(aes(label = genelabels)) +
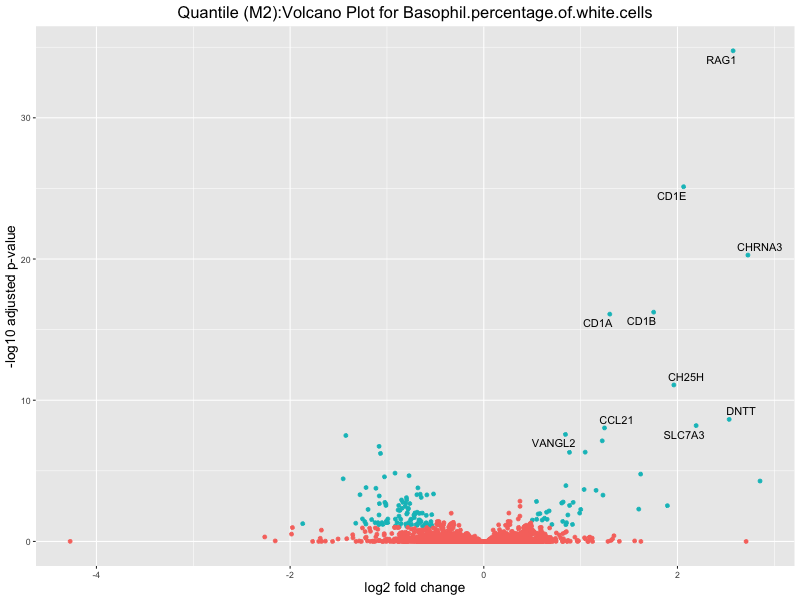
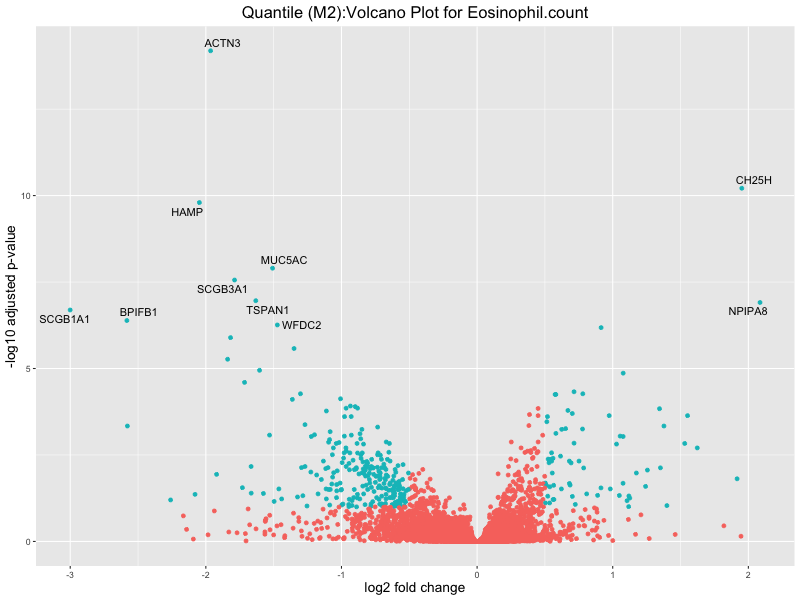
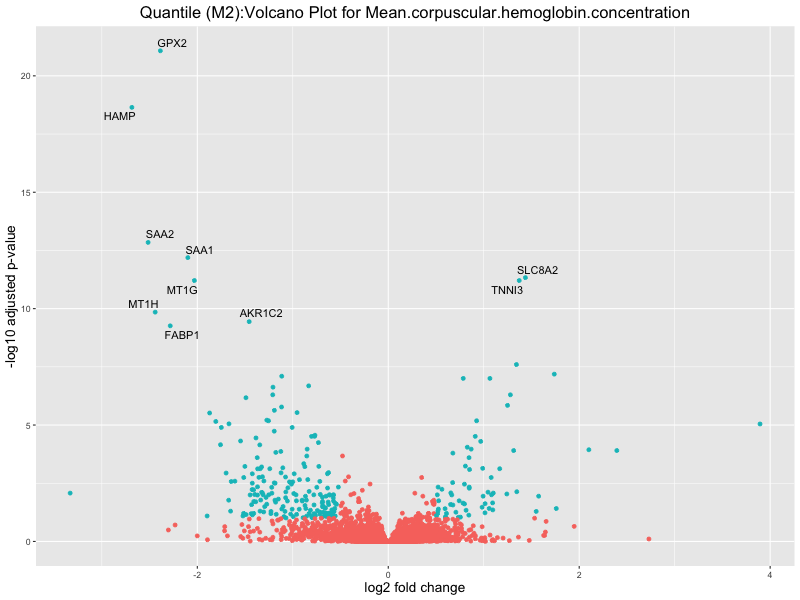
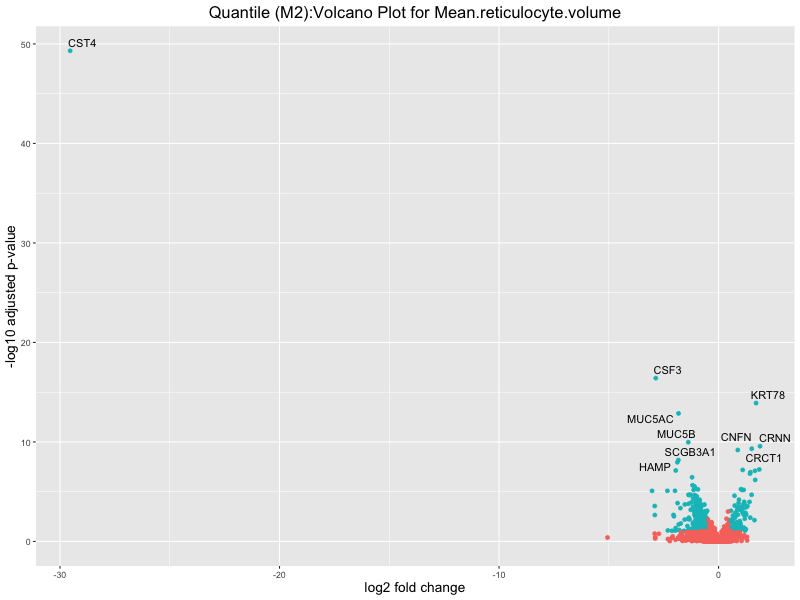
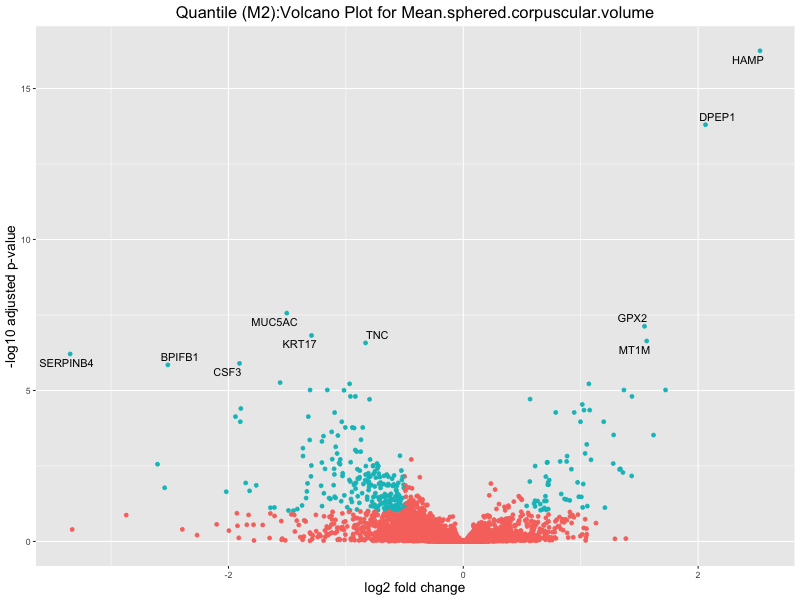
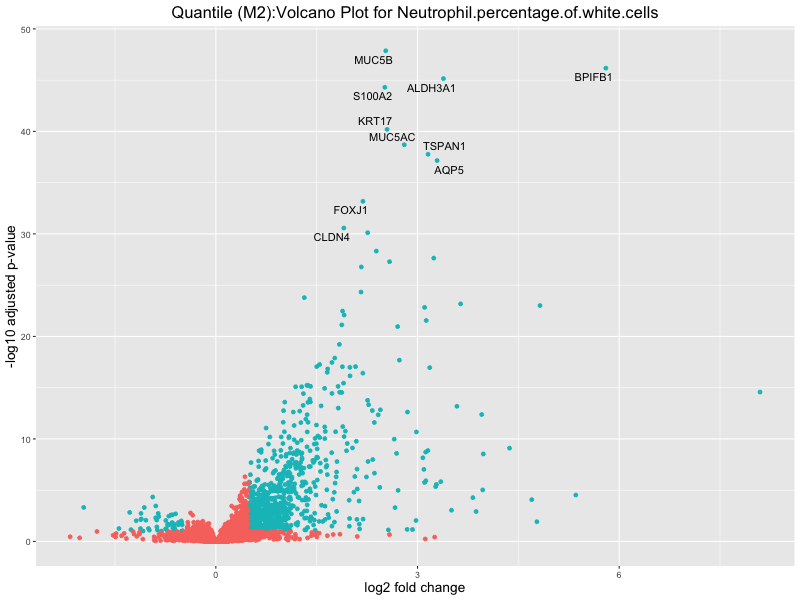
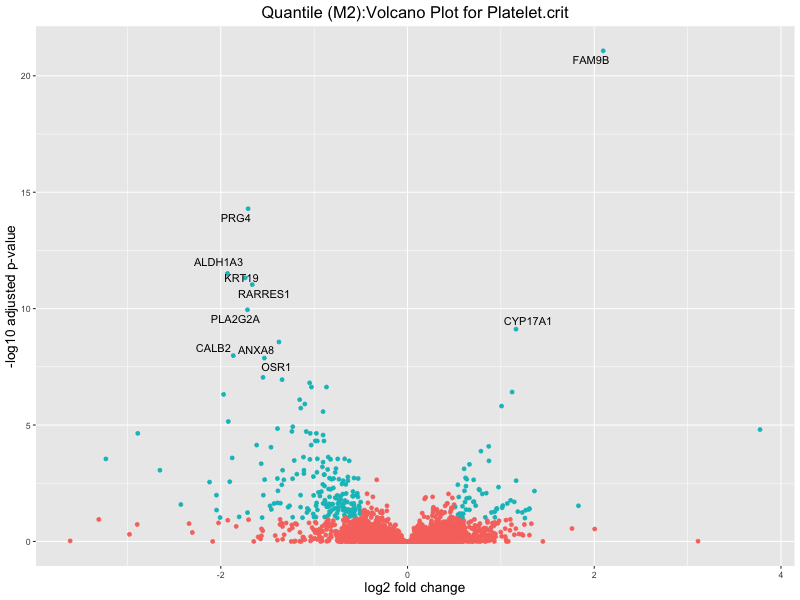
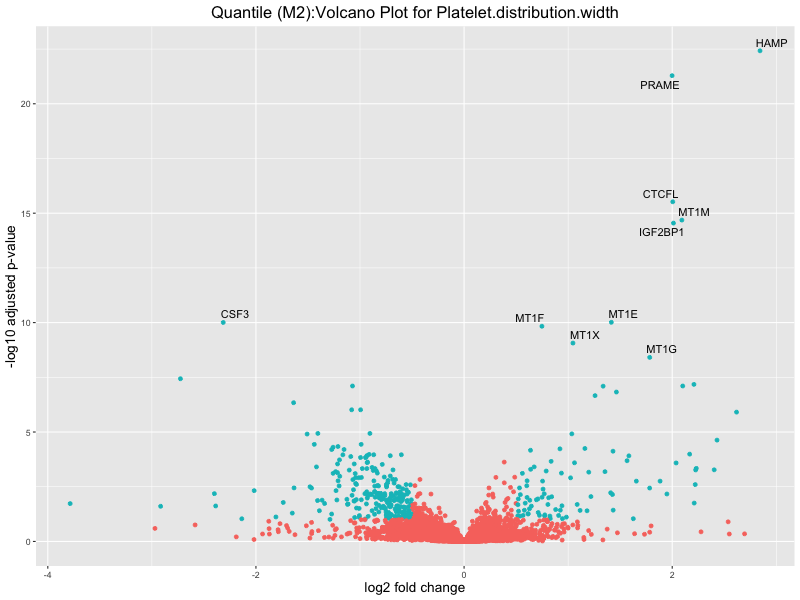
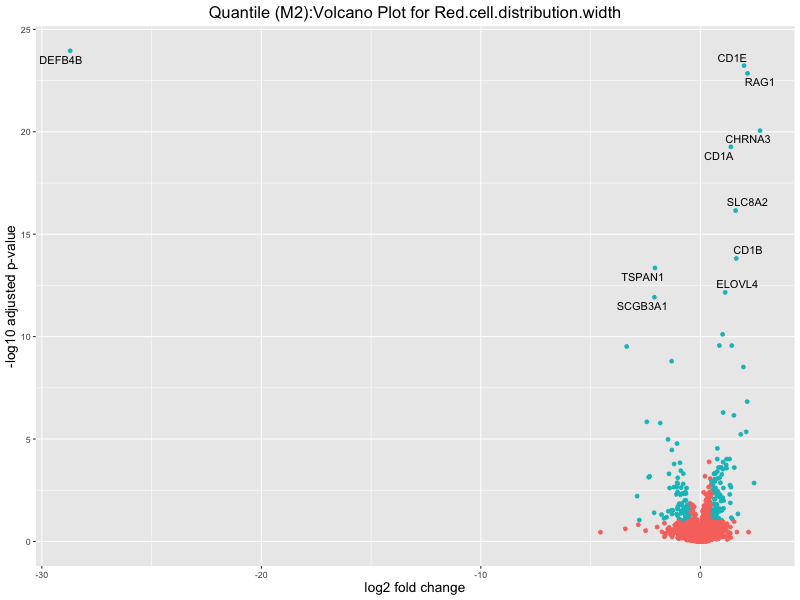
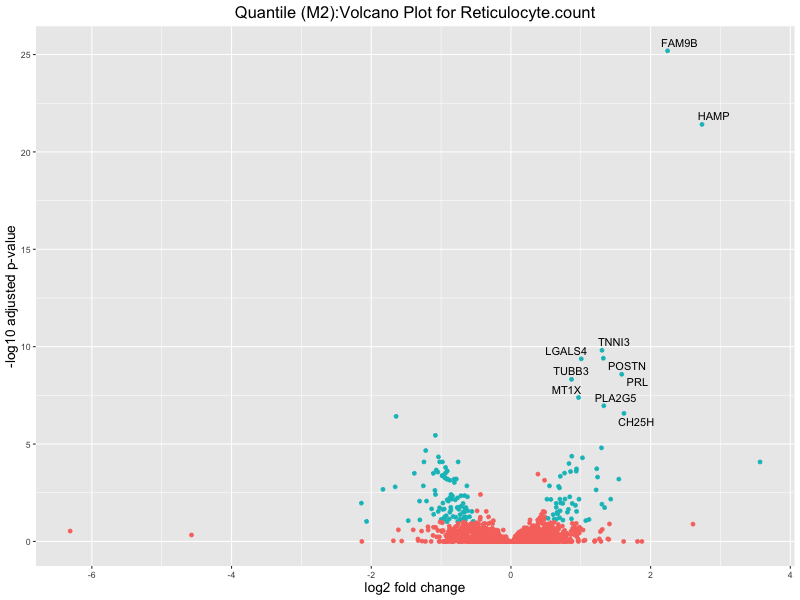
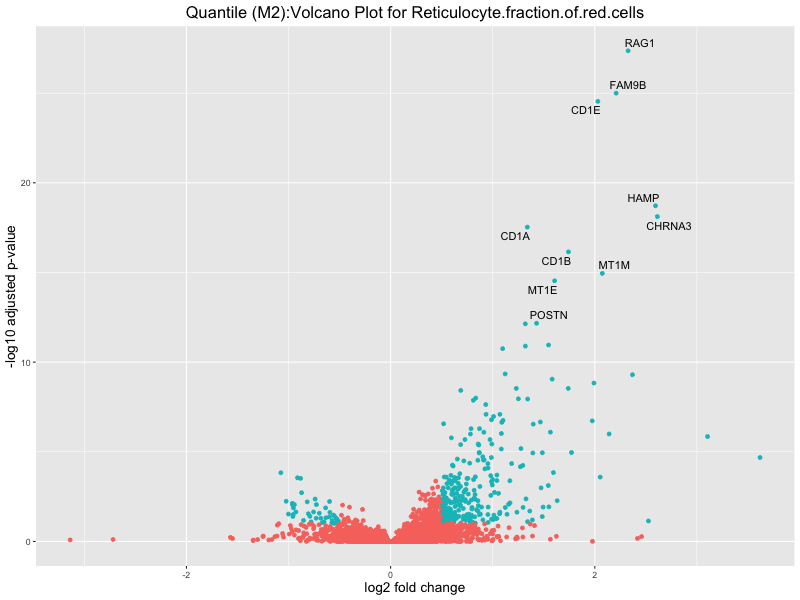
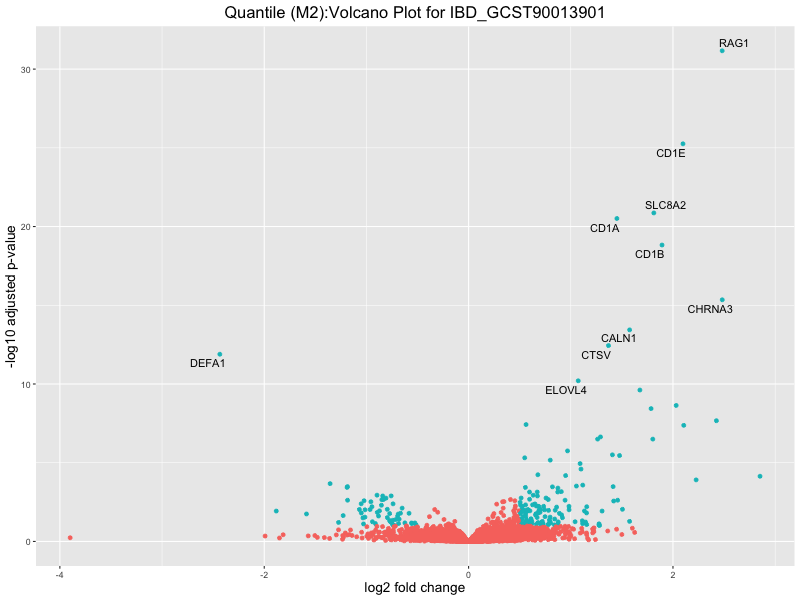
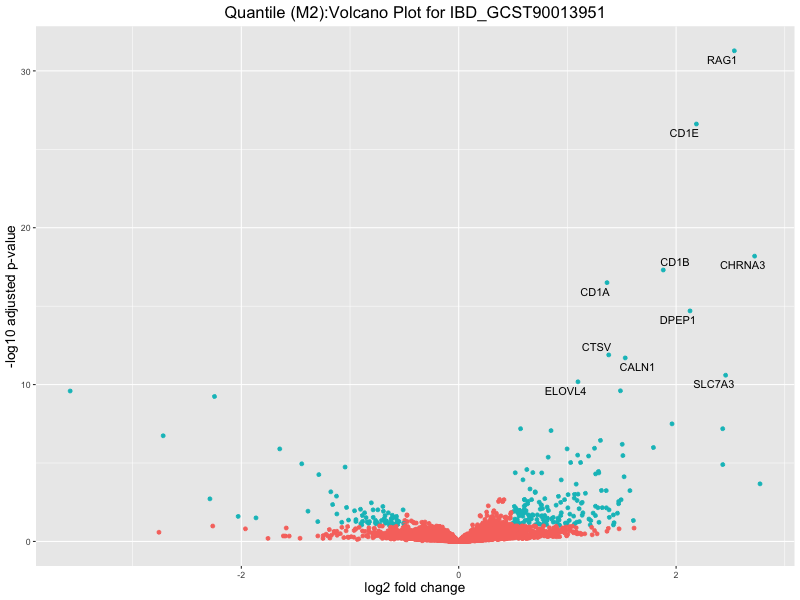
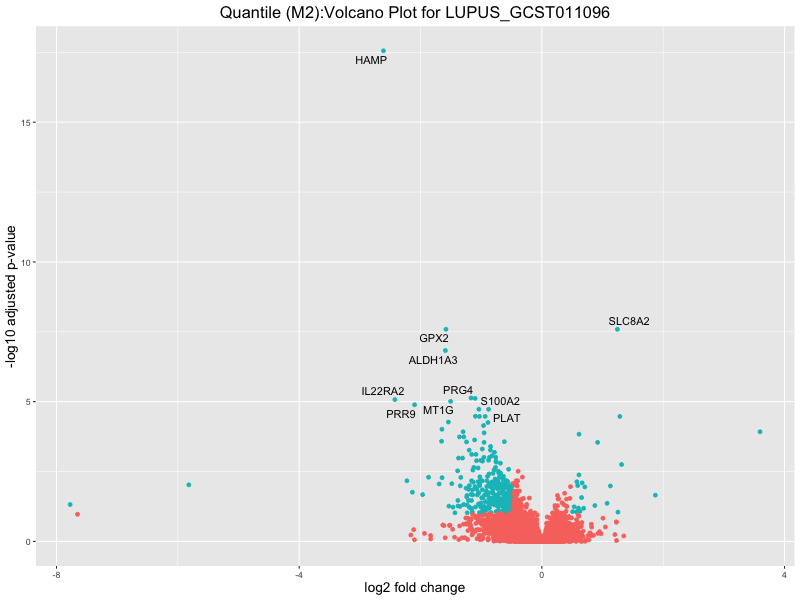
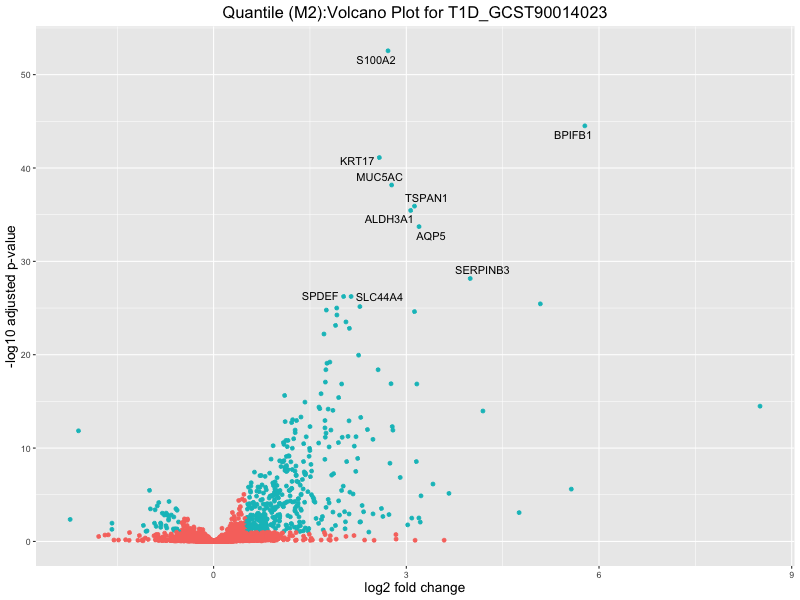
ggtitle(paste("Quantile (M2):Volcano Plot for", trait)) +
xlab("log2 fold change") +
ylab("-log10 adjusted p-value") +
theme(legend.position = "none",
plot.title = element_text(size = rel(1.5), hjust = 0.5),
axis.title = element_text(size = rel(1.25)))
# Save the volcano plot
png(paste0("volcano_plot_quantile_", trait, "_M2.png"), width = 800, height = 600)
print(volcano_plot)
dev.off()
}GO enrichment Analysis
Gene Ontology enrichment is applied to both models, with and without adjusting 2 genotype PCs, for both continuous & quantile PRS.
Continuous PRS:
dir_path <- "analysis/continuous_wb"
files <- list.files(dir_path, pattern = "differential_expression_.*_results.csv",
full.names = TRUE)
for (file in files) {
trait <- gsub("differential_expression_(.*)_results.csv", "\\1", basename(file))
trait
res_tableOE <- read.csv(file, header = T, row.names = 1)
deGenes <- res_tableOE[res_tableOE$padj < 0.1 &
abs(res_tableOE$log2FoldChange) >= 0.5, ]
deGenes$gene_id <- gsub("\\.\\d+$", "", rownames(deGenes))
# Separate upregulated and downregulated genes
upregulated_genes <- deGenes[deGenes$log2FoldChange > 0, ]$gene_id
downregulated_genes <- deGenes[deGenes$log2FoldChange < 0, ]$gene_id
# Run GO enrichment for upregulated genes
gse_up <- enrichGO(gene = upregulated_genes, ont = "BP",
OrgDb = "org.Hs.eg.db", keyType = "ENSEMBL", readable = T)
# Run GO enrichment for downregulated genes
gse_down <- enrichGO(gene = downregulated_genes, ont = "BP",
OrgDb = "org.Hs.eg.db", keyType = "ENSEMBL", readable = T)
# Convert enrichment results to data frames and calculate additional ratios
if (is.null(gse_up)) {
gse_up <- data.frame(ID = character(), Description = character(),
GeneRatio = character(), BgRatio = character(),
pvalue = numeric(), p.adjust = numeric(),
qvalue = numeric(), geneID = character(),
Count = integer(), stringsAsFactors = FALSE)
} else {
gse_up <- as.data.frame(gse_up)}
gse_down <- as.data.frame(gse_down)
gse_up$GeneRatio_num <- as.numeric(sapply(strsplit(gse_up$GeneRatio, "/"), function(x) x[1])) /
as.numeric(sapply(strsplit(gse_up$GeneRatio, "/"), function(x) x[2]))
gse_up$BgRatio_num <- as.numeric(sapply(strsplit(gse_up$BgRatio, "/"), function(x) x[1])) /
as.numeric(sapply(strsplit(gse_up$BgRatio, "/"), function(x) x[2]))
gse_up <- cbind(gse_up, FoldEnrich = gse_up$GeneRatio_num / gse_up$BgRatio_num)
gse_down$GeneRatio_num <- as.numeric(sapply(strsplit(gse_down$GeneRatio, "/"), function(x) x[1])) /
as.numeric(sapply(strsplit(gse_down$GeneRatio, "/"), function(x) x[2]))
gse_down$BgRatio_num <- as.numeric(sapply(strsplit(gse_down$BgRatio, "/"), function(x) x[1])) /
as.numeric(sapply(strsplit(gse_down$BgRatio, "/"), function(x) x[2]))
gse_down <- cbind(gse_down, FoldEnrich = gse_down$GeneRatio_num / gse_down$BgRatio_num)
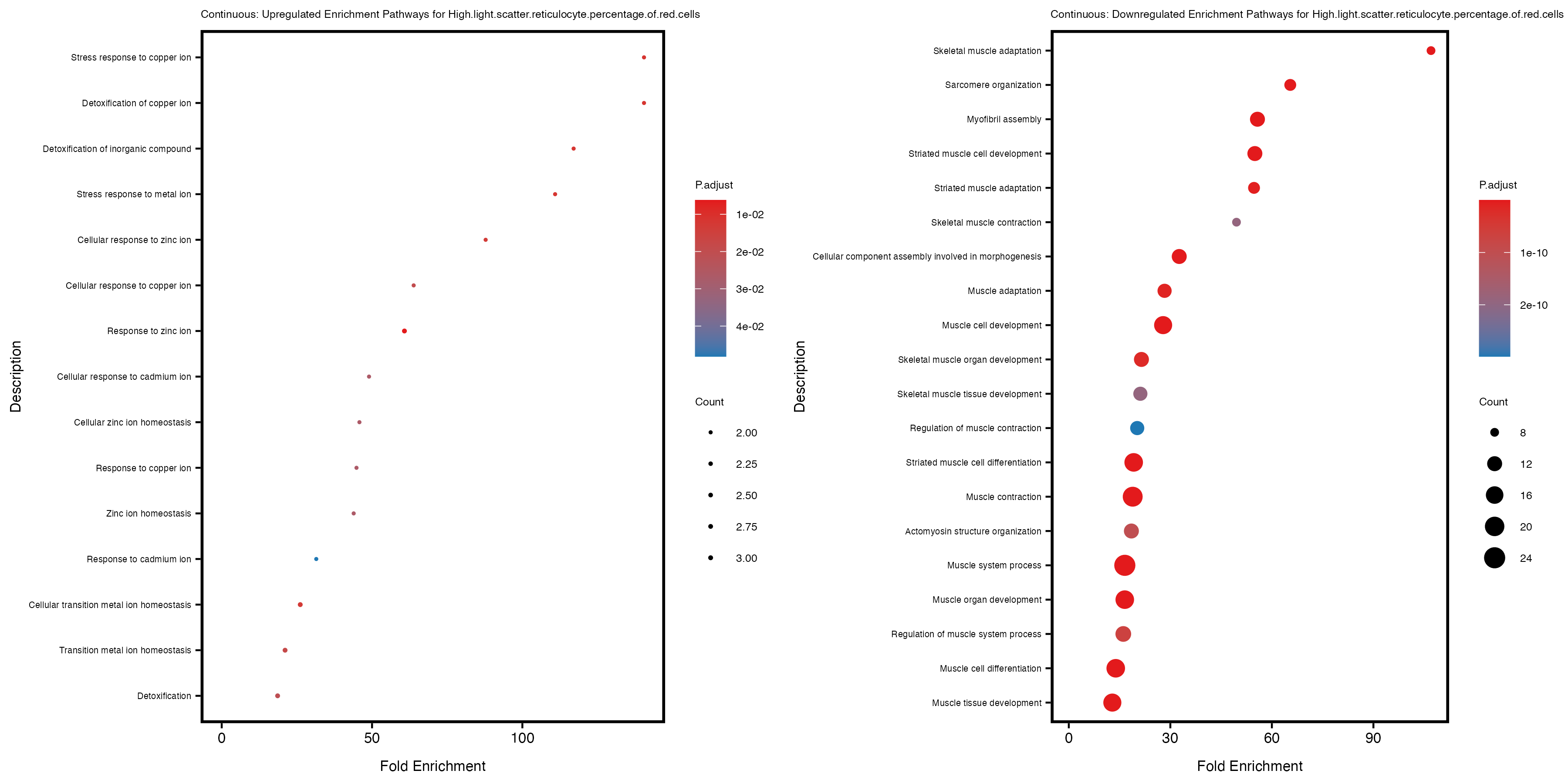
if (nrow(gse_up) >= 20) {
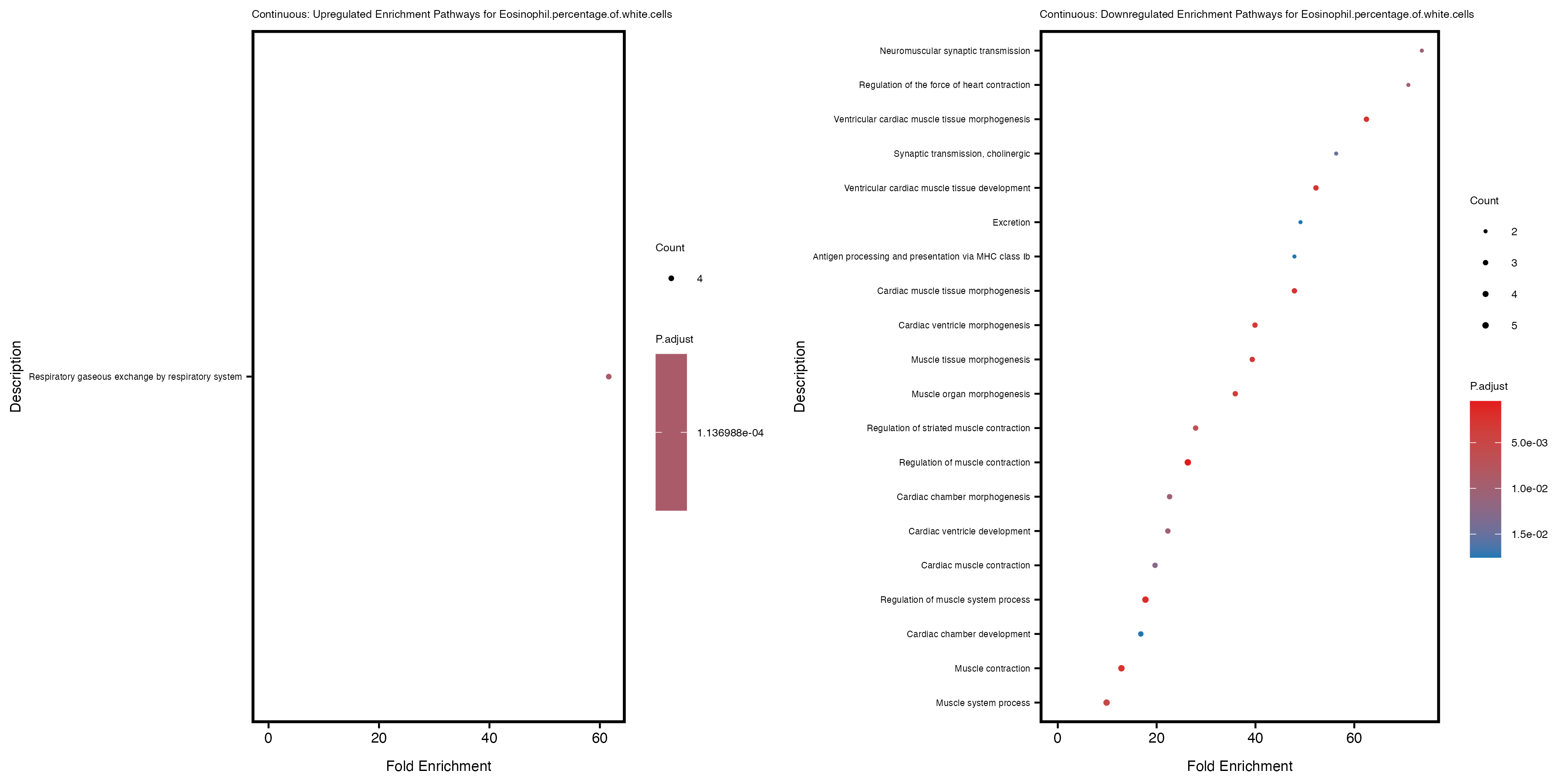
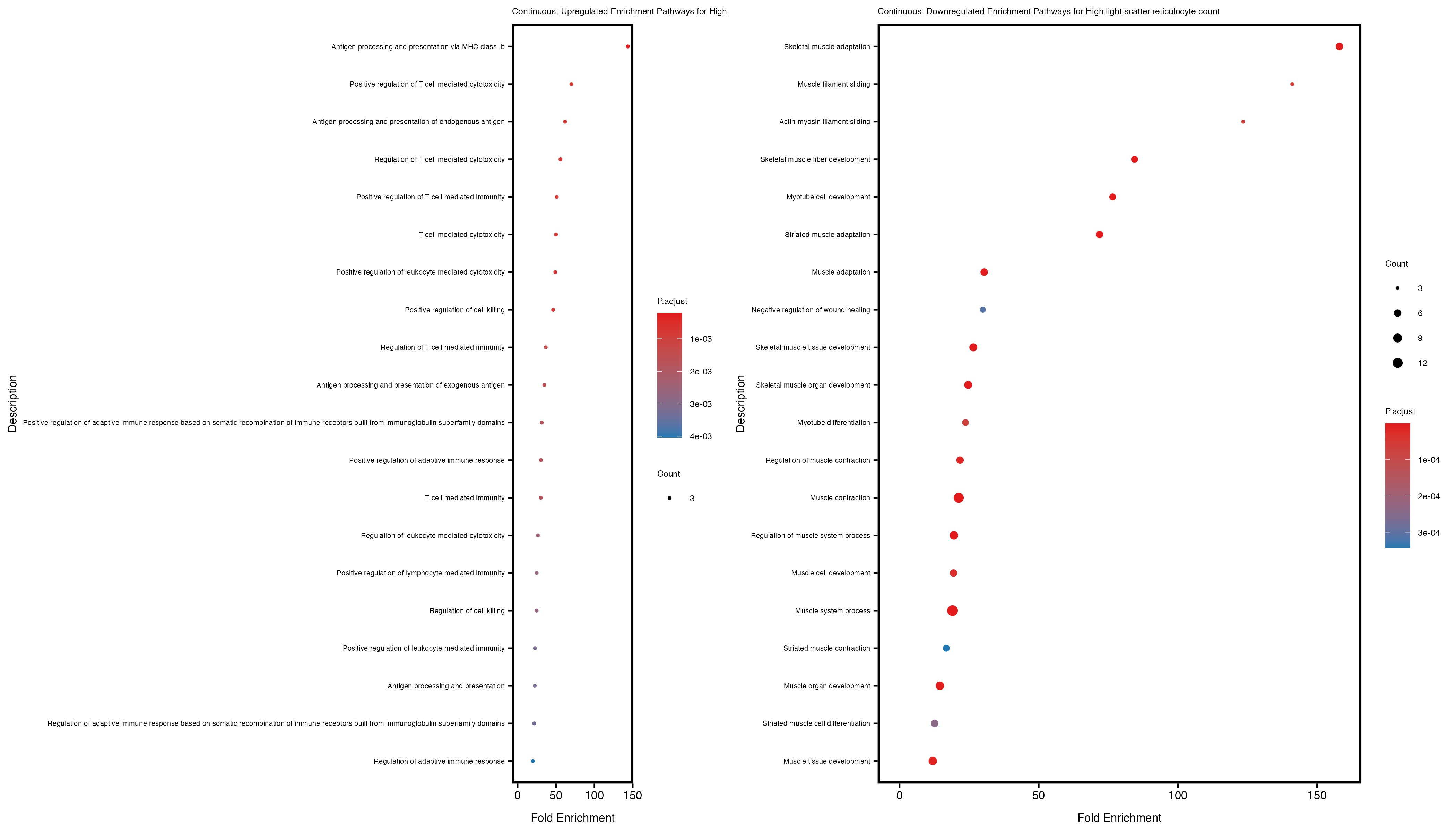
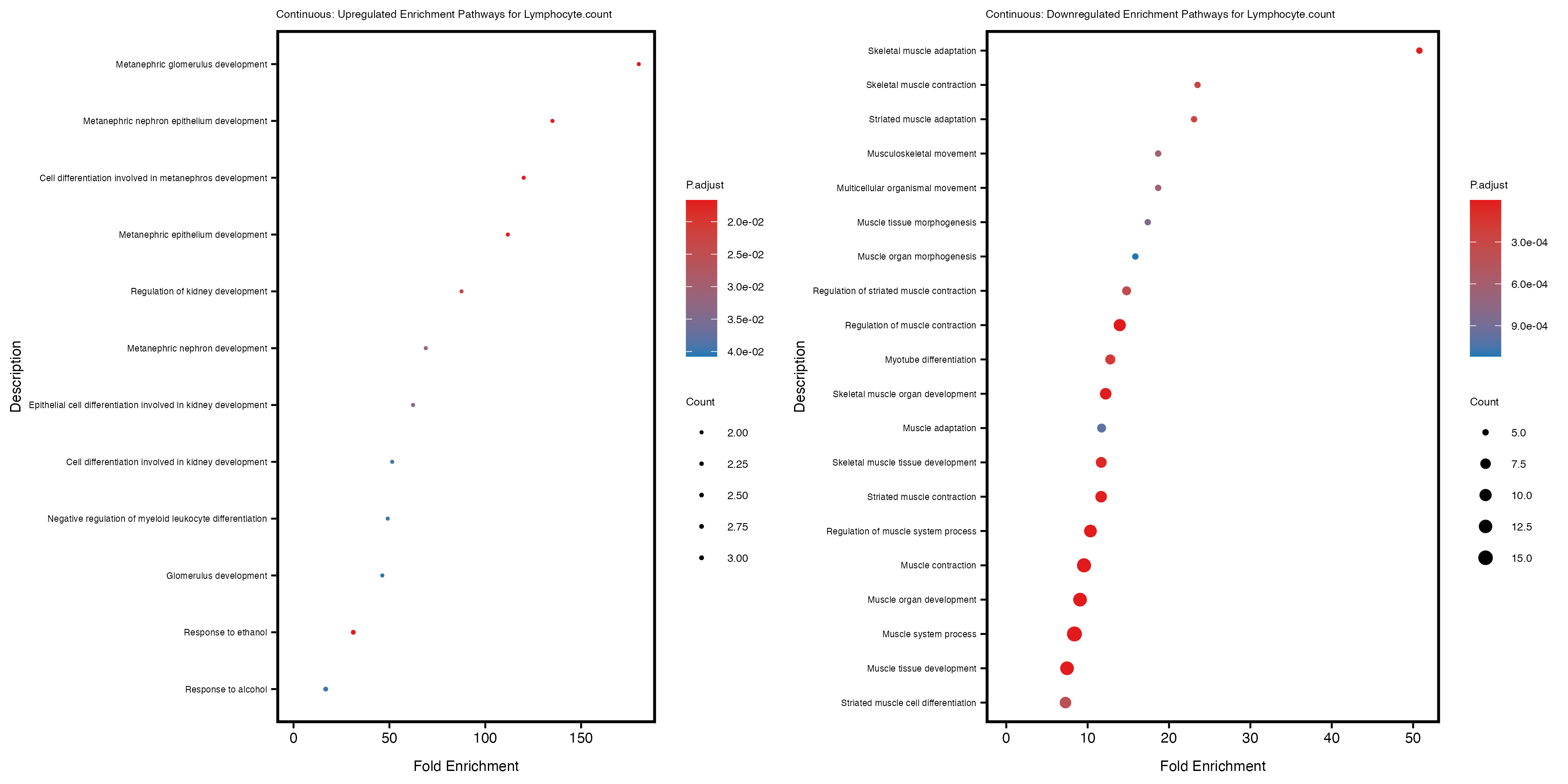
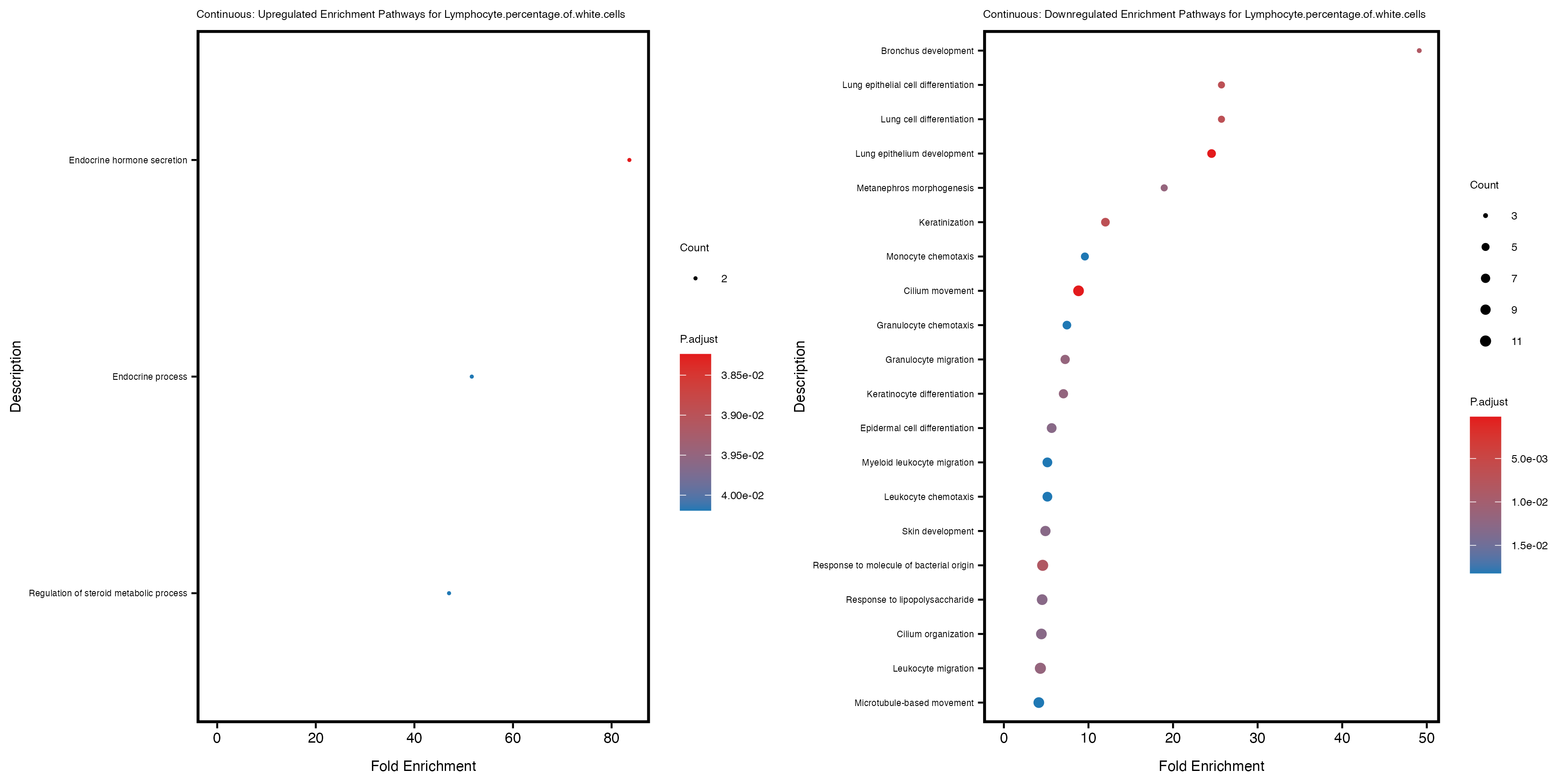
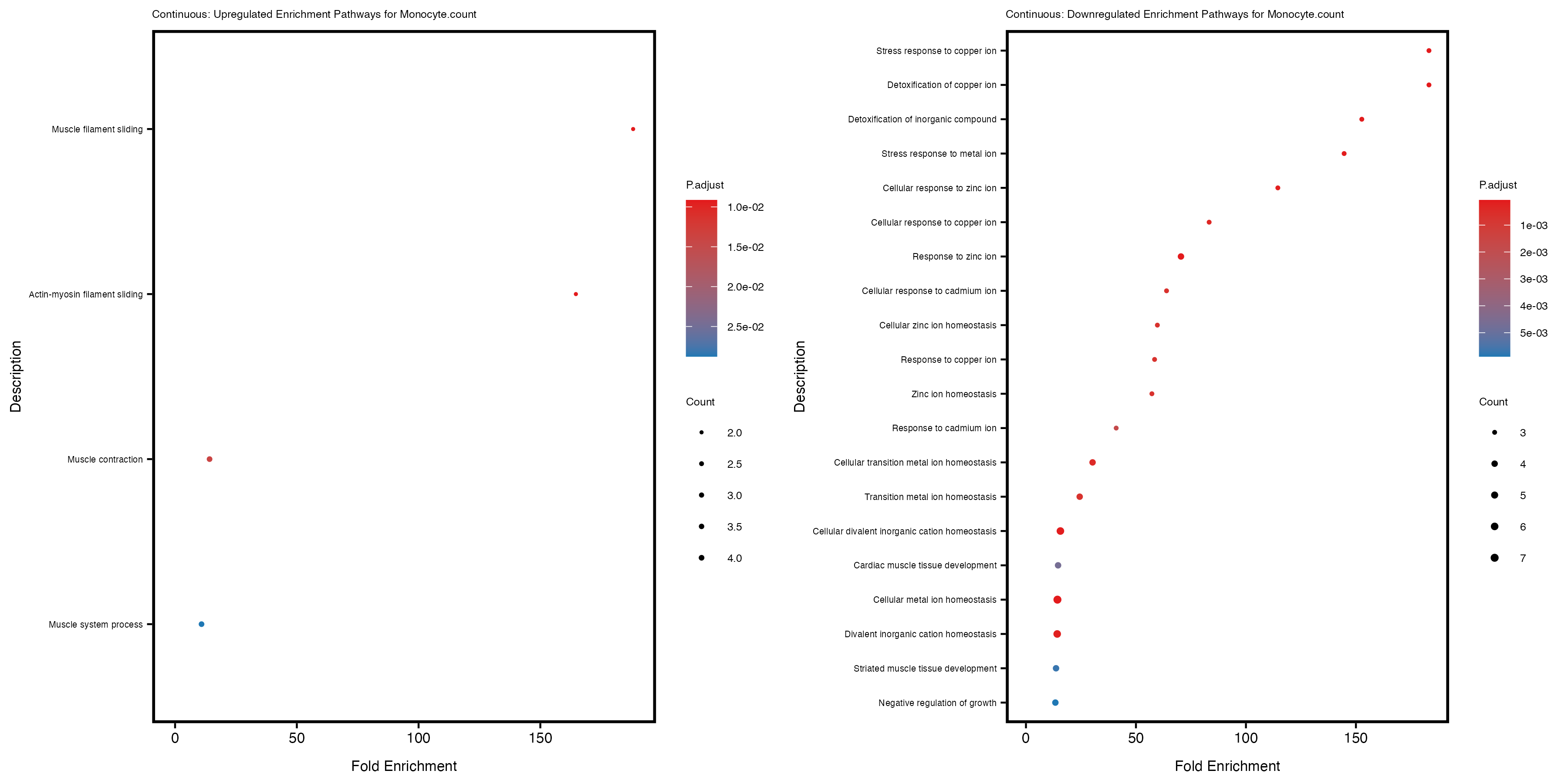
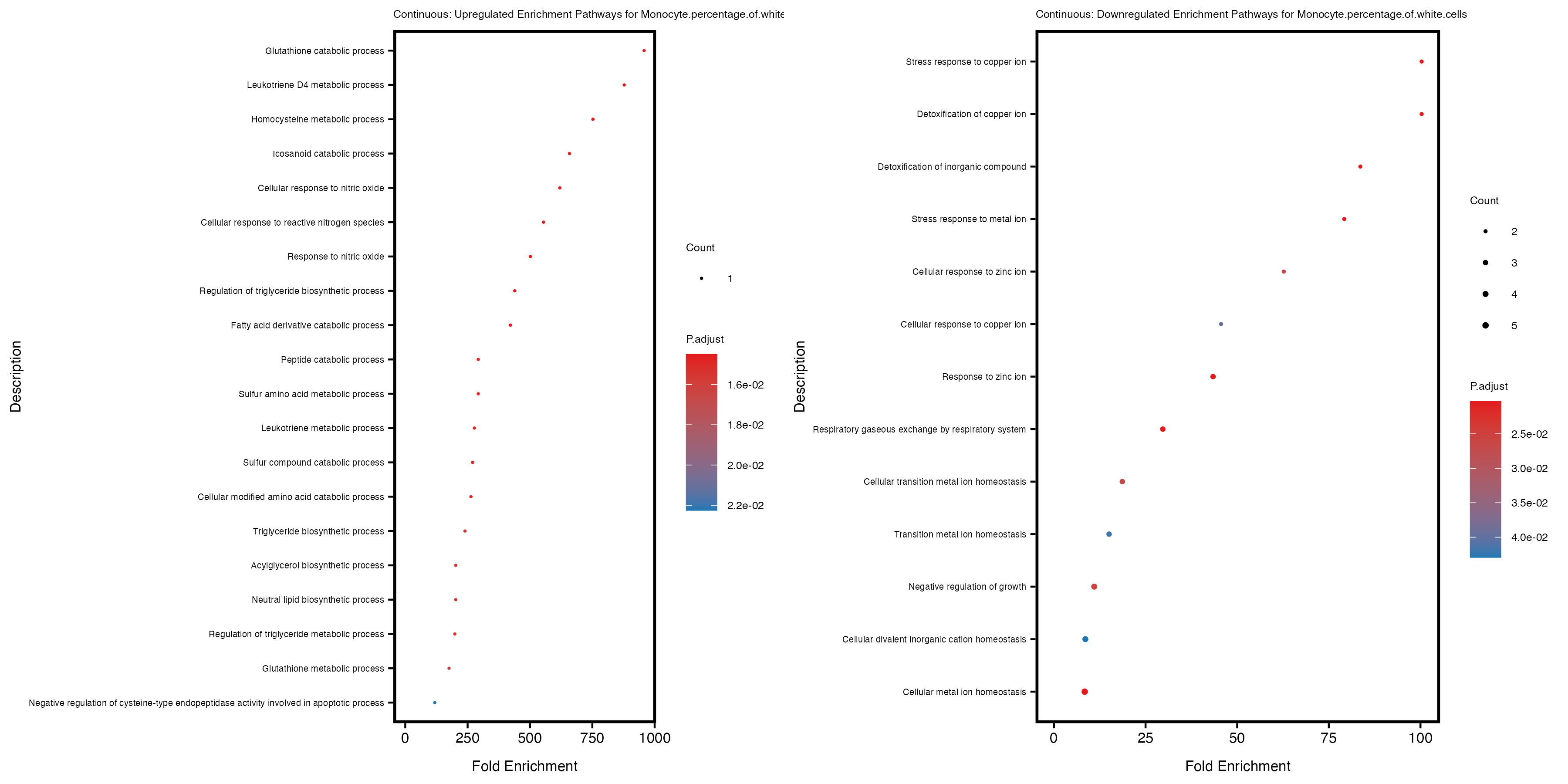
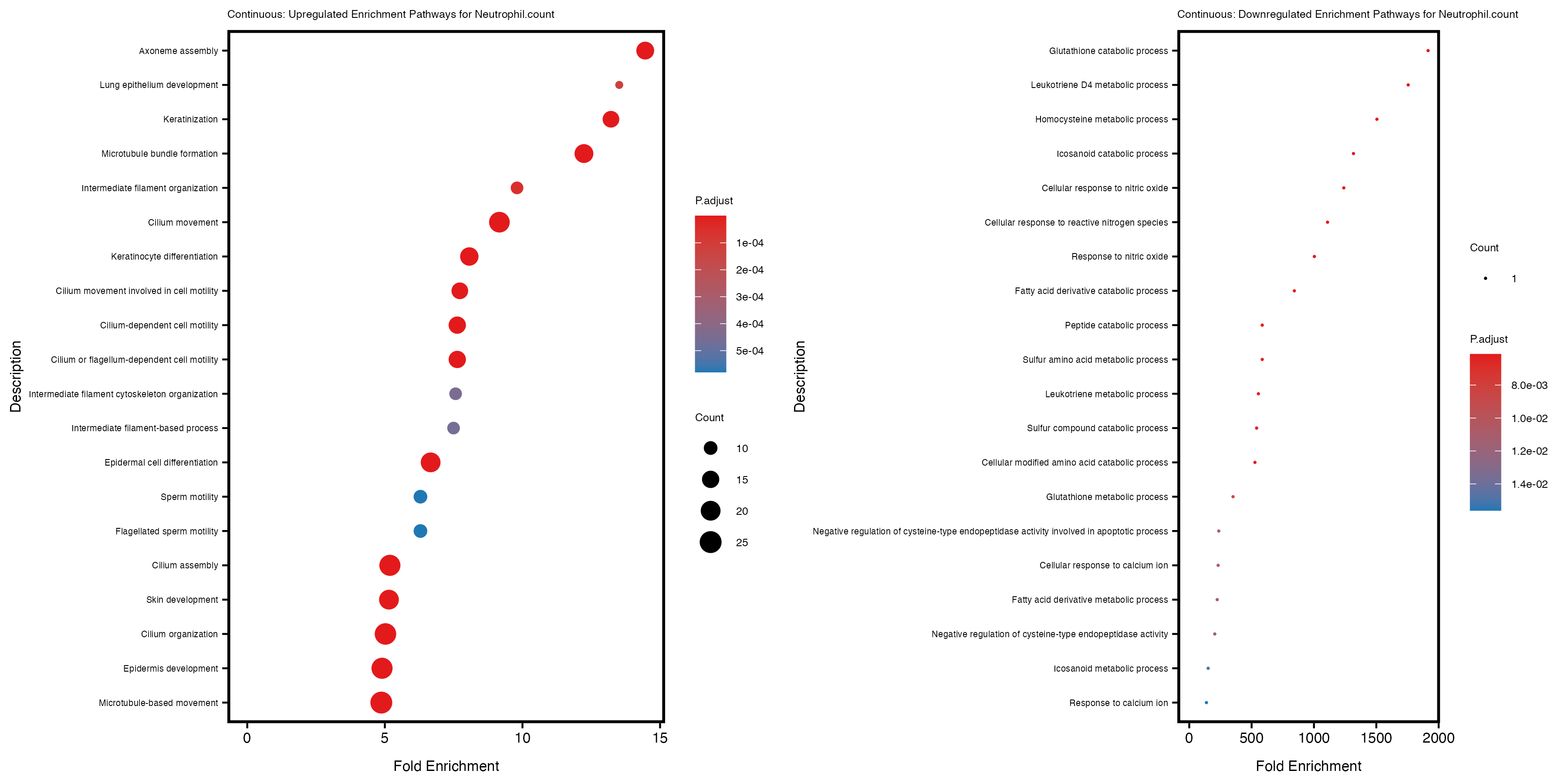
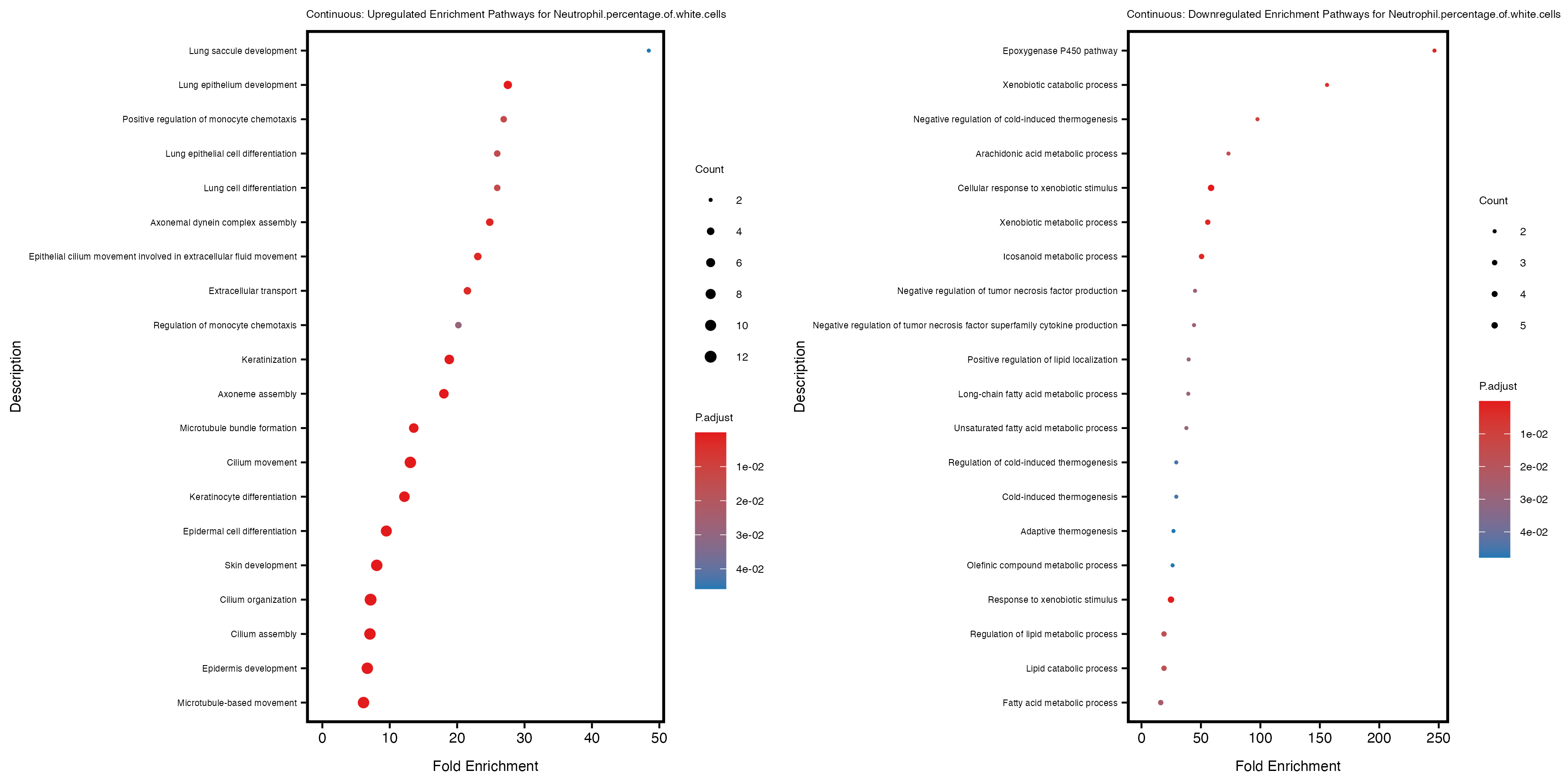
enrich_plot_up <- plotEnrich(gse_up[1:20, ], plot_type = "dot", scale_ratio = 0.4) +
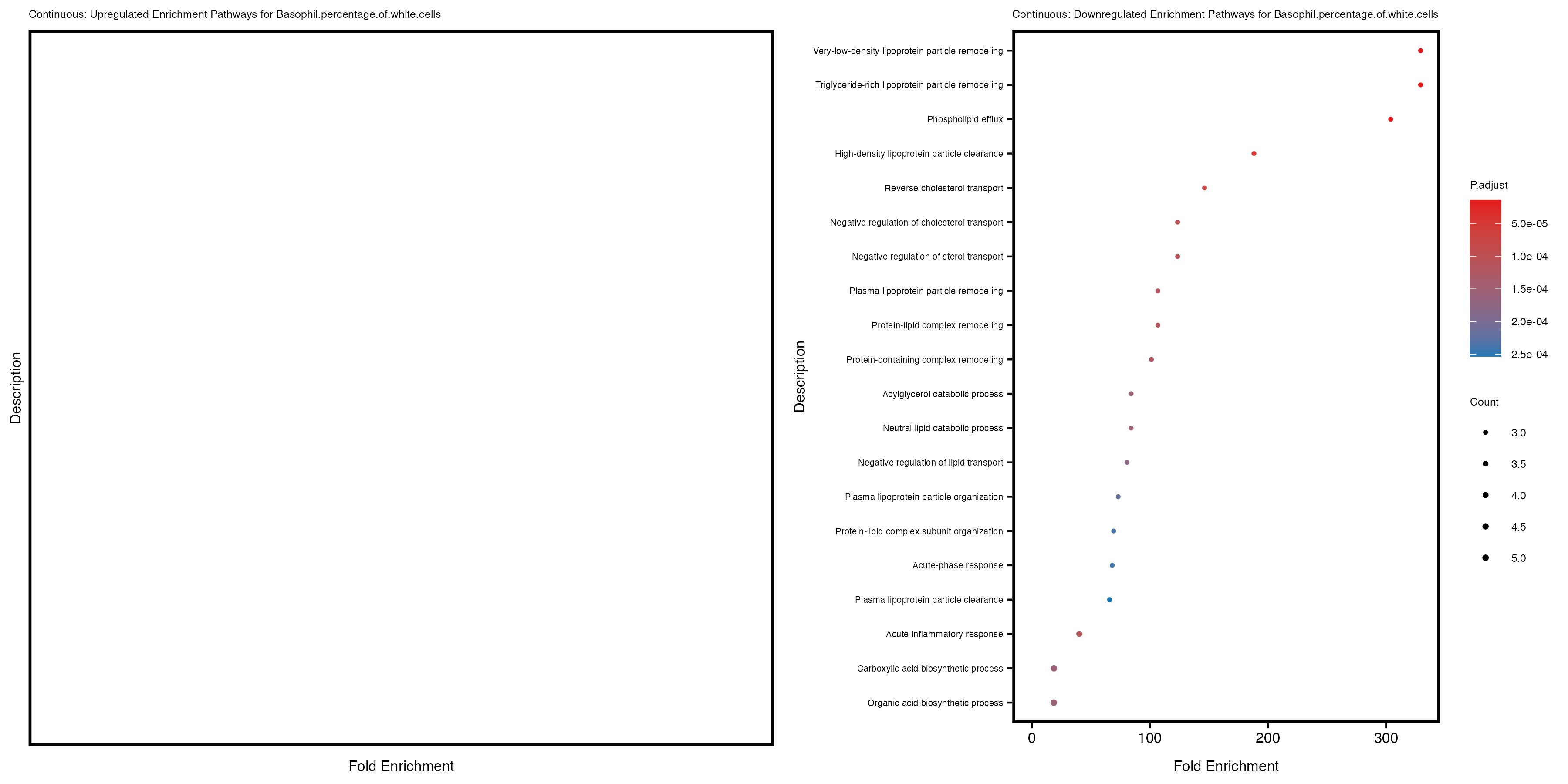
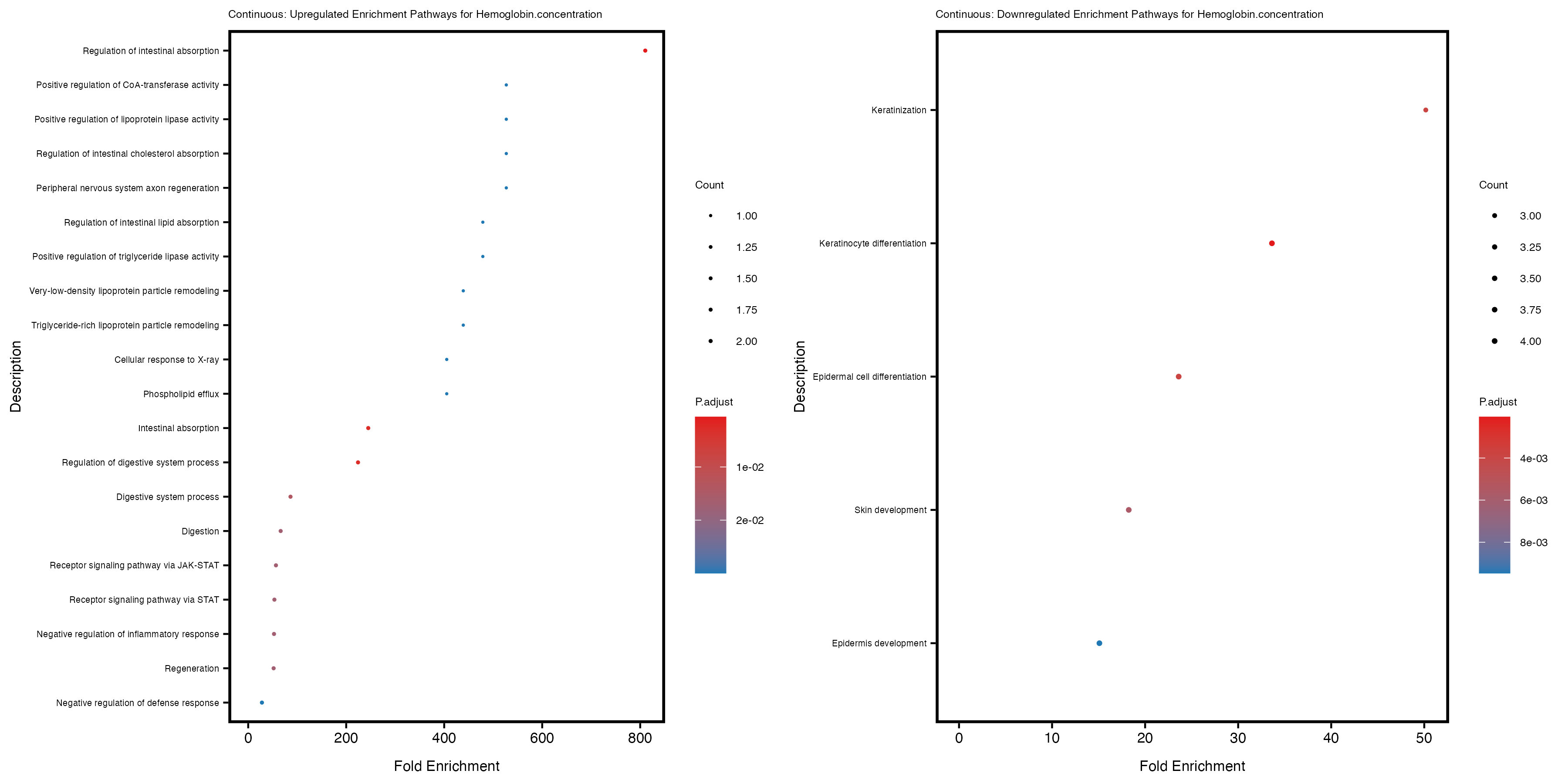
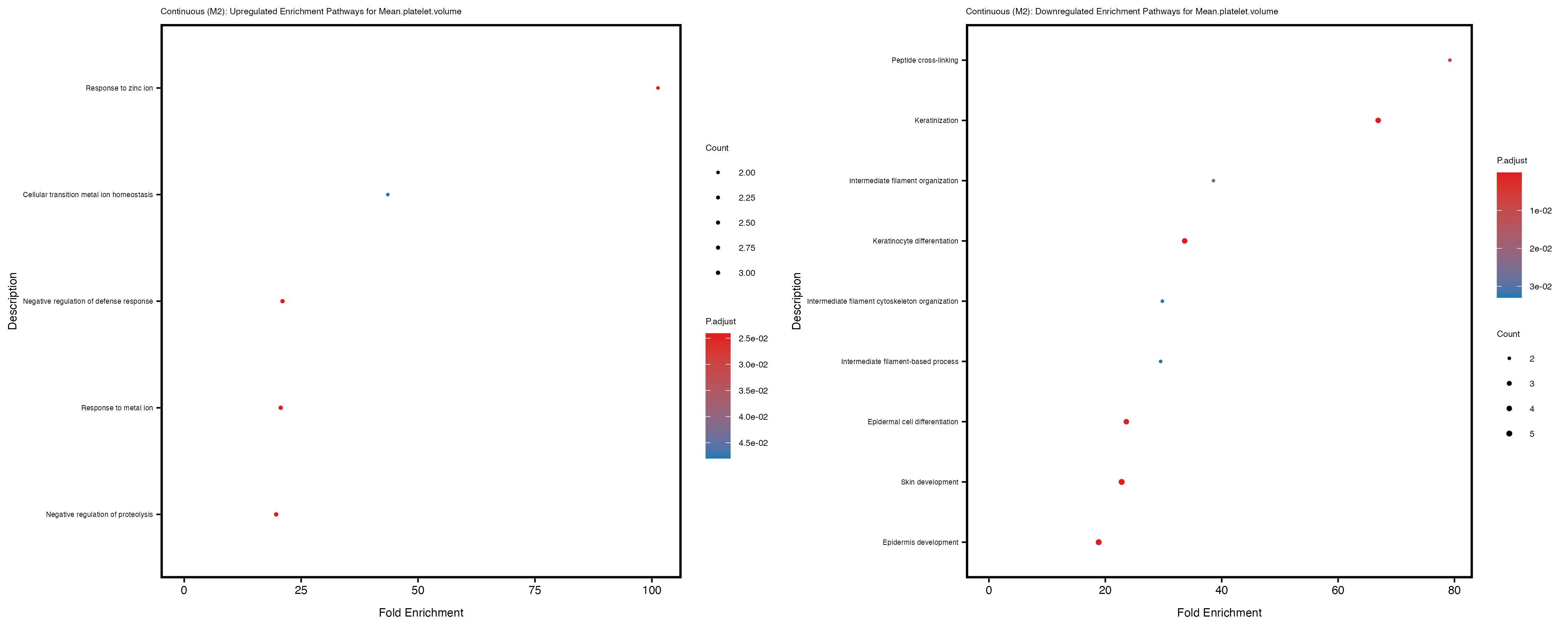
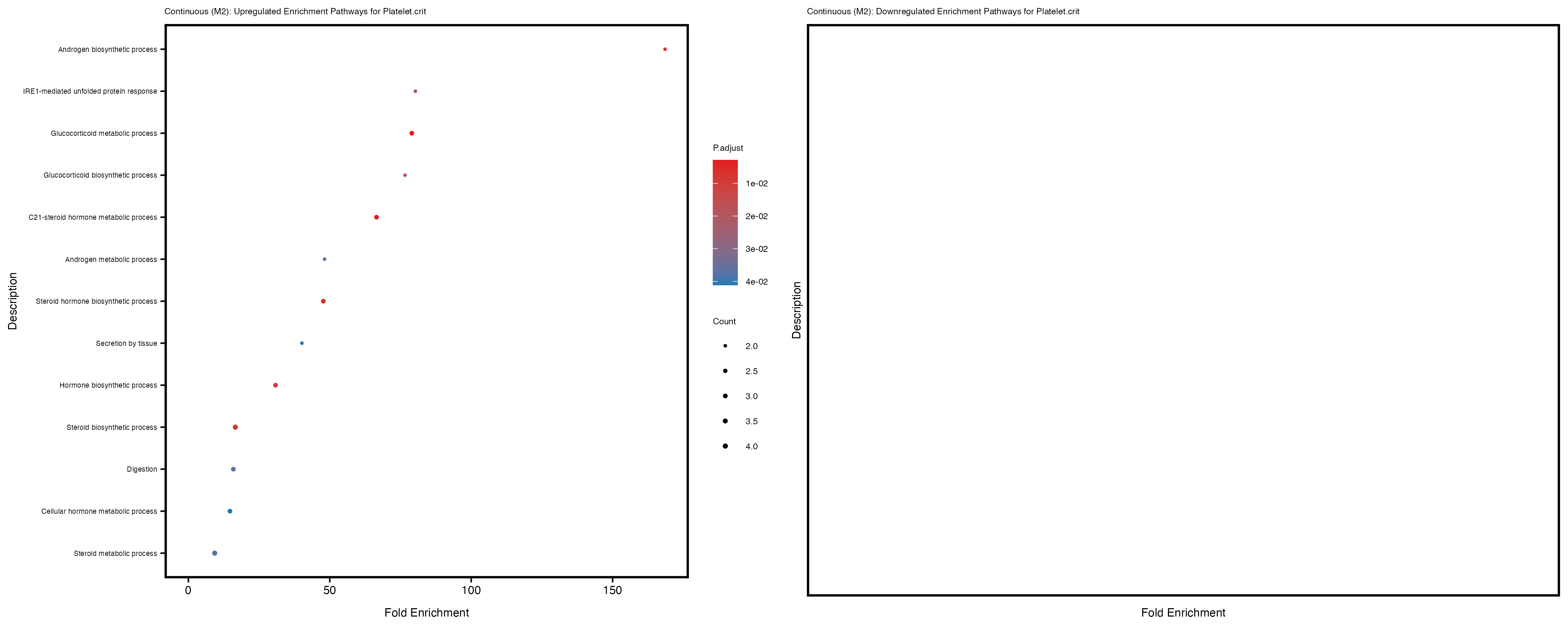
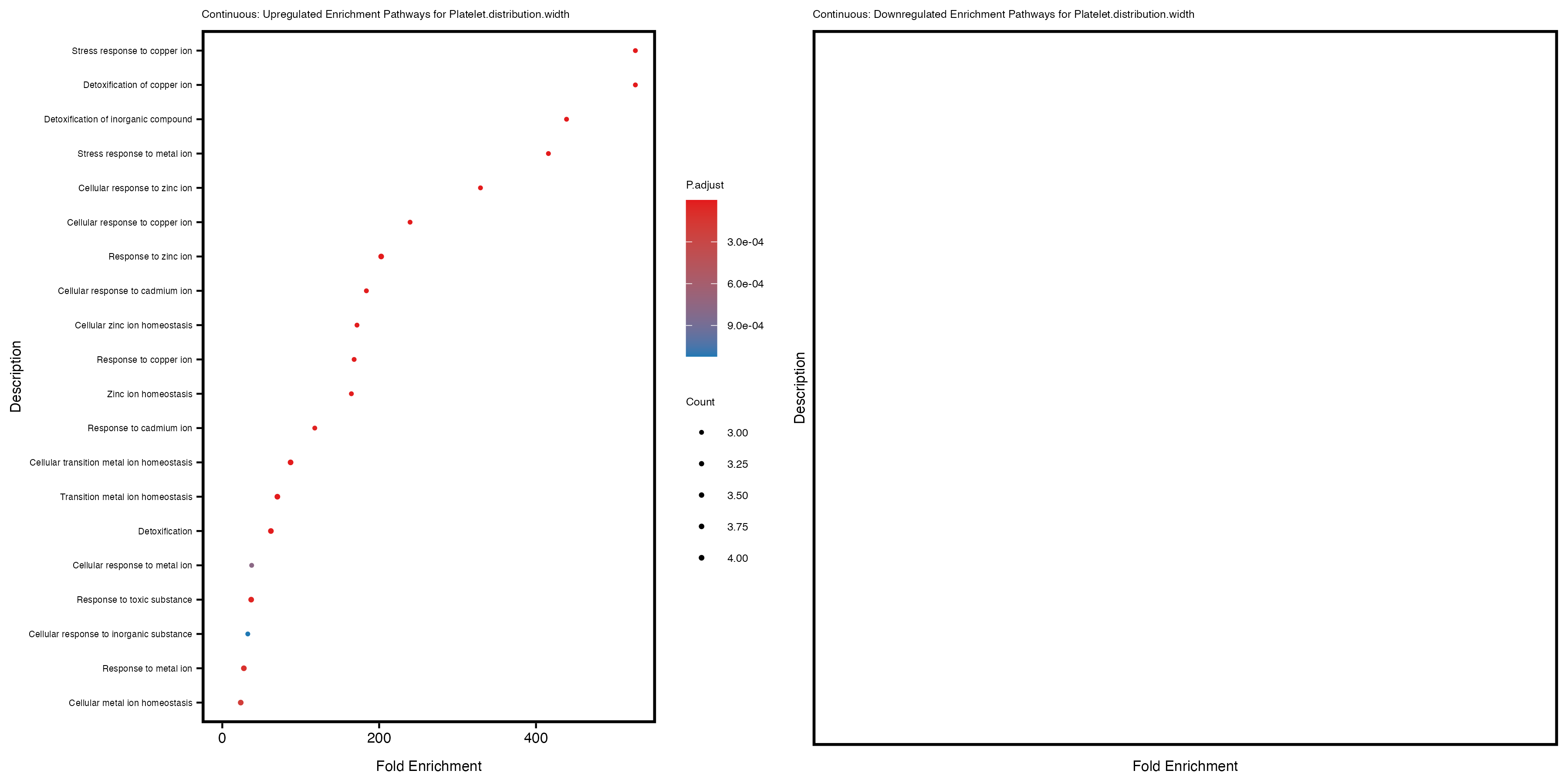
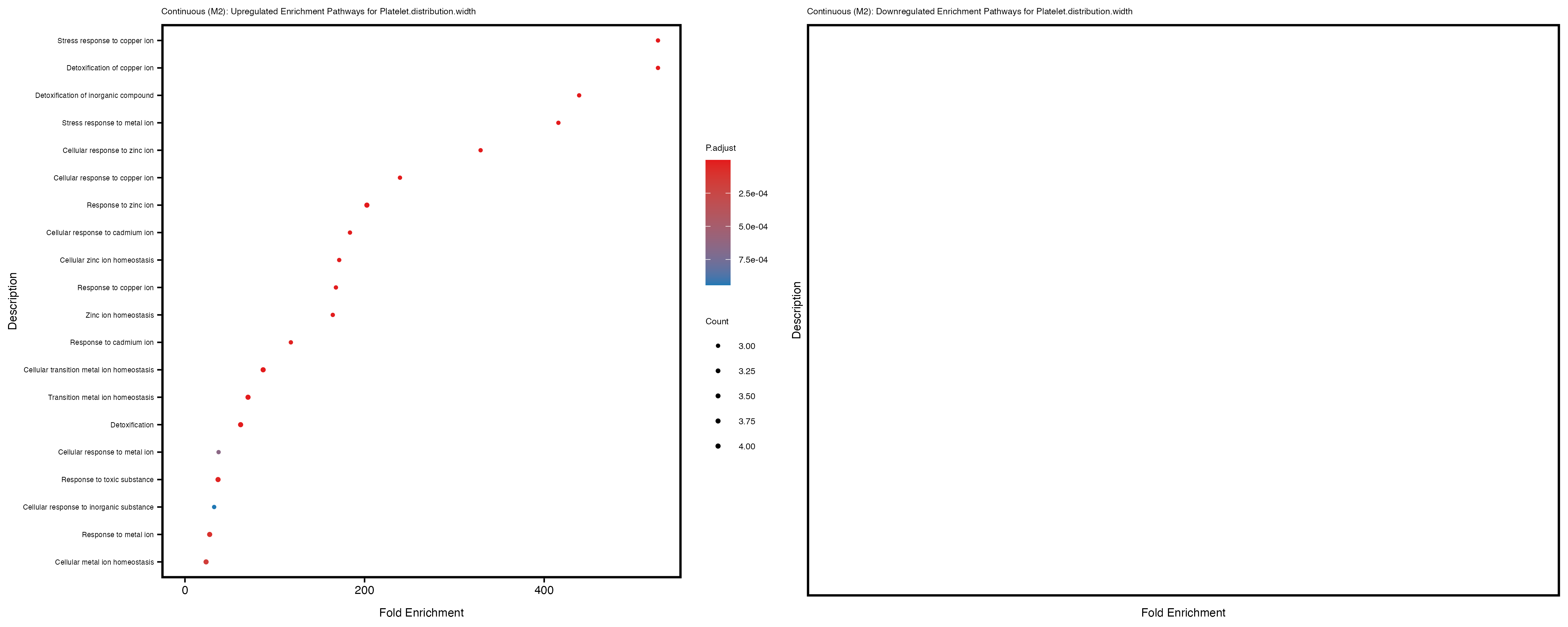
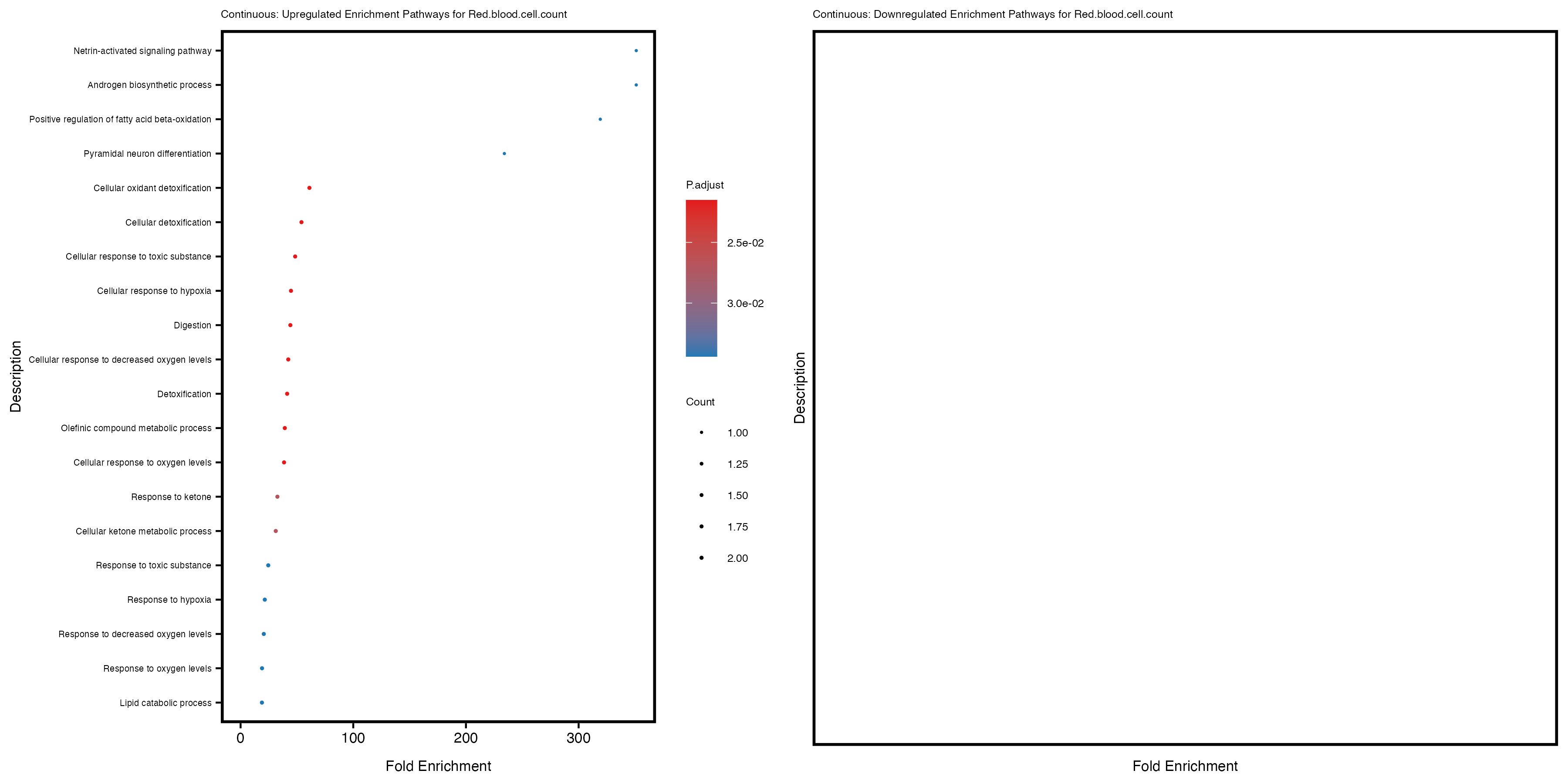
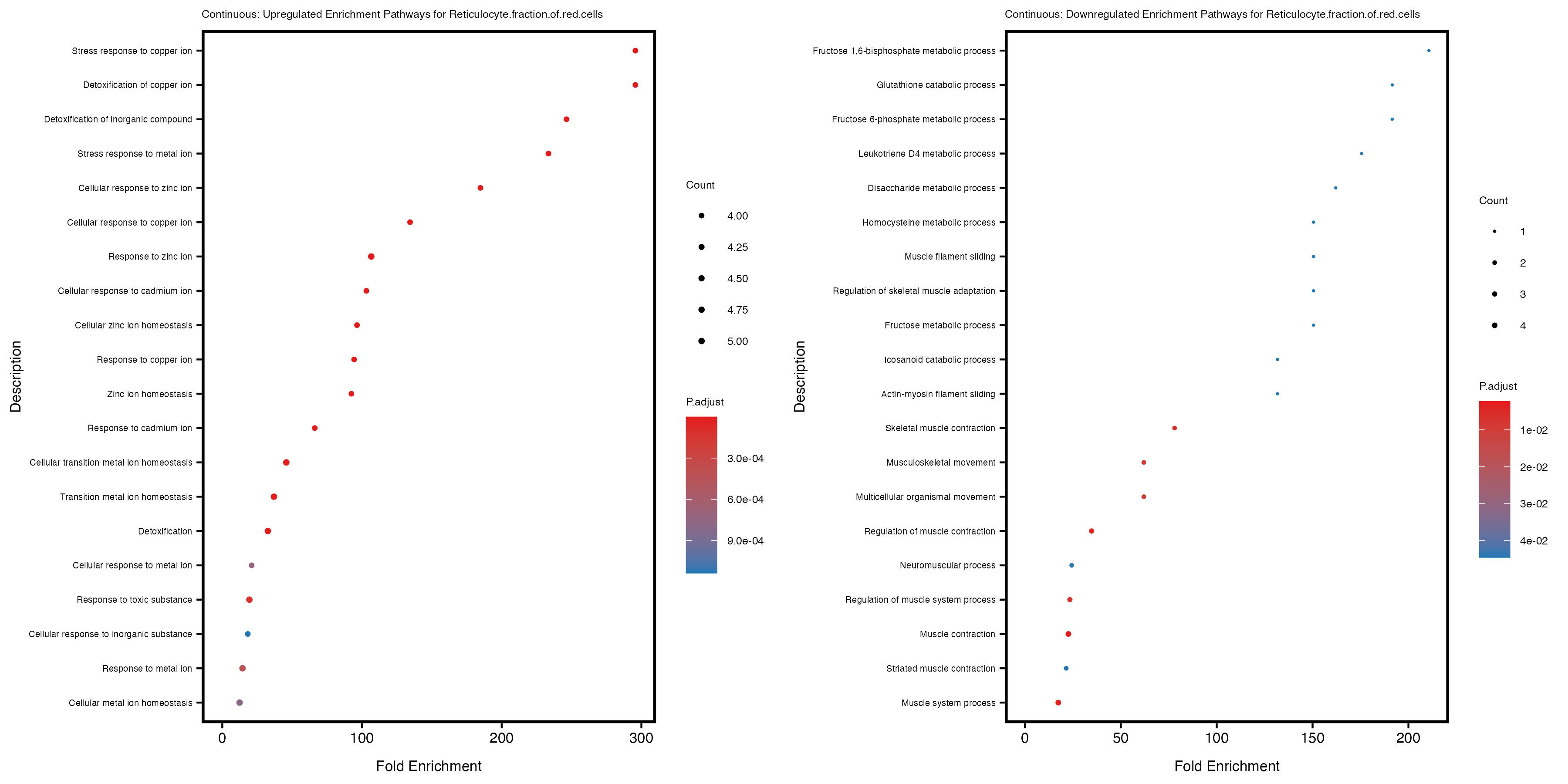
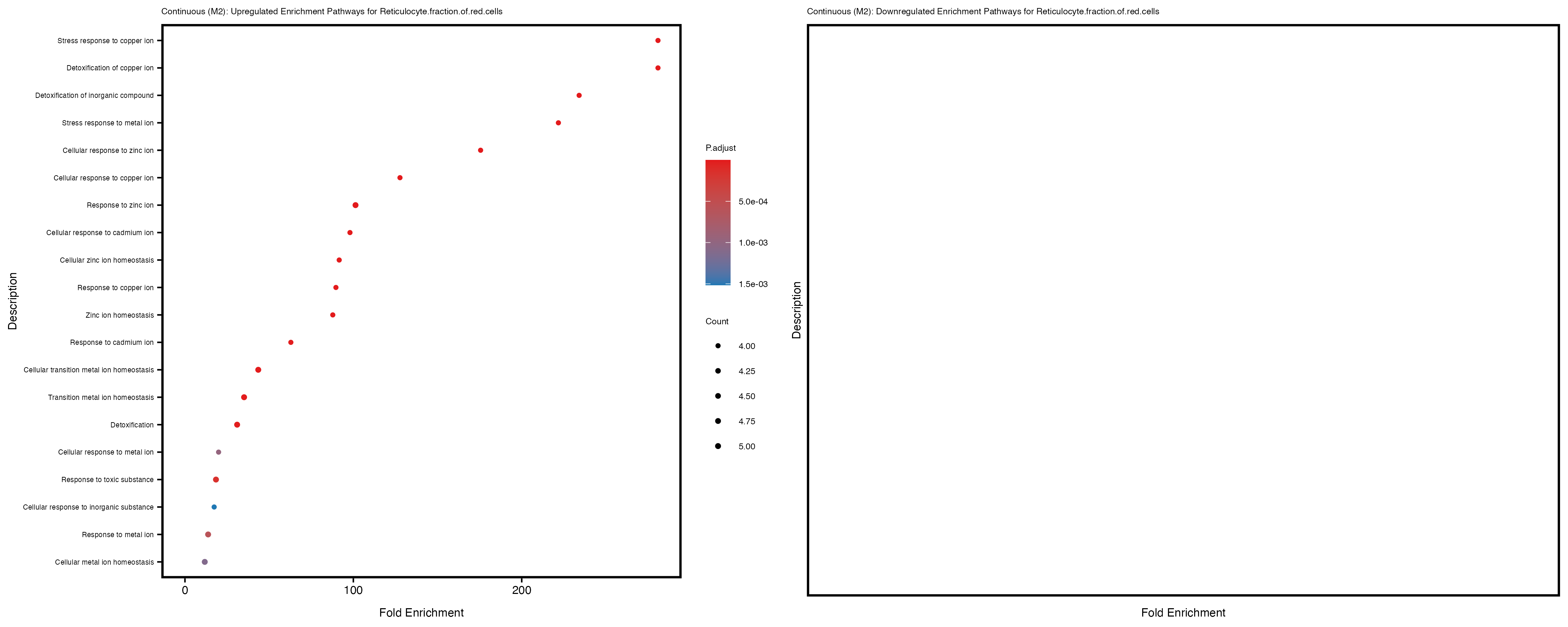
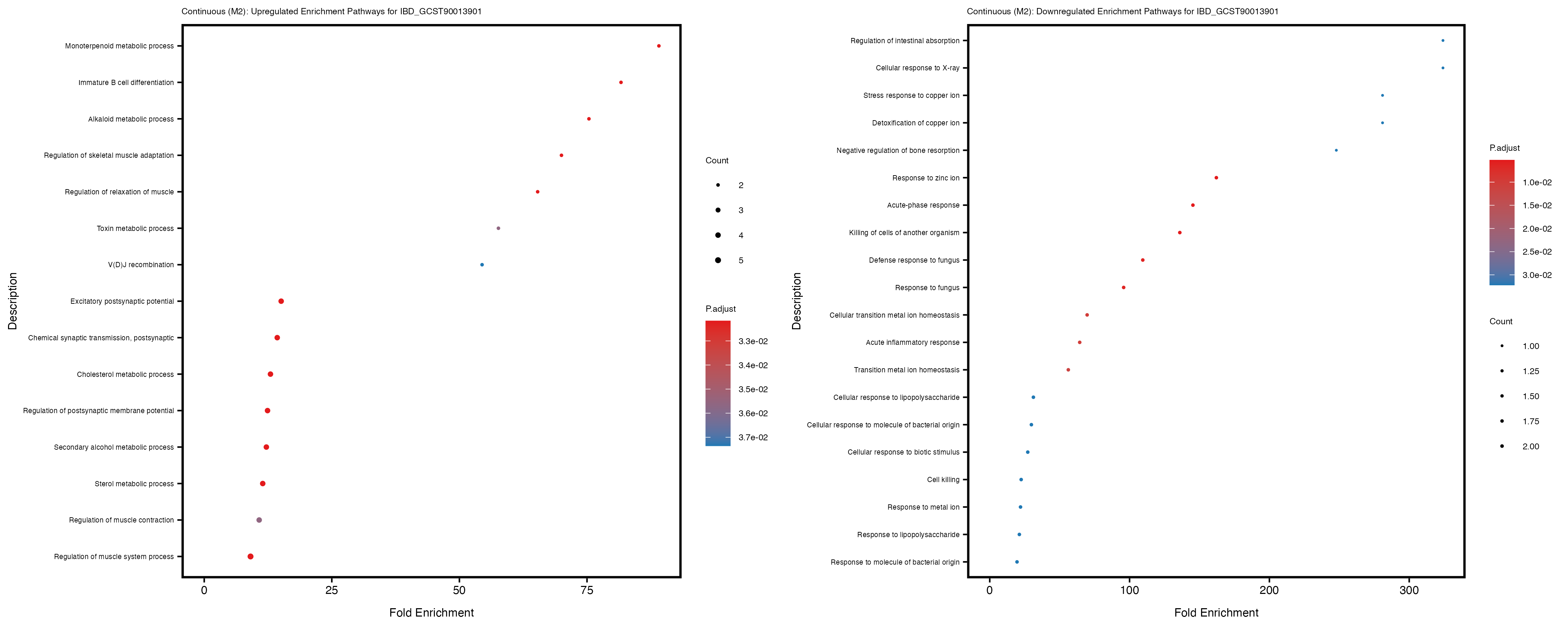
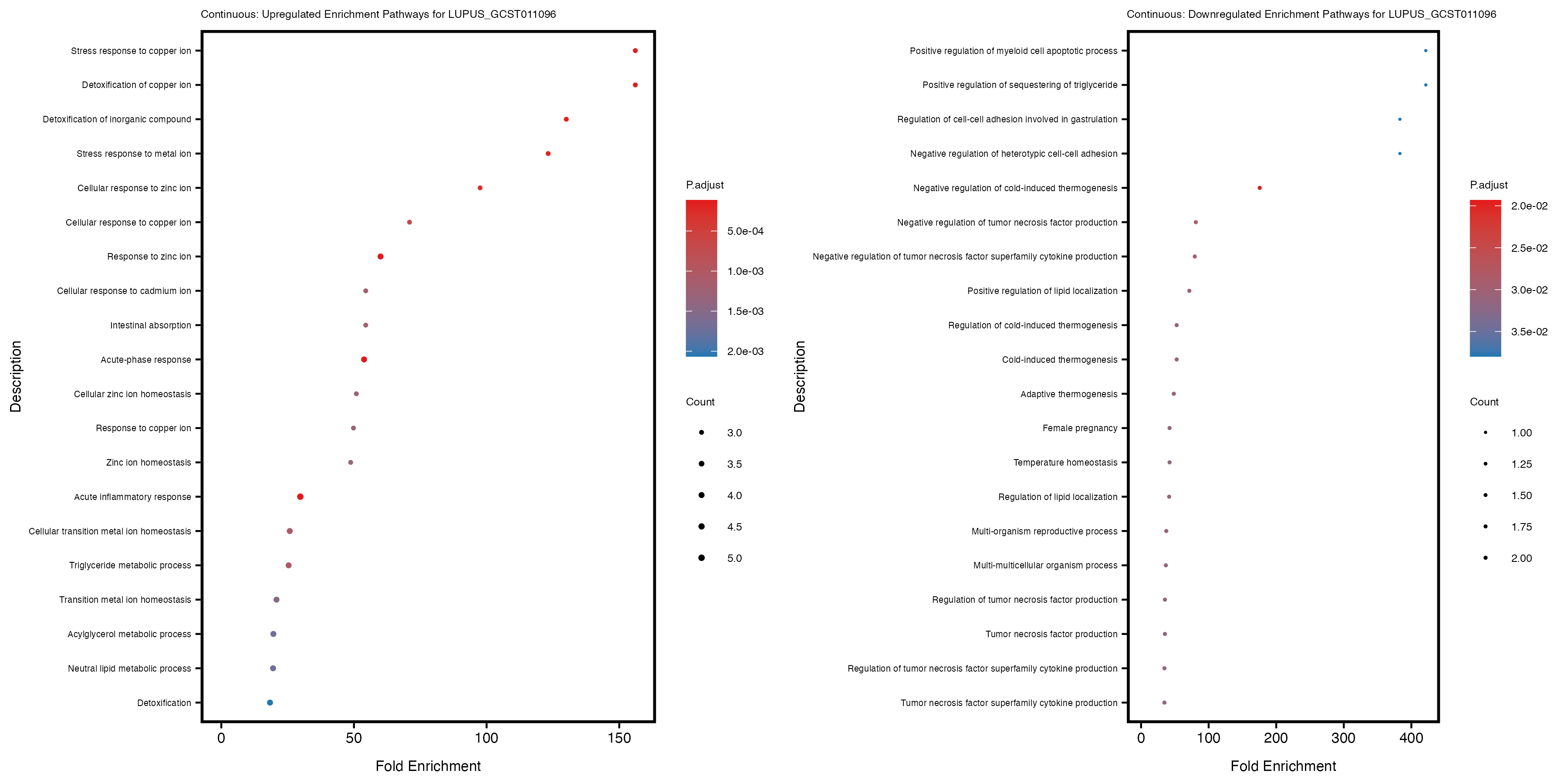
labs(title = paste("Continuous: Upregulated Enrichment Pathways for", trait)) +
theme(plot.title = element_text(size = 6), axis.text.y = element_text(size = 5))
} else {
enrich_plot_up <- plotEnrich(gse_up, plot_type = "dot", scale_ratio = 0.4) +
labs(title = paste("Continuous: Upregulated Enrichment Pathways for", trait)) +
theme(plot.title = element_text(size = 6), axis.text.y = element_text(size = 5))
}
if (nrow(gse_down) >= 20) {
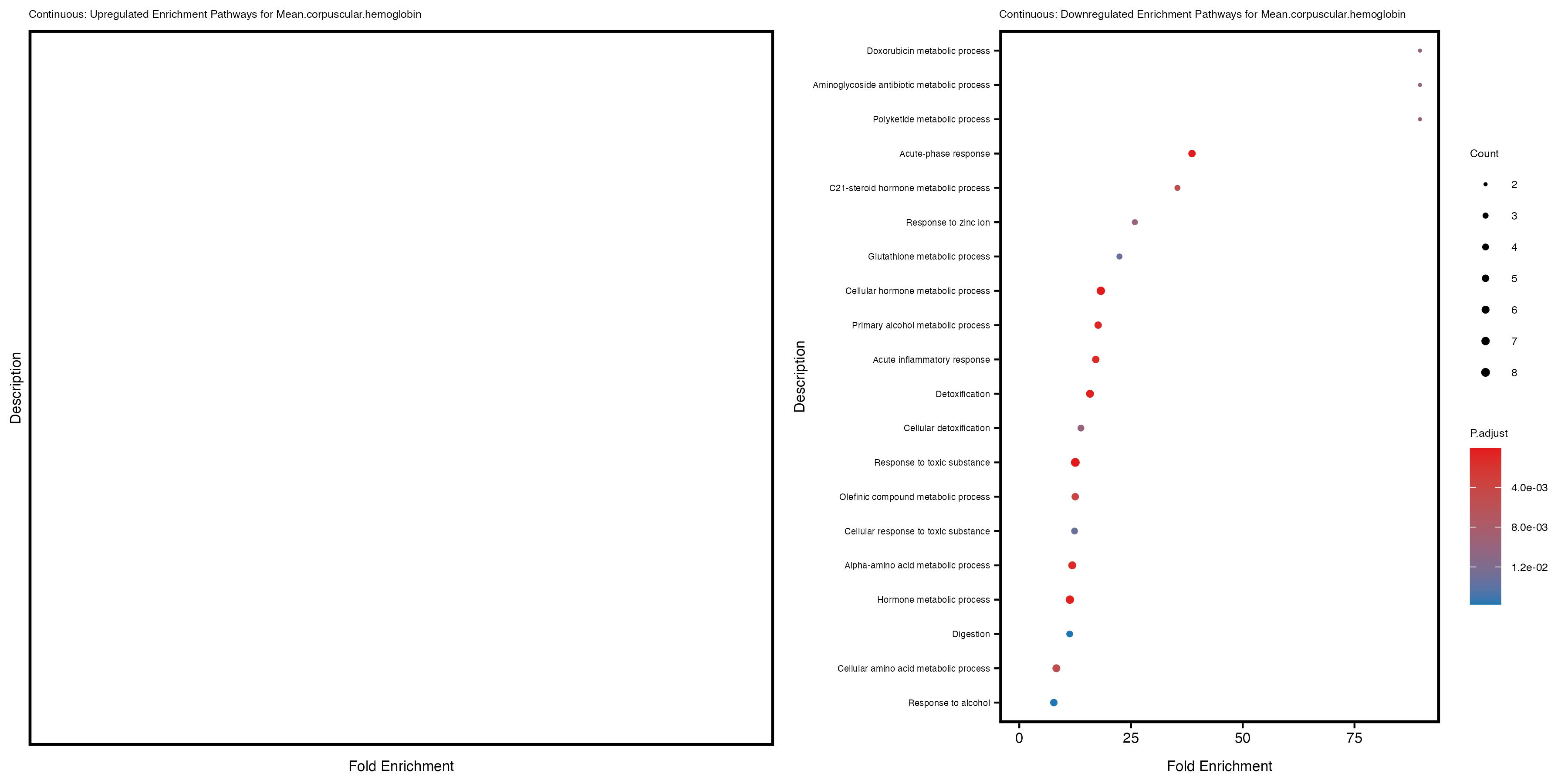
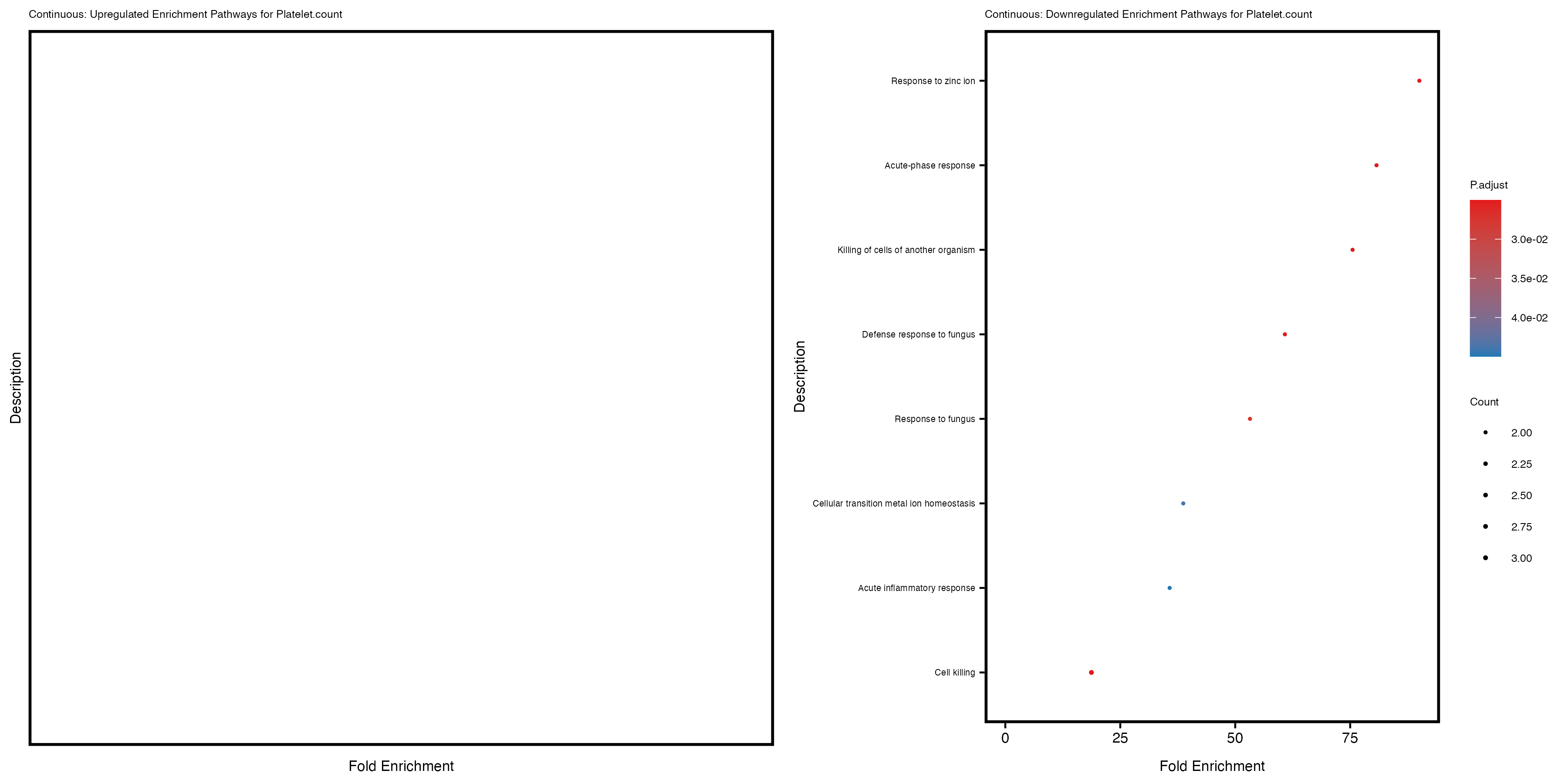
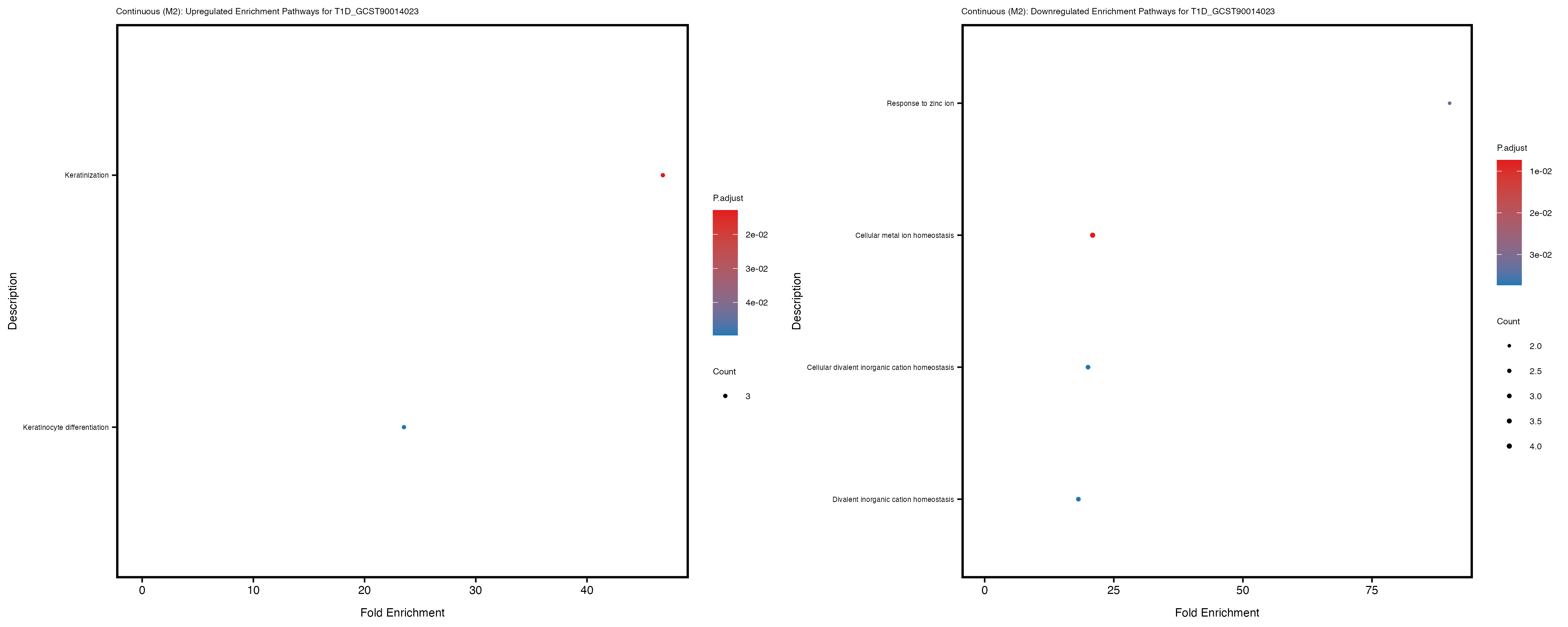
enrich_plot_down <- plotEnrich(gse_down[1:20, ], plot_type = "dot", scale_ratio = 0.4) +
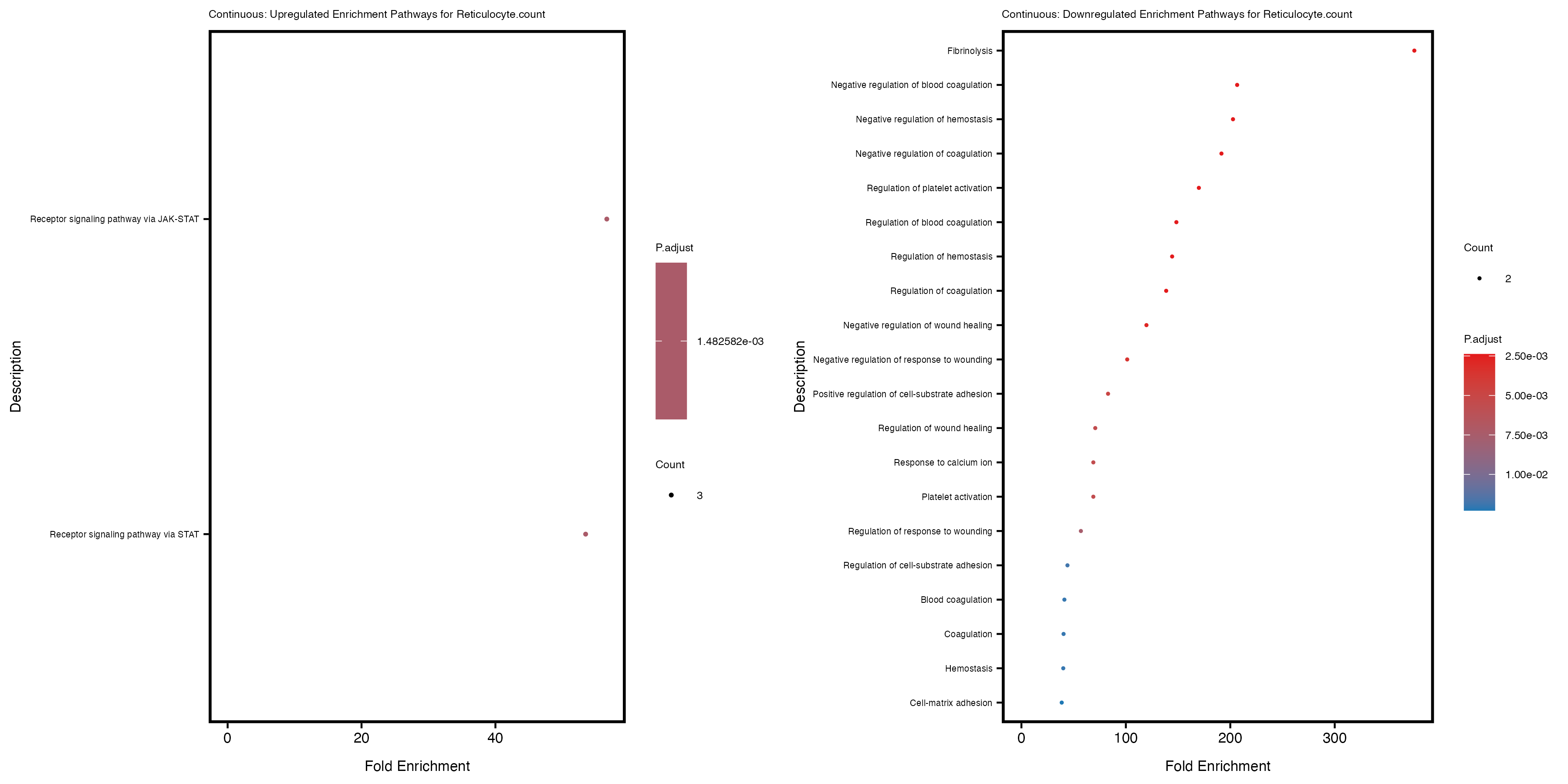
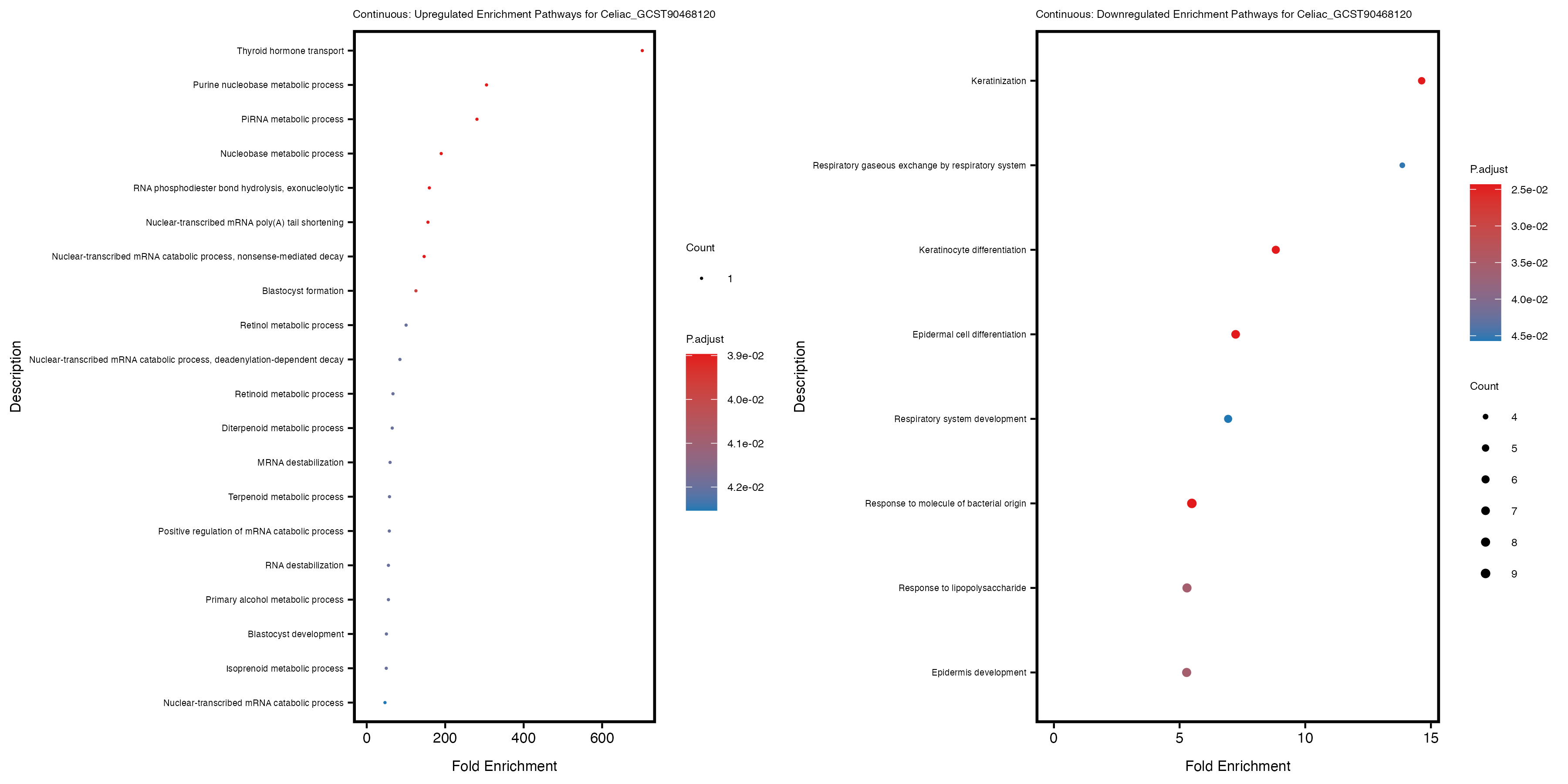
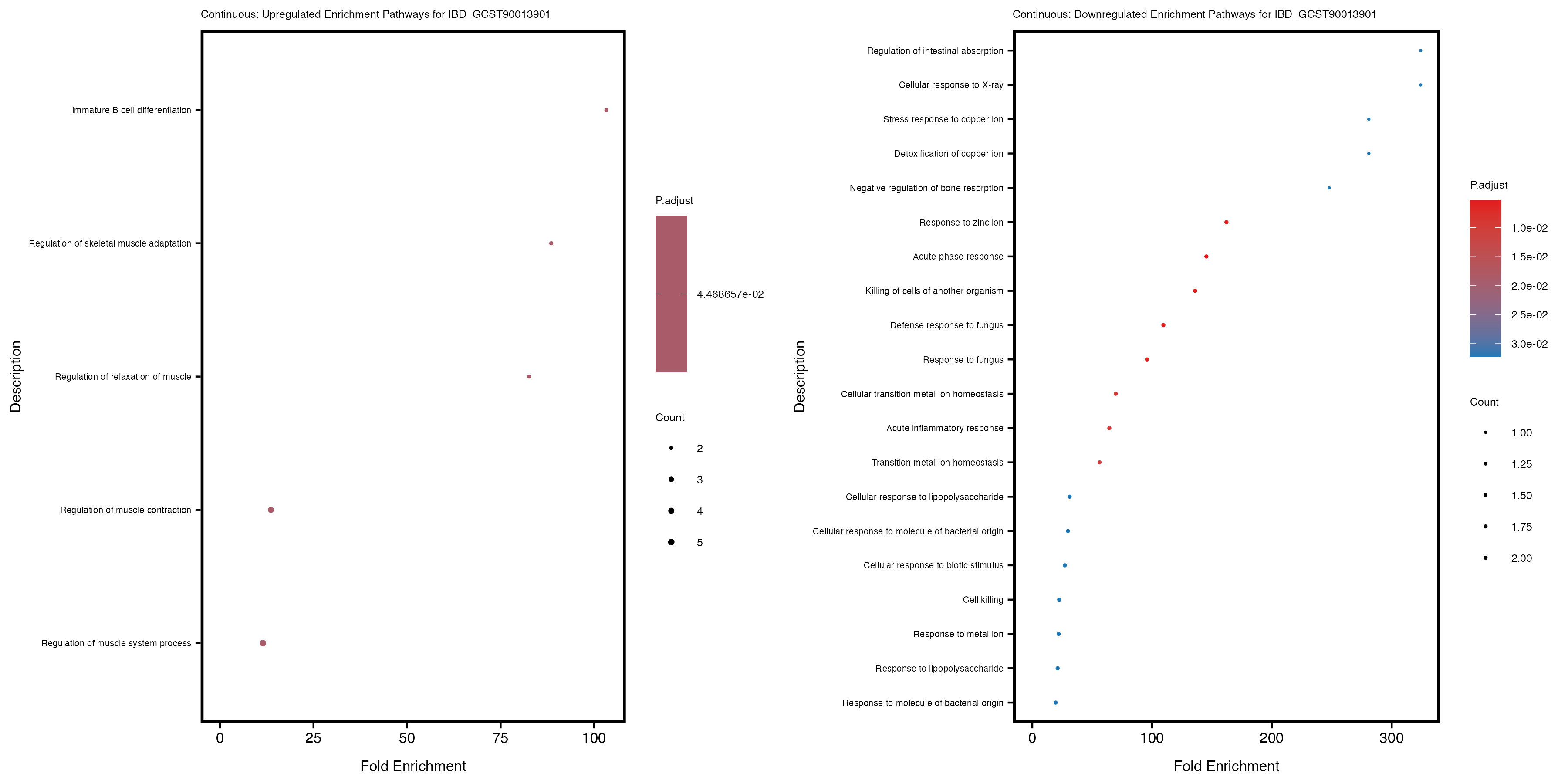
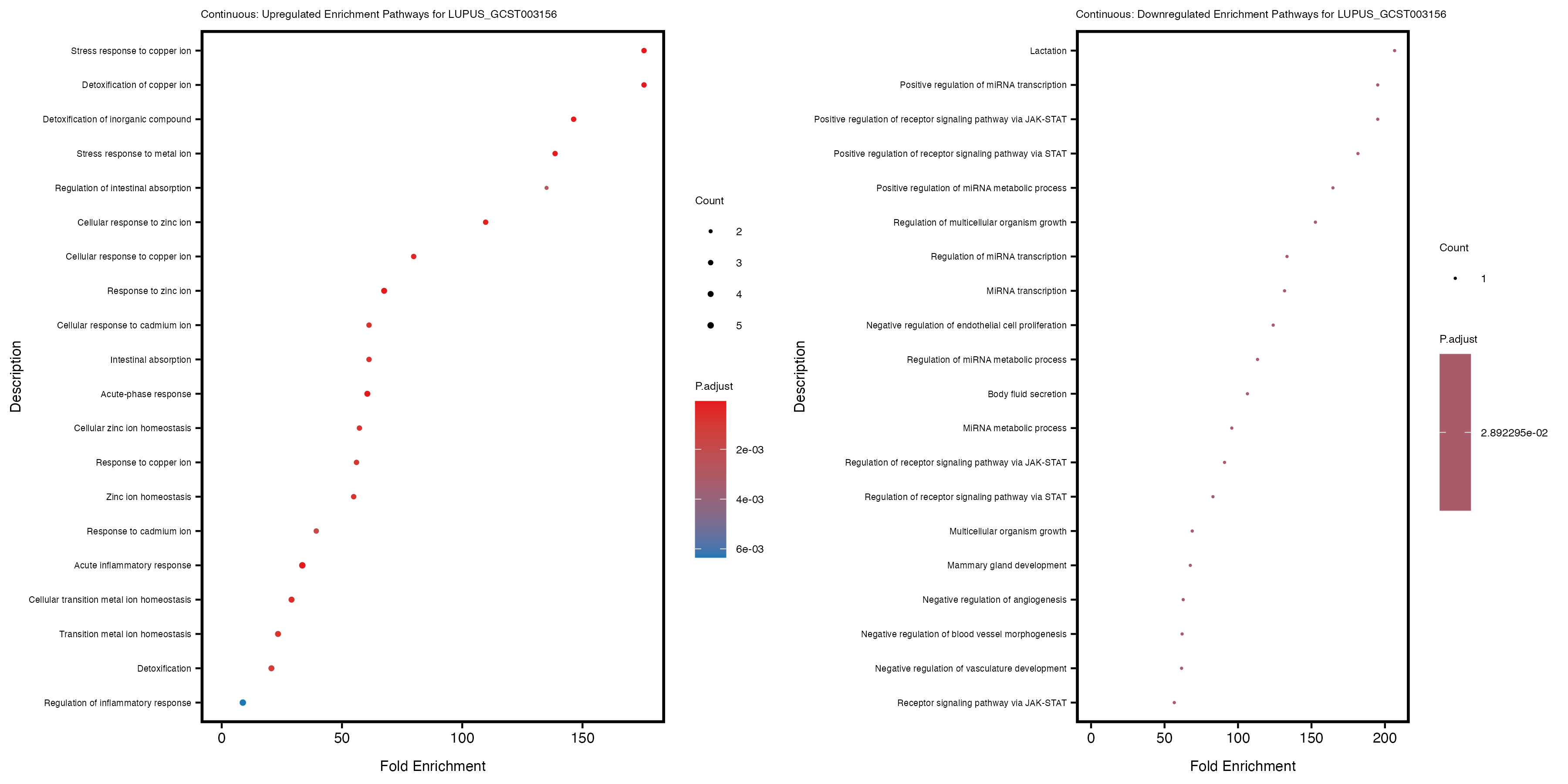
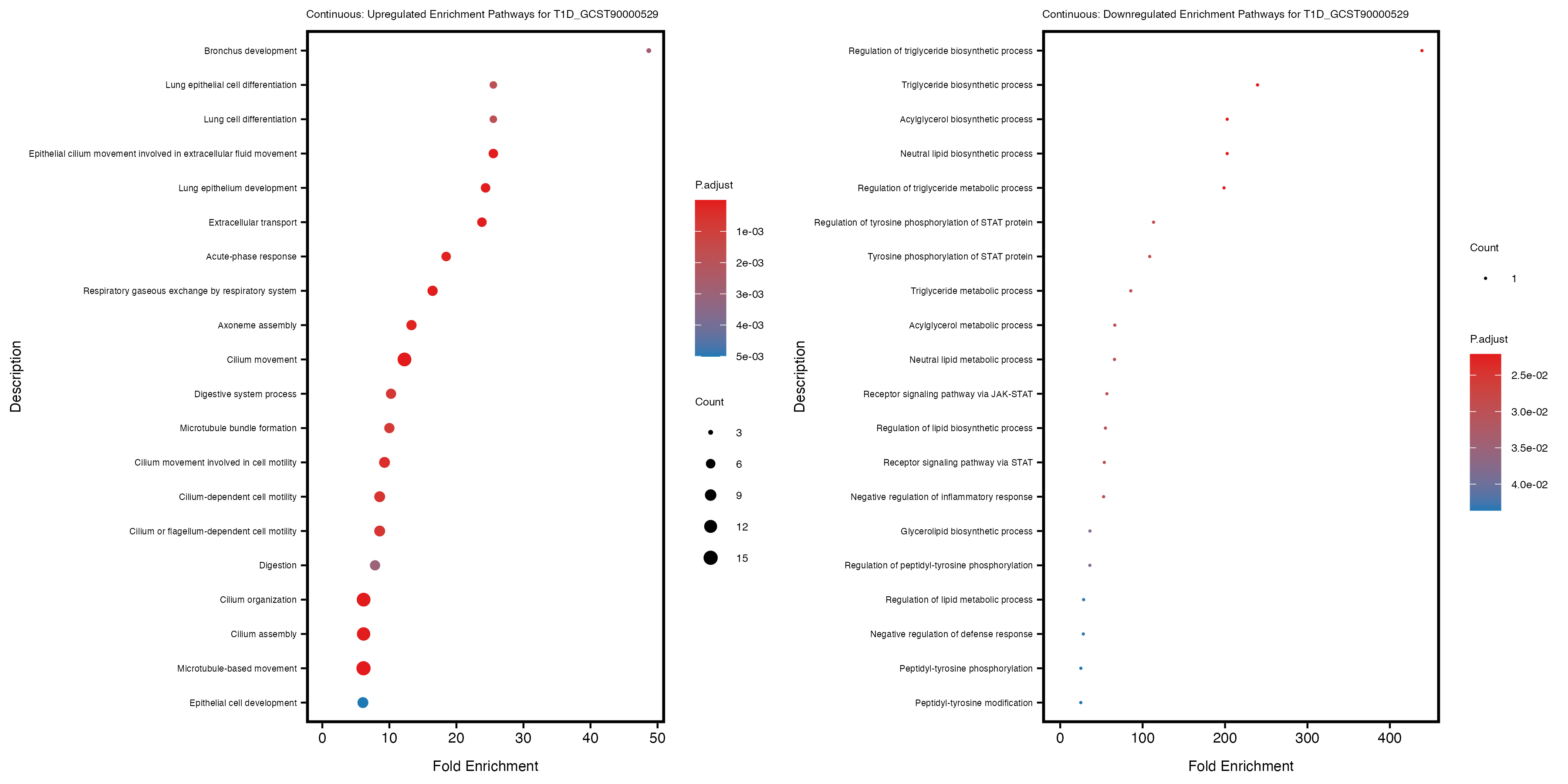
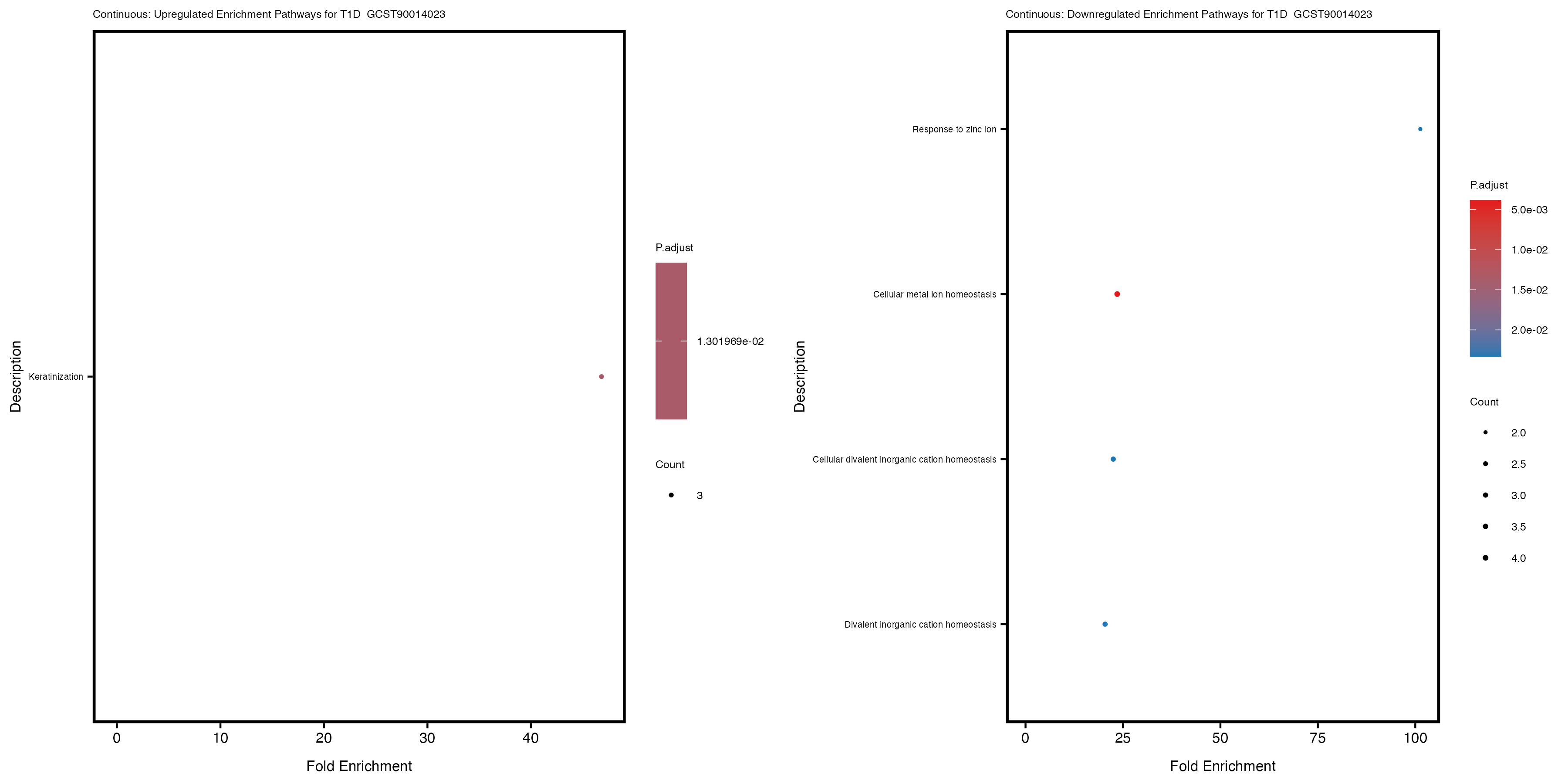
labs(title = paste("Continuous: Downregulated Enrichment Pathways for", trait)) +
theme(plot.title = element_text(size = 6), axis.text.y = element_text(size = 5))
} else {
enrich_plot_down <- plotEnrich(gse_down, plot_type = "dot", scale_ratio = 0.4) +
labs(title = paste("Continuous: Downregulated Enrichment Pathways for", trait)) +
theme(plot.title = element_text(size = 6), axis.text.y = element_text(size = 5))
}
# Arrange the two plots side by side
combined_plot <- grid.arrange(enrich_plot_up, enrich_plot_down, ncol = 2)
# Save the combined plot
ggsave(paste0("enrichment_plot_", trait, ".png"), plot = combined_plot,
width = 15, height = 6)
# Save the GO enrichment results to CSV
write.csv(gse_up, file = paste0("GO_enrichment_", trait, "_upregulated.csv"))
write.csv(gse_down, file = paste0("GO_enrichment_", trait, "_downregulated.csv"))
}Quantile PRS:
dir_path <- "analysis/quantile_wb"
files <- list.files(dir_path, pattern = "differential_expression_.*_quantile_results.csv",
full.names = TRUE)
for (file in files) {
trait <- gsub("differential_expression_(.*)_quantile_results.csv", "\\1", basename(file))
trait
res_tableOE <- read.csv(file, header = T, row.names = 1)
deGenes <- res_tableOE[res_tableOE$padj < 0.1 &
abs(res_tableOE$log2FoldChange) >= 0.5, ]
deGenes$gene_id <- gsub("\\.\\d+$", "", rownames(deGenes))
# Separate upregulated and downregulated genes
upregulated_genes <- deGenes[deGenes$log2FoldChange > 0, ]$gene_id
downregulated_genes <- deGenes[deGenes$log2FoldChange < 0, ]$gene_id
# Run GO enrichment for upregulated genes
gse_up <- enrichGO(gene = upregulated_genes, ont = "BP",
OrgDb = "org.Hs.eg.db", keyType = "ENSEMBL", readable = T)
# Run GO enrichment for downregulated genes
gse_down <- enrichGO(gene = downregulated_genes, ont = "BP",
OrgDb = "org.Hs.eg.db", keyType = "ENSEMBL", readable = T)
# Convert enrichment results to data frames and calculate additional ratios
if (is.null(gse_up)) {
gse_up <- data.frame(ID = character(), Description = character(),
GeneRatio = character(), BgRatio = character(),
pvalue = numeric(), p.adjust = numeric(),
qvalue = numeric(), geneID = character(),
Count = integer(), stringsAsFactors = FALSE)
} else {
gse_up <- as.data.frame(gse_up)}
gse_down <- as.data.frame(gse_down)
gse_up$GeneRatio_num <- as.numeric(sapply(strsplit(gse_up$GeneRatio, "/"), function(x) x[1])) /
as.numeric(sapply(strsplit(gse_up$GeneRatio, "/"), function(x) x[2]))
gse_up$BgRatio_num <- as.numeric(sapply(strsplit(gse_up$BgRatio, "/"), function(x) x[1])) /
as.numeric(sapply(strsplit(gse_up$BgRatio, "/"), function(x) x[2]))
gse_up <- cbind(gse_up, FoldEnrich = gse_up$GeneRatio_num / gse_up$BgRatio_num)
gse_down$GeneRatio_num <- as.numeric(sapply(strsplit(gse_down$GeneRatio, "/"), function(x) x[1])) /
as.numeric(sapply(strsplit(gse_down$GeneRatio, "/"), function(x) x[2]))
gse_down$BgRatio_num <- as.numeric(sapply(strsplit(gse_down$BgRatio, "/"), function(x) x[1])) /
as.numeric(sapply(strsplit(gse_down$BgRatio, "/"), function(x) x[2]))
gse_down <- cbind(gse_down, FoldEnrich = gse_down$GeneRatio_num / gse_down$BgRatio_num)
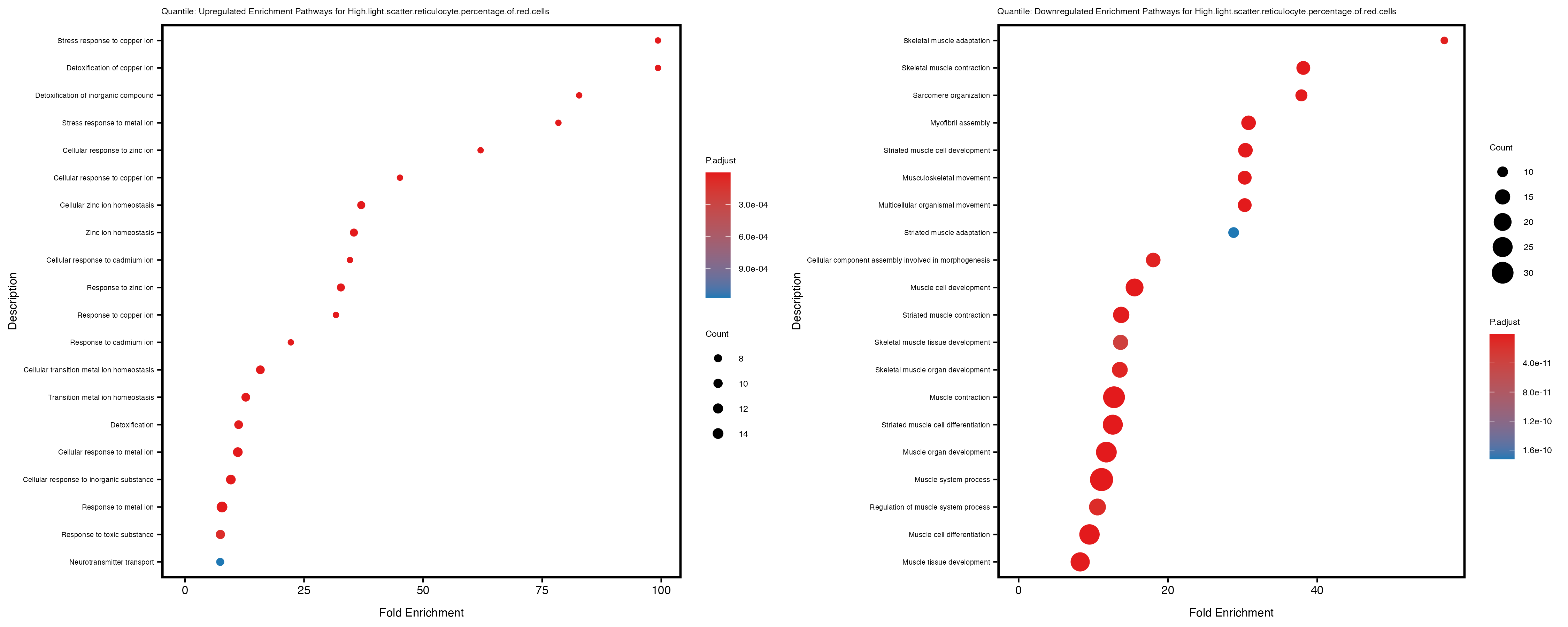
if (nrow(gse_up) >= 20) {
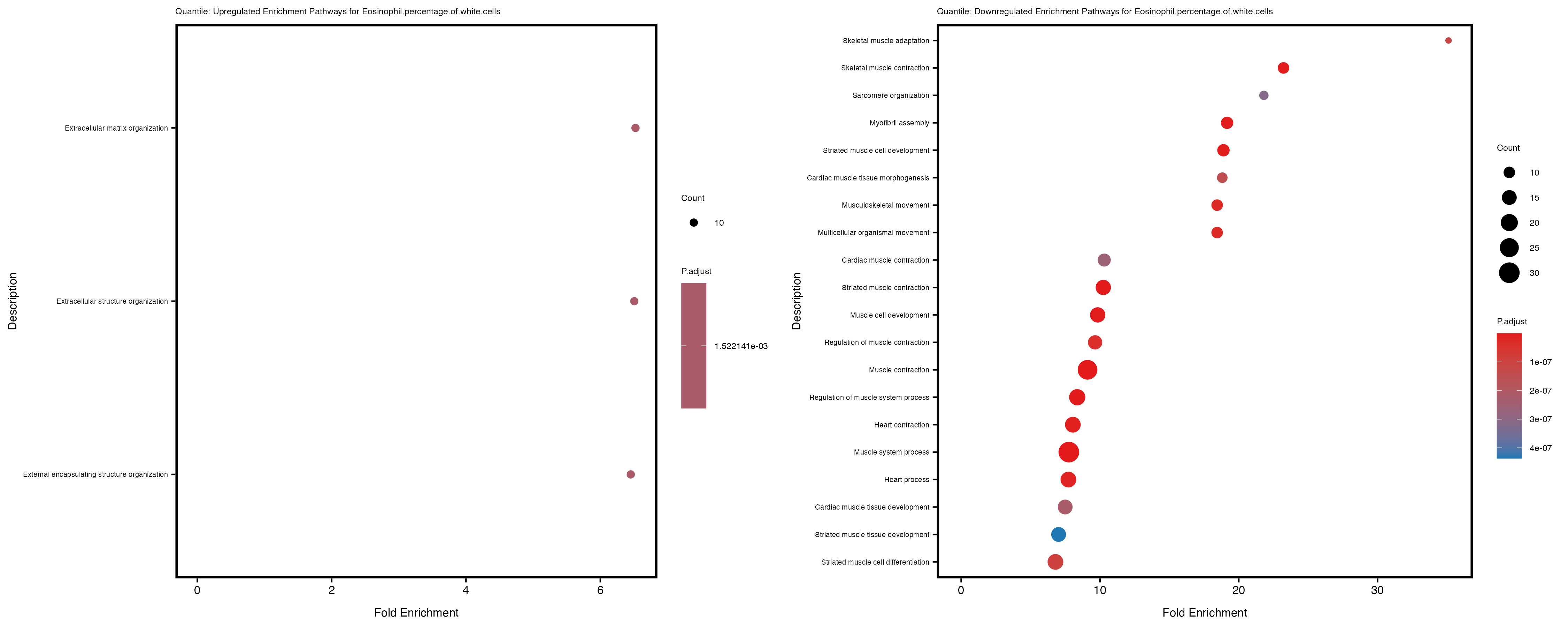
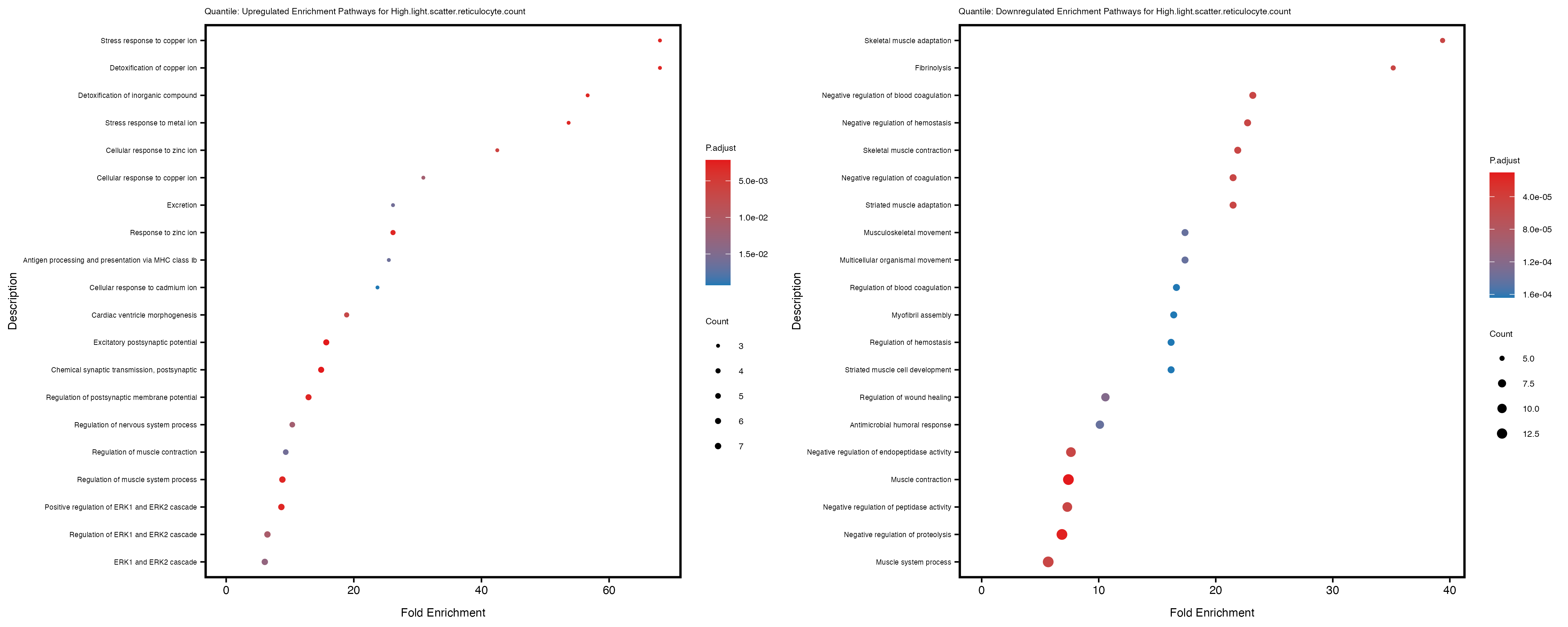
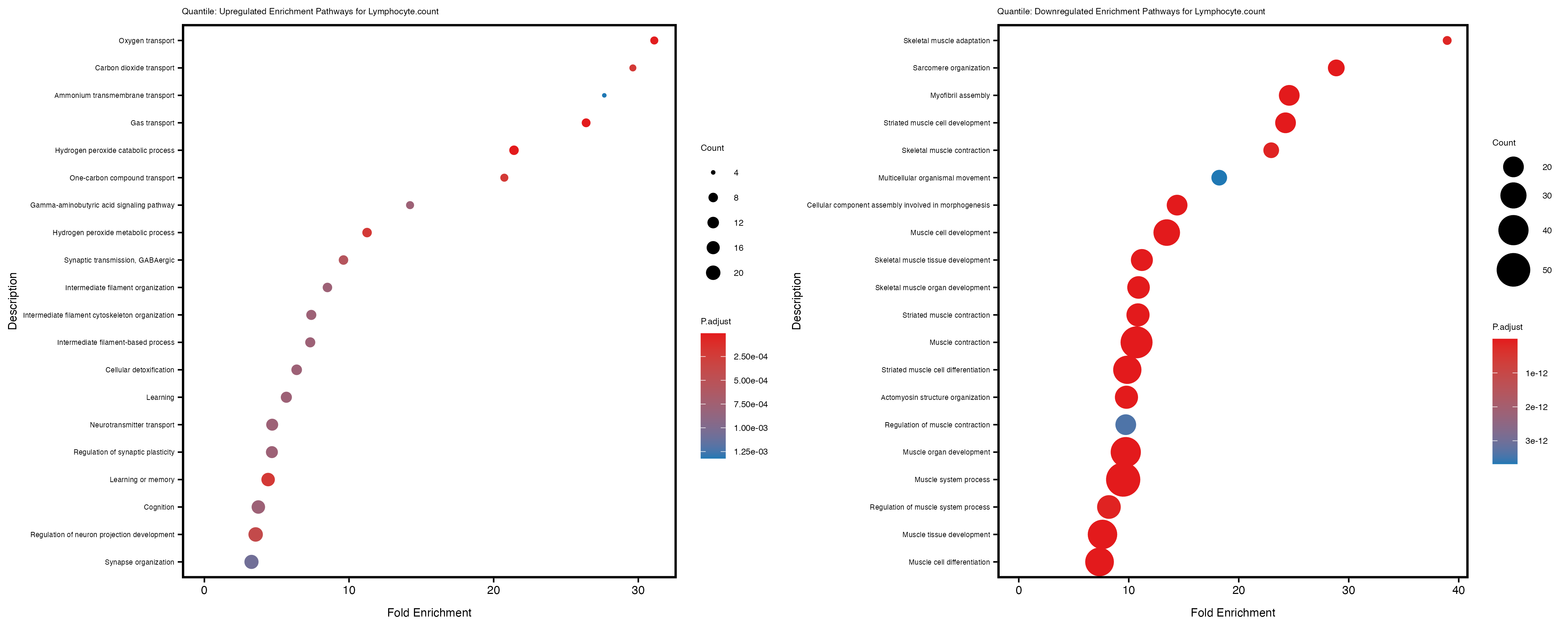
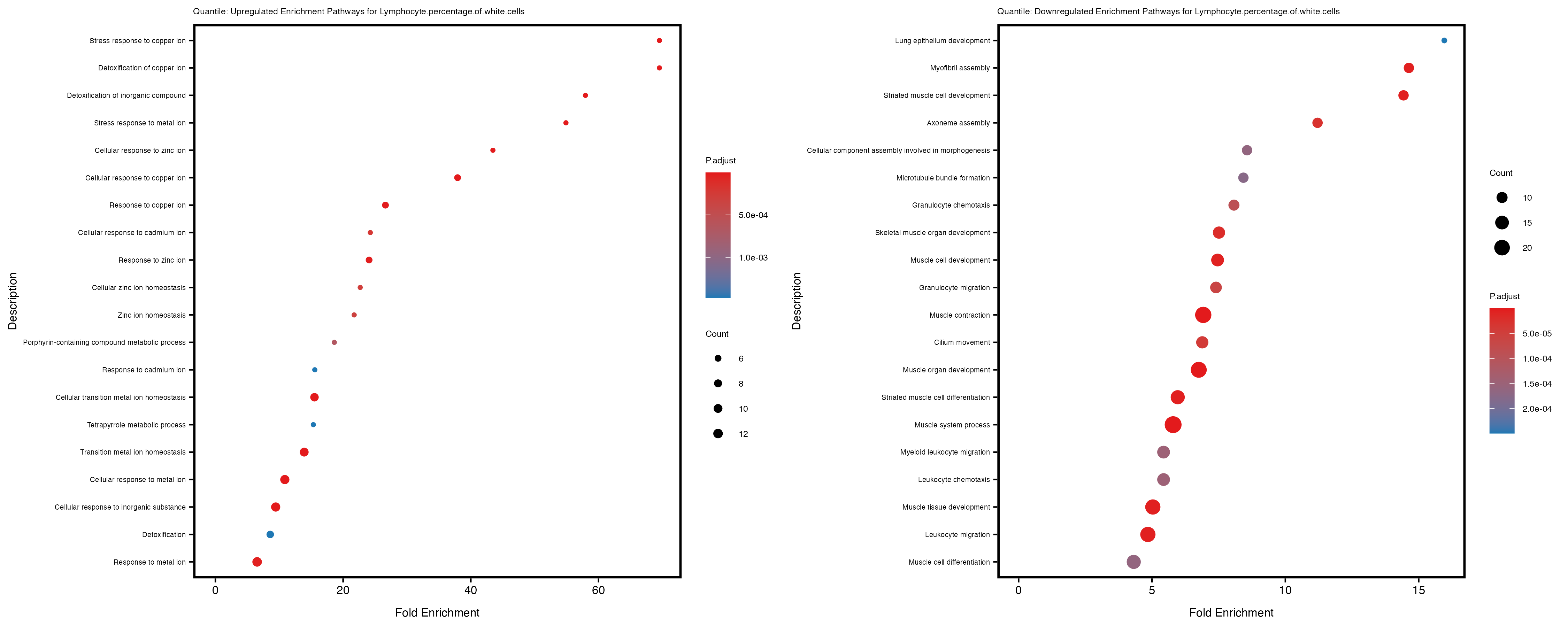
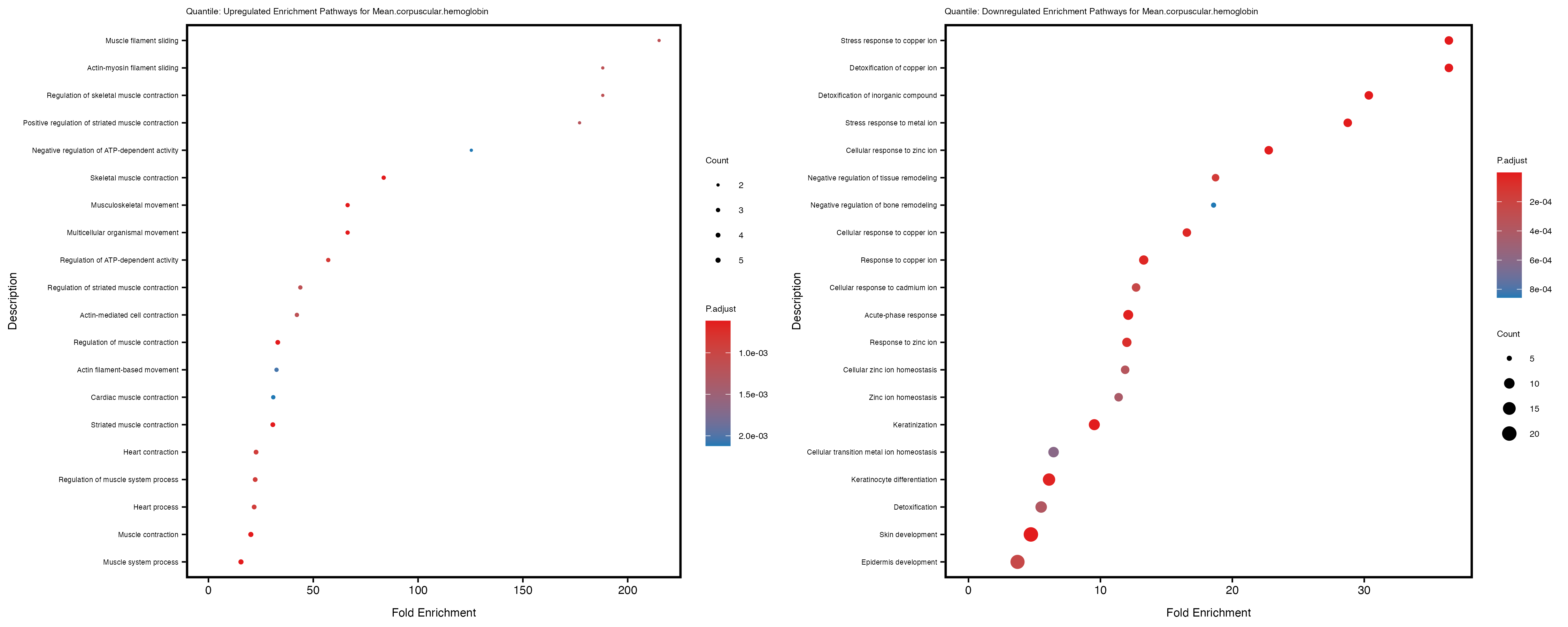
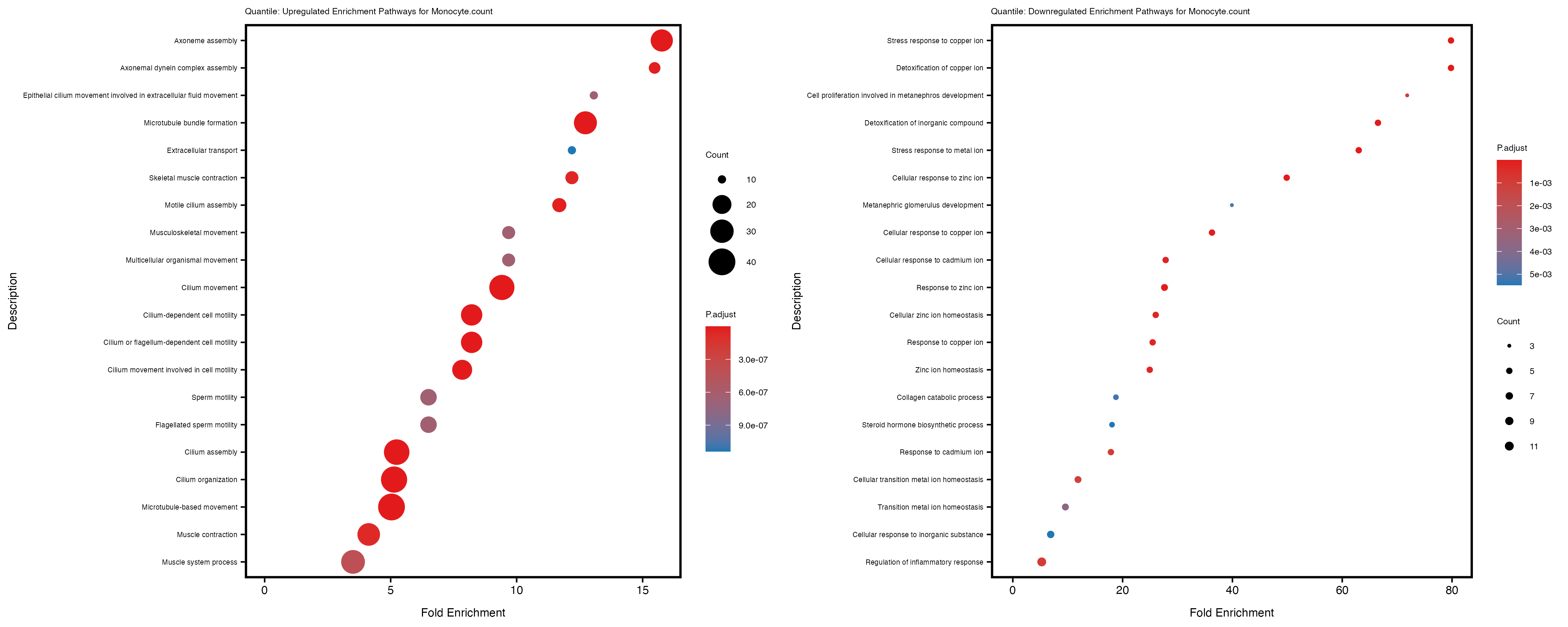
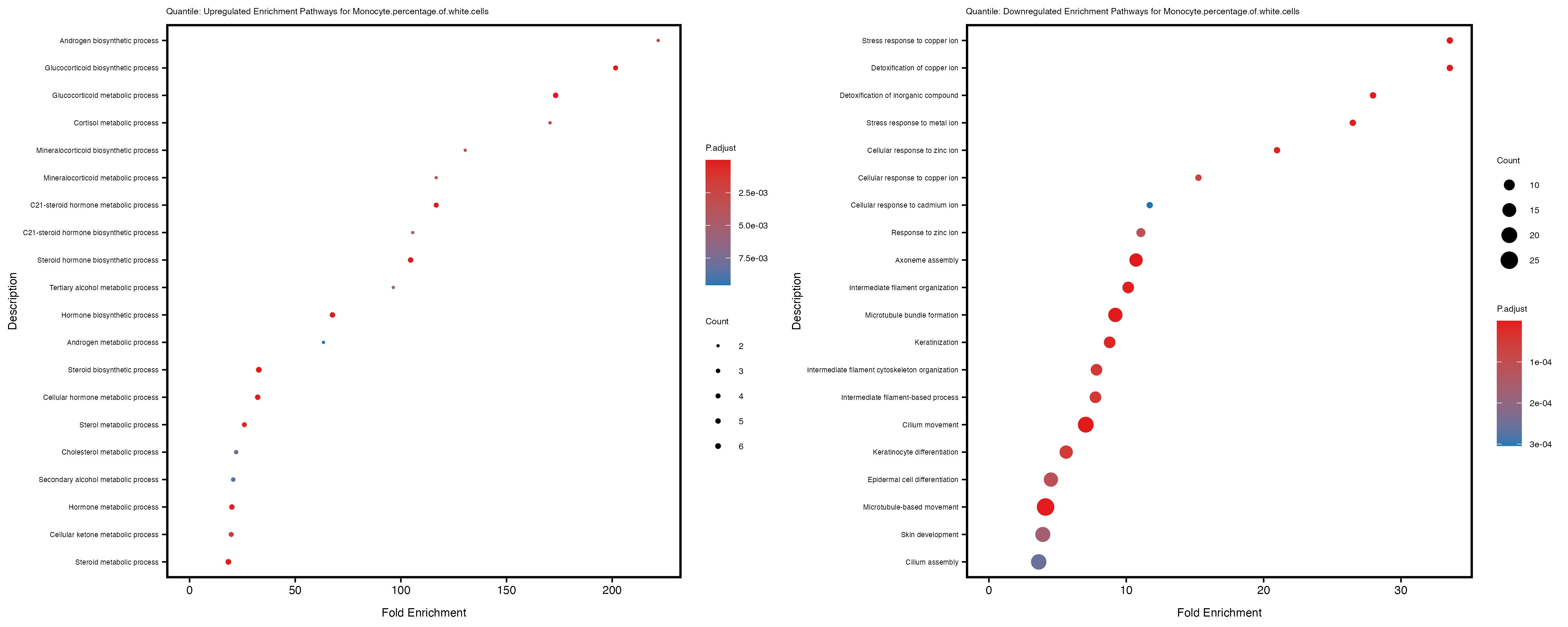
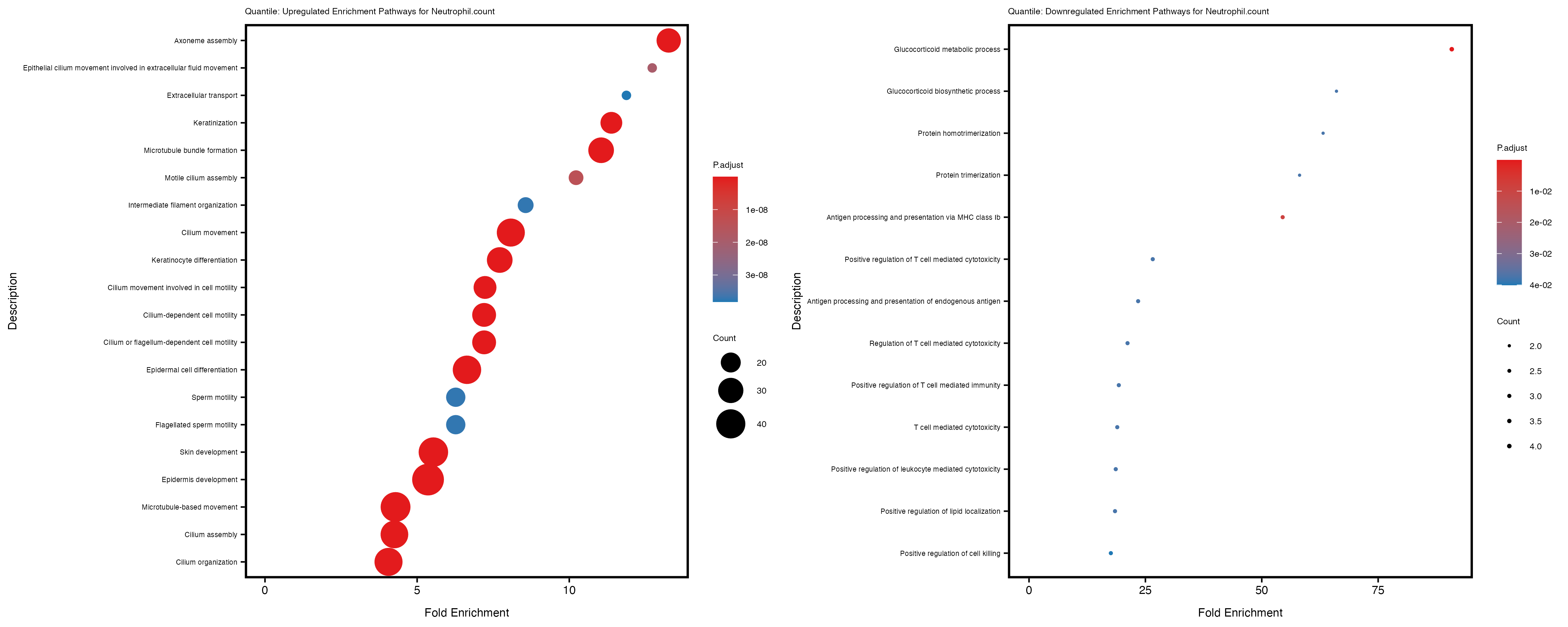
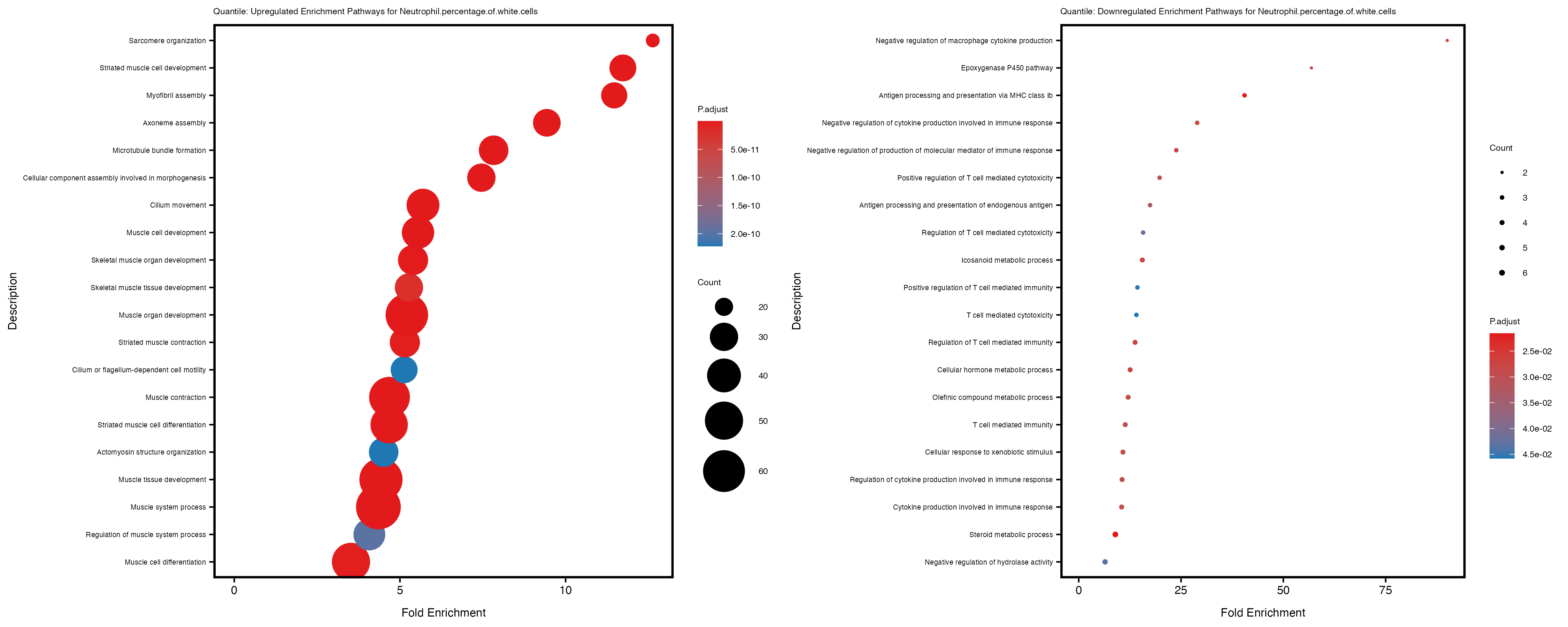
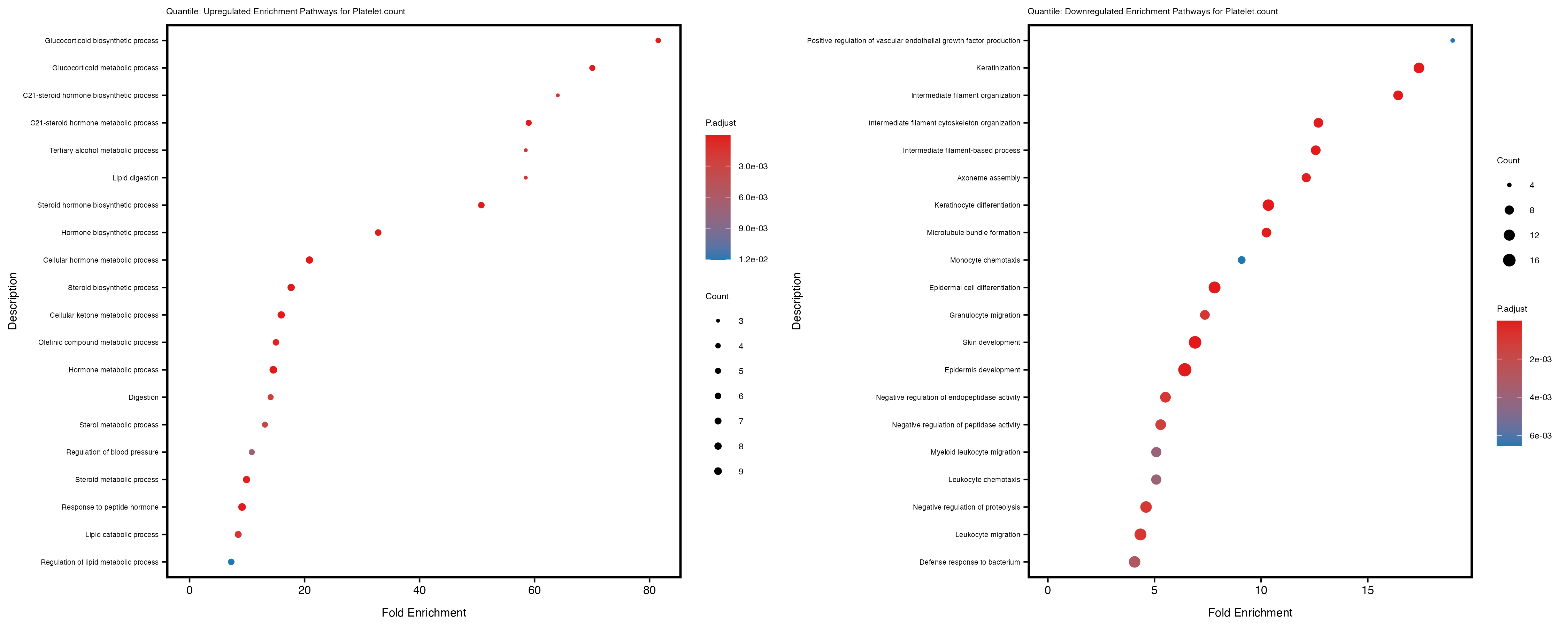
enrich_plot_up <- plotEnrich(gse_up[1:20, ], plot_type = "dot", scale_ratio = 0.4) +
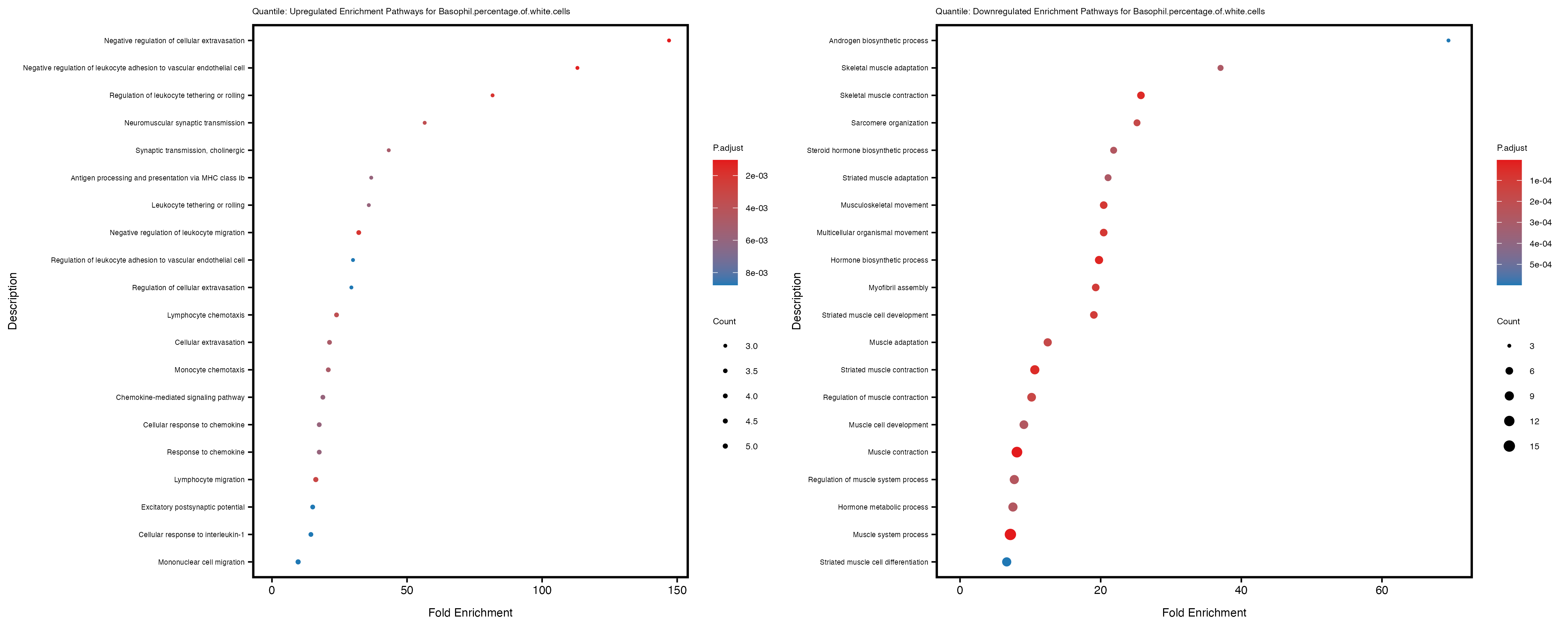
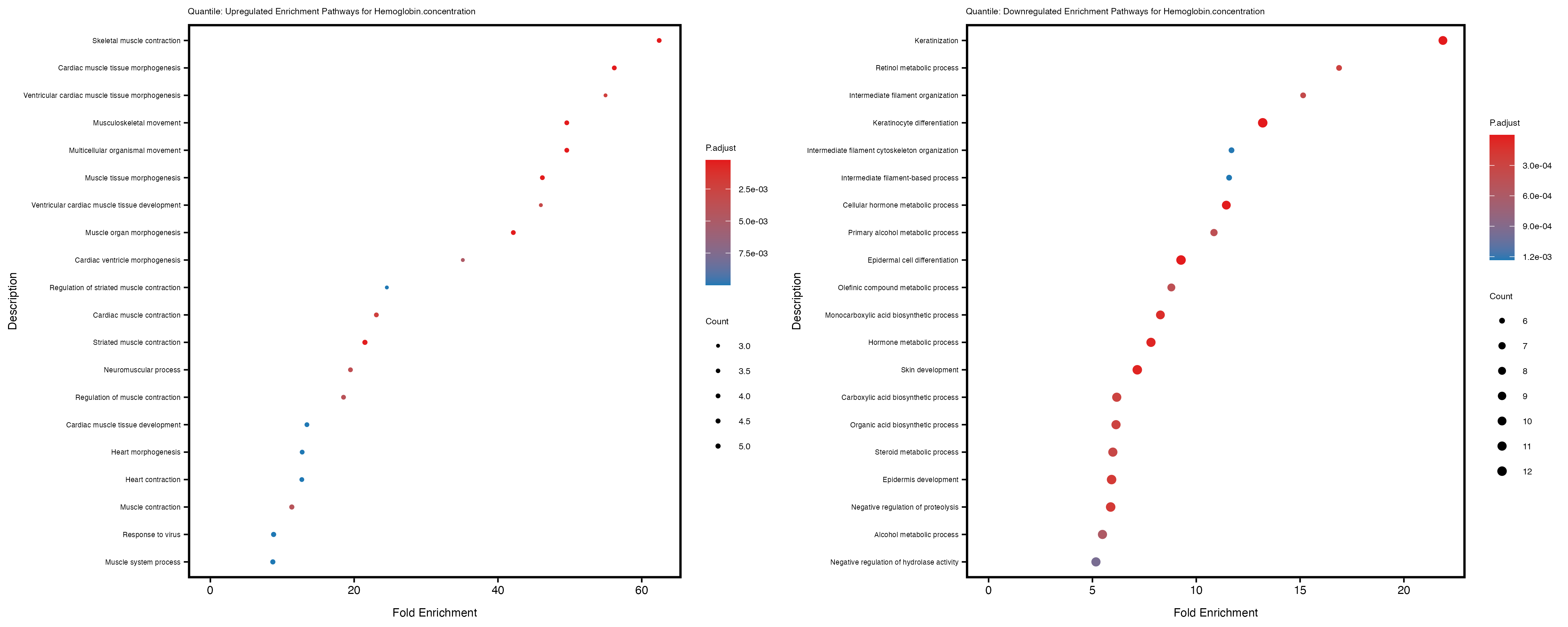
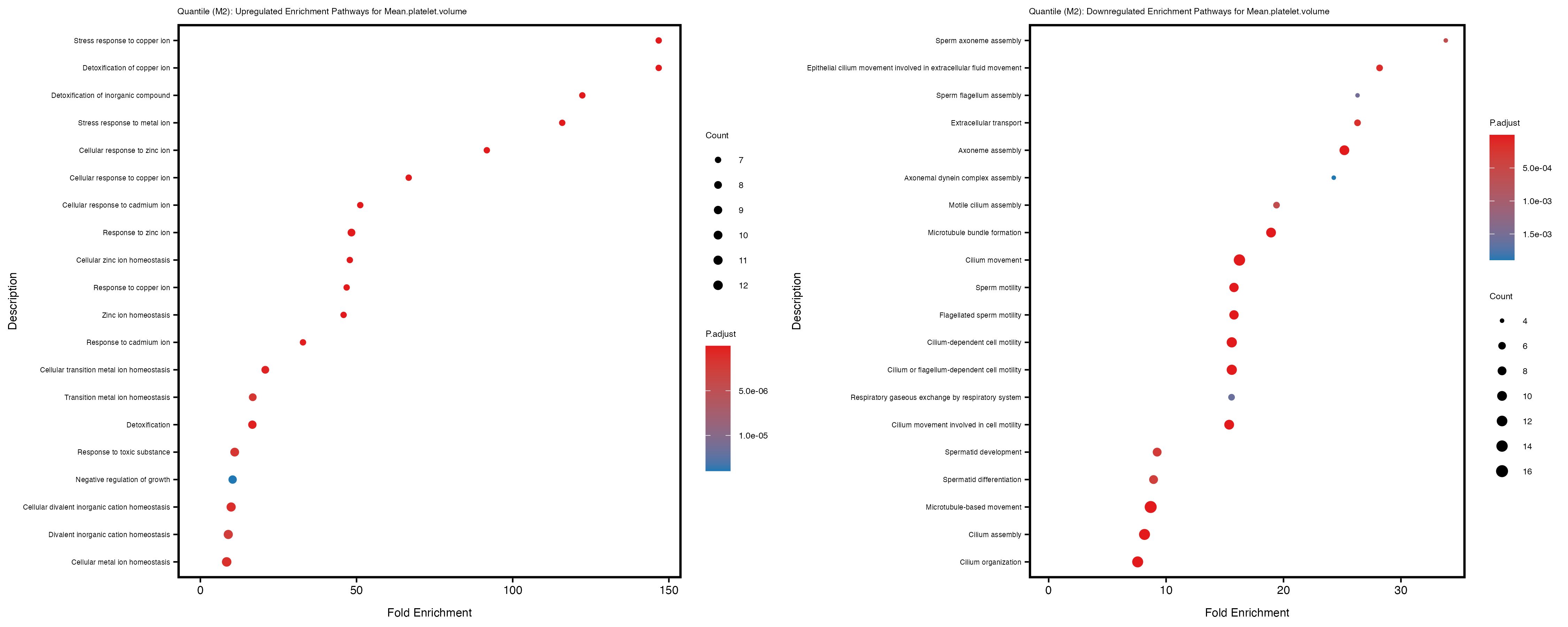
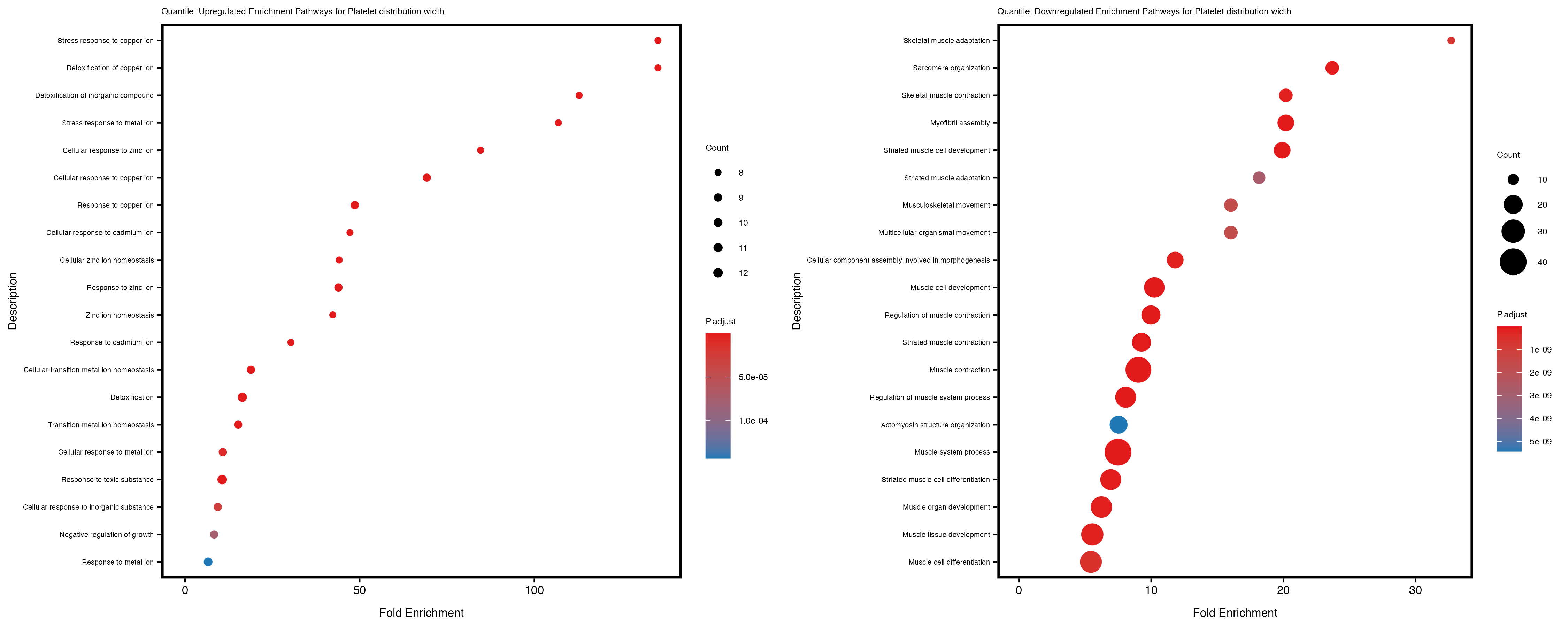
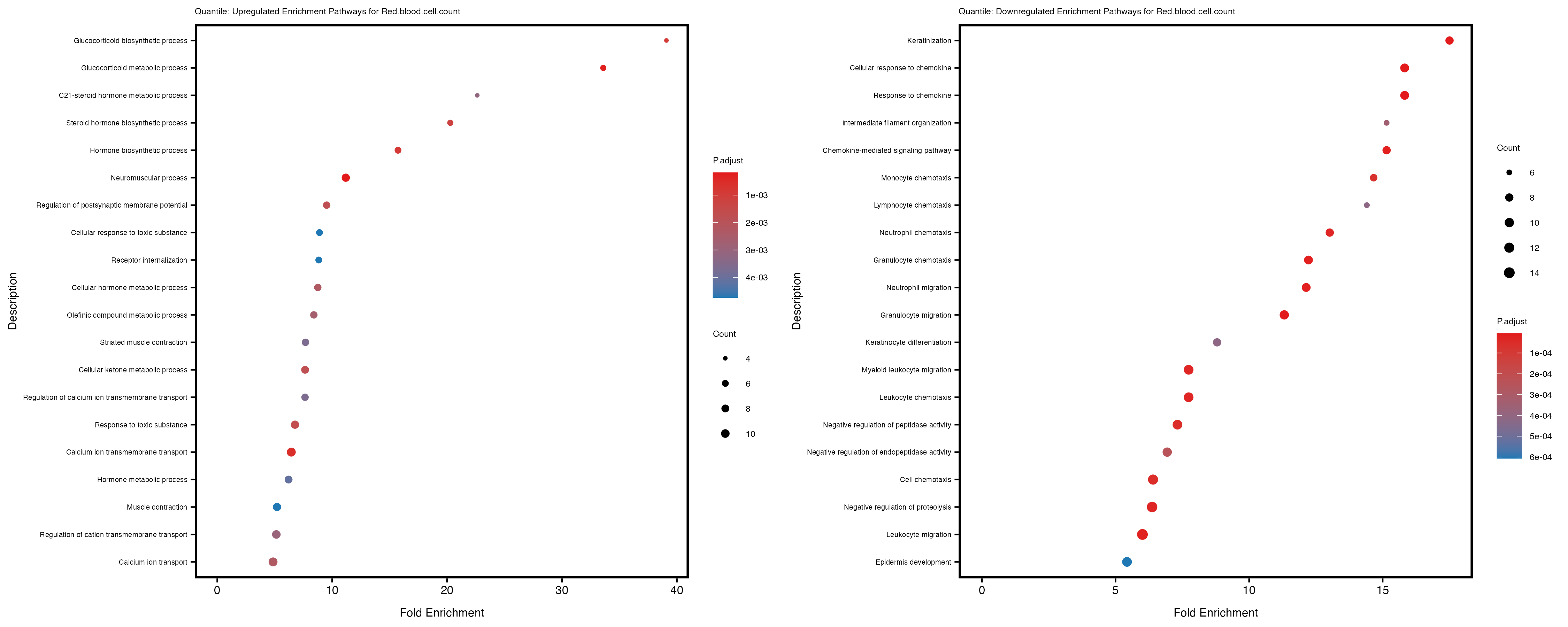
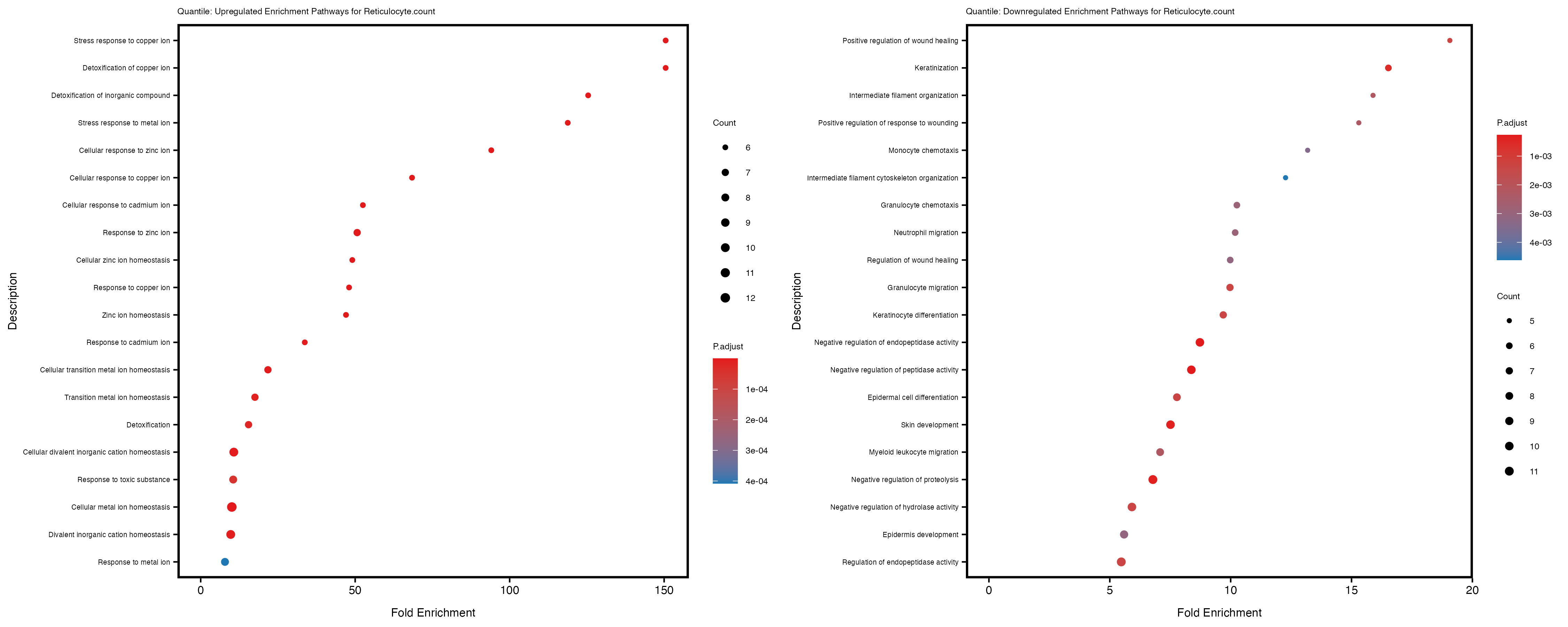
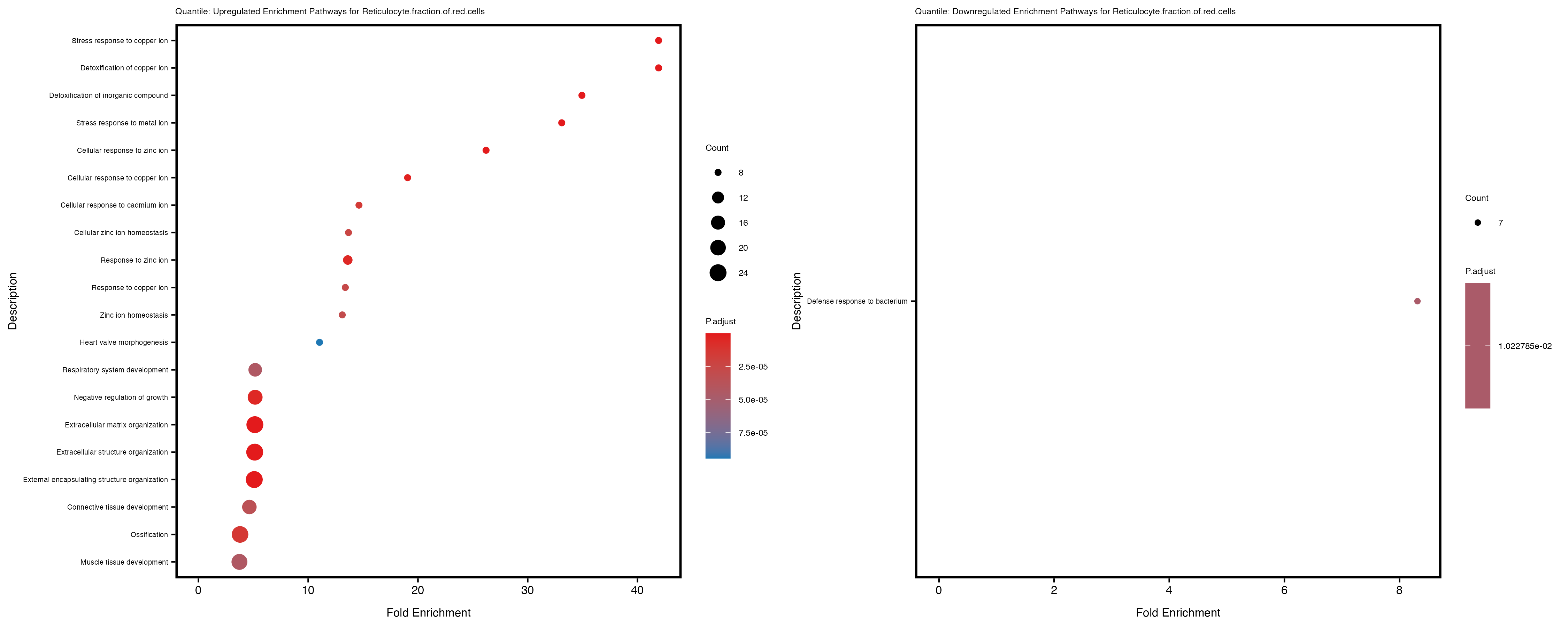
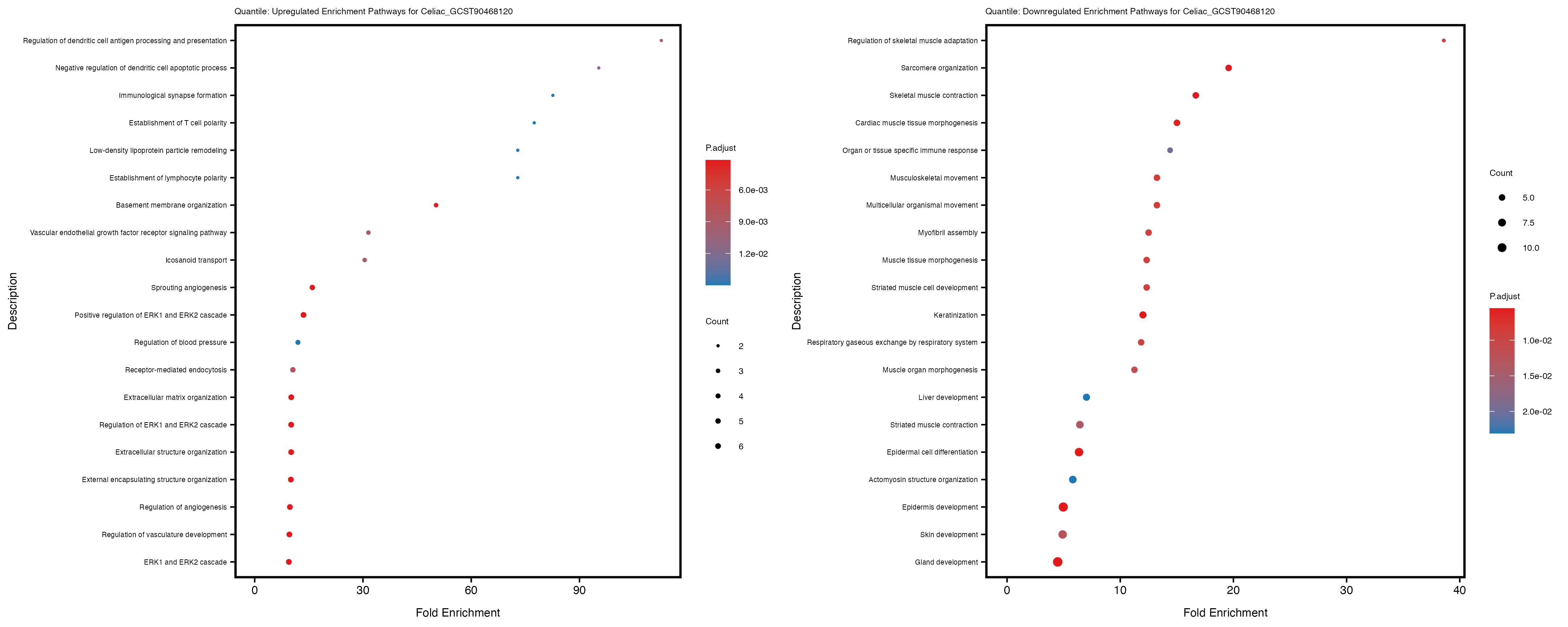
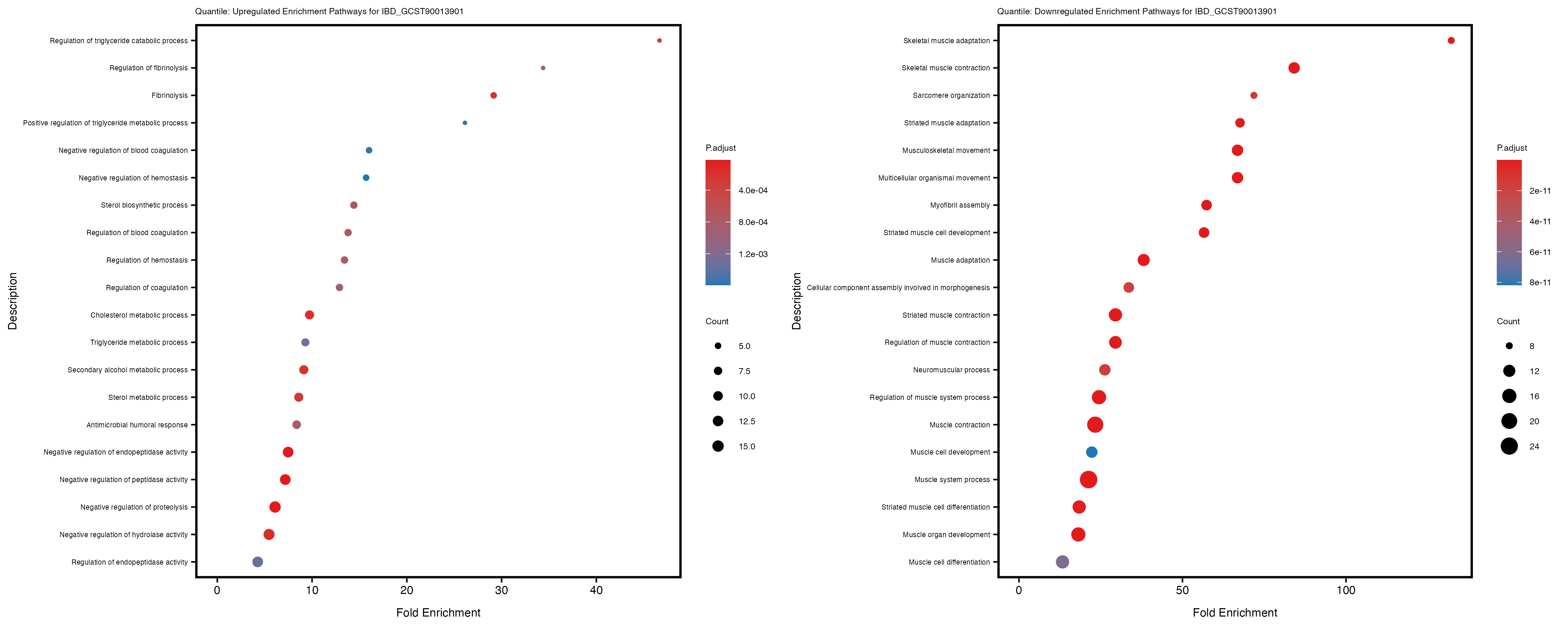
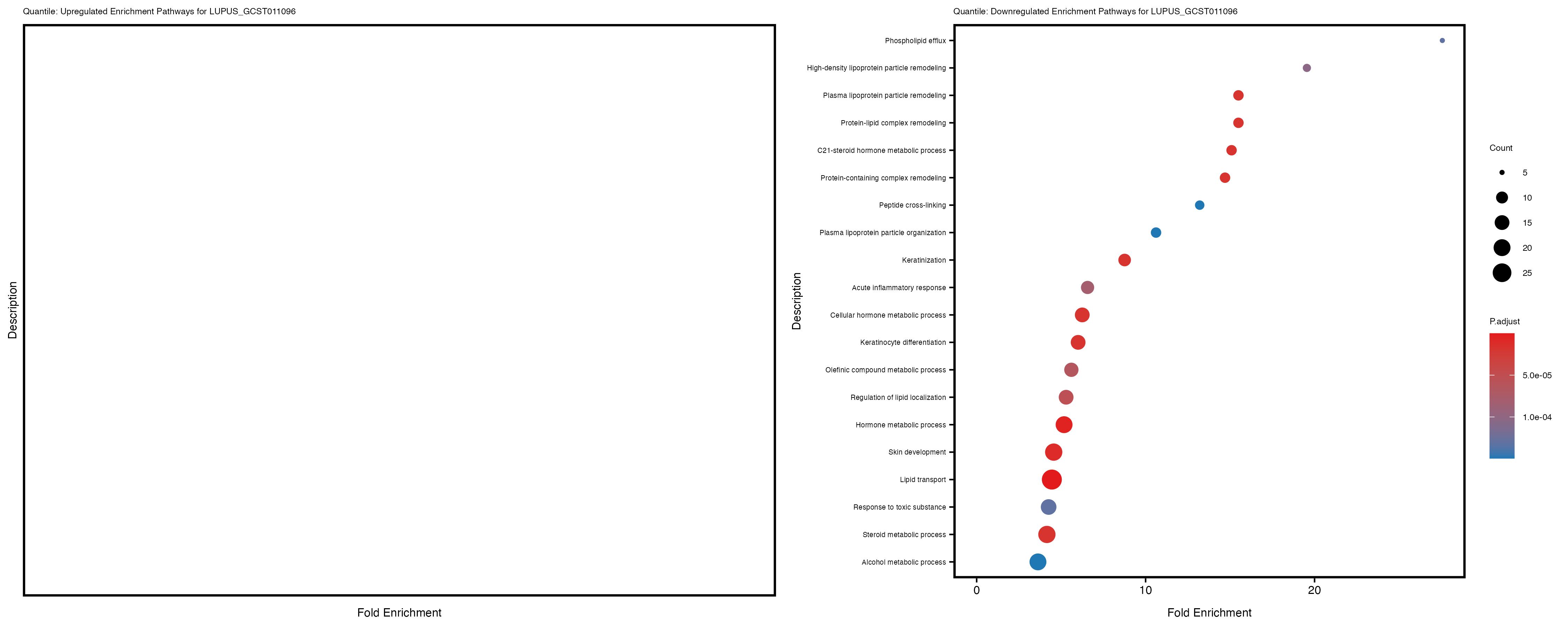
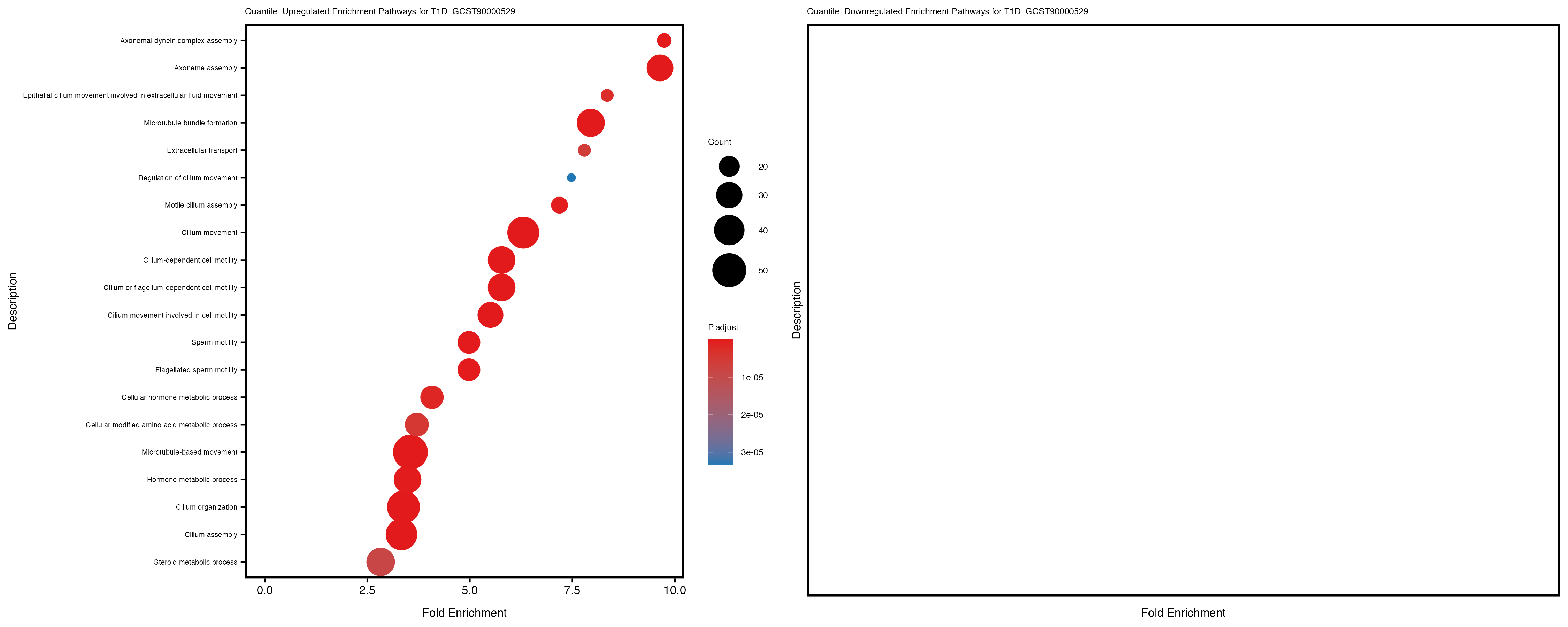
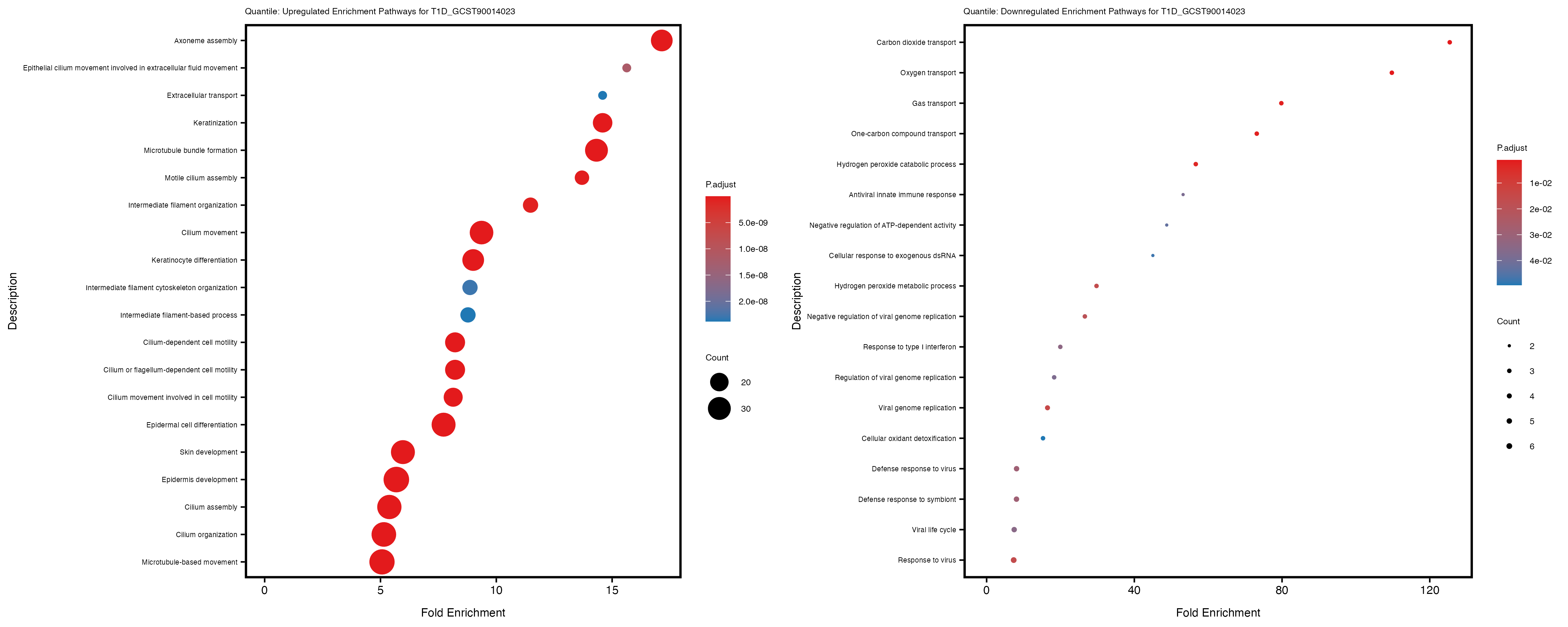
labs(title = paste("Quantile: Upregulated Enrichment Pathways for", trait)) +
theme(plot.title = element_text(size = 6), axis.text.y = element_text(size = 5))
} else {
enrich_plot_up <- plotEnrich(gse_up, plot_type = "dot", scale_ratio = 0.4) +
labs(title = paste("Quantile: Upregulated Enrichment Pathways for", trait)) +
theme(plot.title = element_text(size = 6), axis.text.y = element_text(size = 5))
}
if (nrow(gse_down) >= 20) {
enrich_plot_down <- plotEnrich(gse_down[1:20, ], plot_type = "dot", scale_ratio = 0.4) +
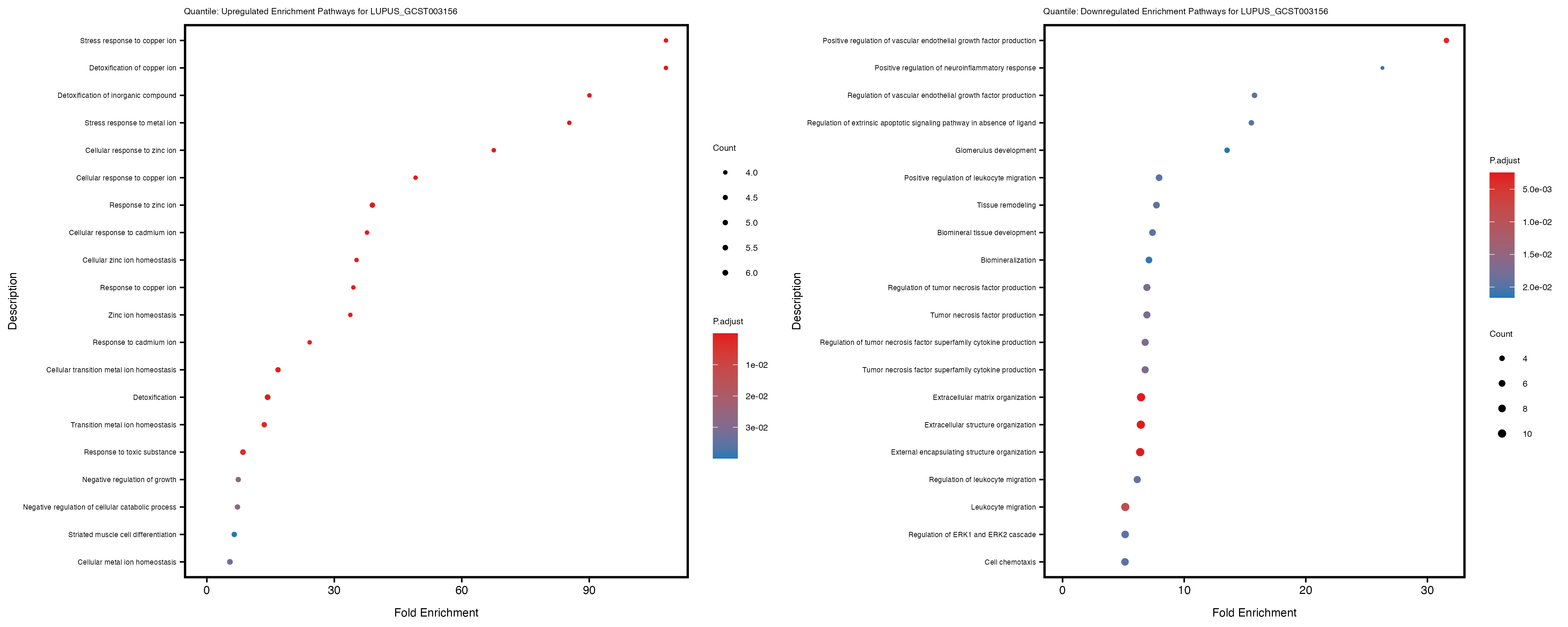
labs(title = paste("Quantile: Downregulated Enrichment Pathways for", trait)) +
theme(plot.title = element_text(size = 6), axis.text.y = element_text(size = 5))
} else {
enrich_plot_down <- plotEnrich(gse_down, plot_type = "dot", scale_ratio = 0.4) +
labs(title = paste("Quantile: Downregulated Enrichment Pathways for", trait)) +
theme(plot.title = element_text(size = 6), axis.text.y = element_text(size = 5))
}
# Arrange the two plots side by side
combined_plot <- grid.arrange(enrich_plot_up, enrich_plot_down, ncol = 2)
# Save the combined plot
ggsave(paste0("enrichment_plot_quantile_", trait, ".png"), plot = combined_plot,
width = 15, height = 6)
# Save the GO enrichment results to CSV
write.csv(gse_up, file = paste0("GO_enrichment_", trait, "_quantile_upregulated.csv"))
write.csv(gse_down, file = paste0("GO_enrichment_", trait, "_quantile_downregulated.csv"))
}Result
Summary table
| Trait | Significant DE genes | Up-regulated Genes | Down-regulated Genes | Up-regulated GO pathways | Down-regulated GO pathways |
|---|---|---|---|---|---|
| Basophil.count | 402 | 363 | 39 | 0 | 27 |
| Basophil.percentage.of.white.cells | 200 | 89 | 111 | 0 | 167 |
| Celiac_GCST90014442 | 97 | 55 | 42 | 7 | 73 |
| Celiac_GCST90468120 | 493 | 25 | 468 | 27 | 8 |
| Eosinophil.count | 338 | 128 | 210 | 0 | 88 |
| Eosinophil.percentage.of.white.cells | 209 | 85 | 124 | 1 | 28 |
| Hematocrit | 337 | 57 | 280 | 8 | 0 |
| Hemoglobin.concentration | 132 | 57 | 75 | 194 | 5 |
| High.light.scatter.reticulocyte.count | 239 | 52 | 187 | 34 | 73 |
| High.light.scatter.reticulocyte.percentage.of.red.cells | 325 | 94 | 231 | 15 | 161 |
| IBD_GCST90013901 | 559 | 522 | 37 | 5 | 96 |
| IBD_GCST90013951 | 380 | 338 | 42 | 199 | 71 |
| Immature.fraction.of.reticulocytes | 288 | 134 | 154 | 43 | 73 |
| LUPUS_GCST003156 | 119 | 79 | 40 | 48 | 39 |
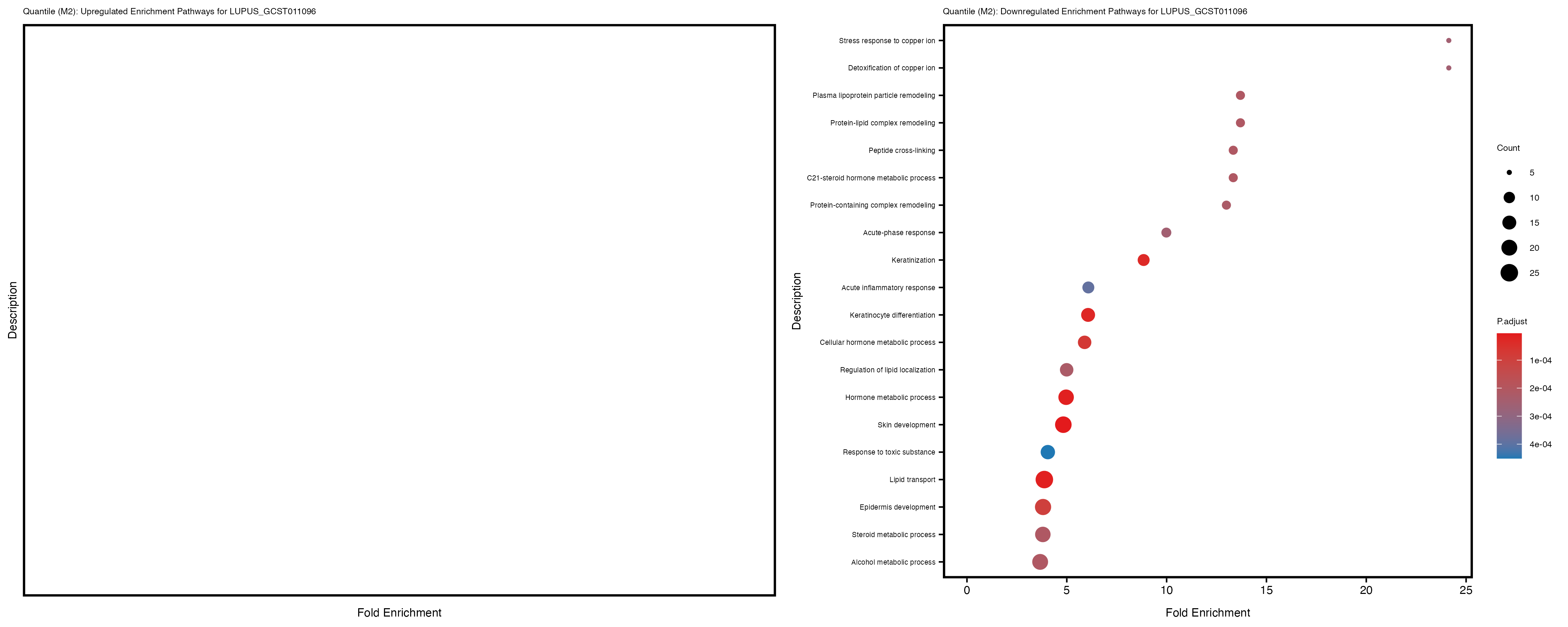
| LUPUS_GCST011096 | 131 | 89 | 42 | 66 | 132 |
| Lymphocyte.count | 729 | 347 | 382 | 12 | 71 |
| Lymphocyte.percentage.of.white.cells | 927 | 305 | 622 | 3 | 33 |
| Mean.corpuscular.hemoglobin | 402 | 22 | 380 | 0 | 55 |
| Mean.corpuscular.hemoglobin.concentration | 788 | 69 | 719 | 20 | 193 |
| Mean.corpuscular.volume | 162 | 22 | 140 | 0 | 44 |
| Mean.platelet.volume | 146 | 30 | 116 | 39 | 7 |
| Mean.reticulocyte.volume | 312 | 148 | 164 | 110 | 4 |
| Mean.sphered.corpuscular.volume | 533 | 81 | 452 | 89 | 0 |
| Monocyte.count | 348 | 180 | 168 | 4 | 33 |
| Monocyte.percentage.of.white.cells | 179 | 42 | 137 | 55 | 13 |
| Neutrophil.count | 1358 | 1260 | 98 | 46 | 43 |
| Neutrophil.percentage.of.white.cells | 421 | 357 | 64 | 21 | 50 |
| Platelet.count | 78 | 51 | 27 | 0 | 8 |
| Platelet.crit | 113 | 82 | 31 | 13 | 0 |
| Platelet.distribution.width | 105 | 32 | 73 | 38 | 0 |
| Red.blood.cell.count | 130 | 81 | 49 | 56 | 0 |
| Red.cell.distribution.width | 999 | 396 | 603 | 77 | 18 |
| Reticulocyte.count | 140 | 60 | 80 | 2 | 164 |
| Reticulocyte.fraction.of.red.cells | 225 | 122 | 103 | 23 | 25 |
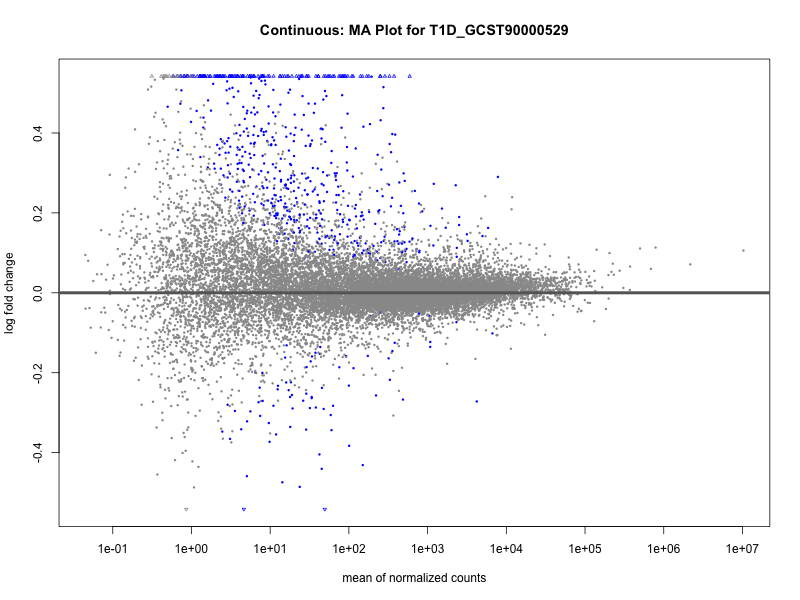
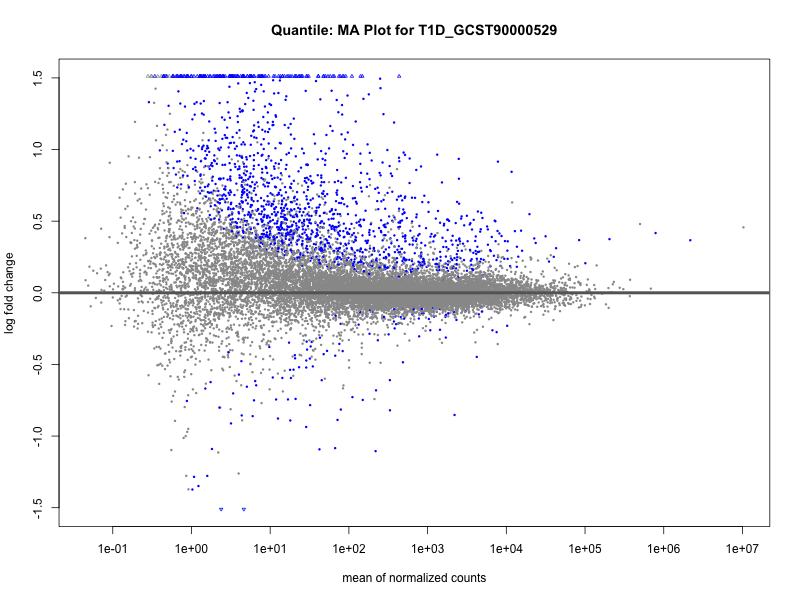
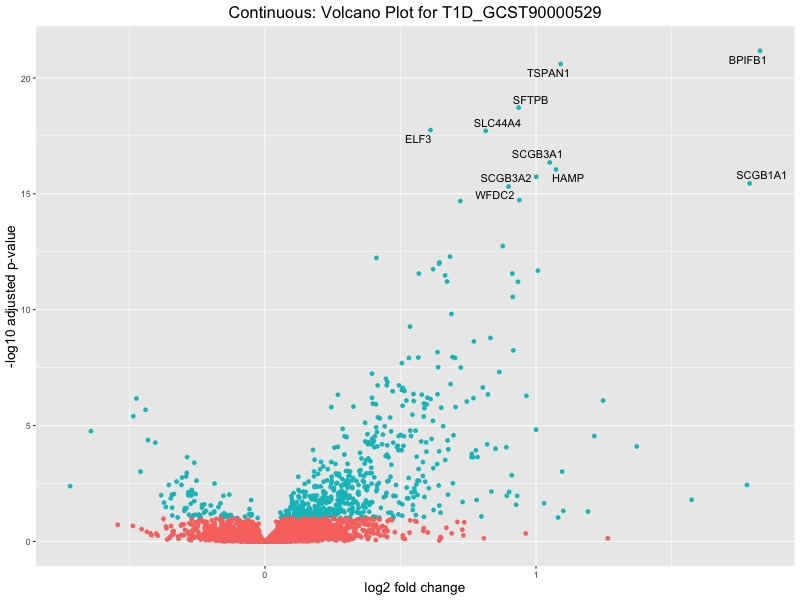
| T1D_GCST90000529 | 618 | 540 | 78 | 30 | 22 |
| T1D_GCST90014023 | 205 | 135 | 70 | 1 | 4 |
| White.blood.cell.count | 1314 | 1118 | 196 | 40 | 0 |
| Trait | Significant DE genes | Up-regulated Genes | Down-regulated Genes | Up-regulated GO pathways | Down-regulated GO pathways |
|---|---|---|---|---|---|
| Basophil.count | 385 | 345 | 40 | 0 | 47 |
| Basophil.percentage.of.white.cells | 203 | 93 | 110 | 0 | 167 |
| Celiac_GCST90014442 | 94 | 55 | 39 | 2 | 114 |
| Celiac_GCST90468120 | 493 | 25 | 468 | 2 | 9 |
| Eosinophil.count | 333 | 126 | 207 | 0 | 89 |
| Eosinophil.percentage.of.white.cells | 211 | 87 | 124 | 1 | 21 |
| Hematocrit | 318 | 51 | 267 | 8 | 0 |
| Hemoglobin.concentration | 120 | 54 | 66 | 0 | 0 |
| High.light.scatter.reticulocyte.count | 237 | 52 | 185 | 34 | 74 |
| High.light.scatter.reticulocyte.percentage.of.red.cells | 328 | 101 | 227 | 14 | 153 |
| IBD_GCST90013901 | 561 | 525 | 36 | 15 | 96 |
| IBD_GCST90013951 | 378 | 340 | 38 | 183 | 71 |
| Immature.fraction.of.reticulocytes | 294 | 142 | 152 | 44 | 73 |
| LUPUS_GCST003156 | 123 | 82 | 41 | 42 | 279 |
| LUPUS_GCST011096 | 138 | 94 | 44 | 60 | 236 |
| Lymphocyte.count | 718 | 341 | 377 | 8 | 74 |
| Lymphocyte.percentage.of.white.cells | 923 | 309 | 614 | 3 | 30 |
| Mean.corpuscular.hemoglobin | 381 | 22 | 359 | 0 | 63 |
| Mean.corpuscular.hemoglobin.concentration | 785 | 71 | 714 | 0 | 244 |
| Mean.corpuscular.volume | 162 | 22 | 140 | 0 | 40 |
| Mean.platelet.volume | 147 | 31 | 116 | 5 | 9 |
| Mean.reticulocyte.volume | 312 | 147 | 165 | 105 | 2 |
| Mean.sphered.corpuscular.volume | 534 | 84 | 450 | 43 | 0 |
| Monocyte.count | 366 | 188 | 178 | 10 | 33 |
| Monocyte.percentage.of.white.cells | 182 | 43 | 139 | 30 | 21 |
| Neutrophil.count | 1374 | 1271 | 103 | 44 | 43 |
| Neutrophil.percentage.of.white.cells | 428 | 361 | 67 | 19 | 18 |
| Platelet.count | 73 | 47 | 26 | 0 | 8 |
| Platelet.crit | 110 | 80 | 30 | 13 | 0 |
| Platelet.distribution.width | 93 | 31 | 62 | 39 | 0 |
| Red.blood.cell.count | 110 | 67 | 43 | 29 | 0 |
| Red.cell.distribution.width | 990 | 401 | 589 | 16 | 19 |
| Reticulocyte.count | 129 | 57 | 72 | 2 | 150 |
| Reticulocyte.fraction.of.red.cells | 213 | 120 | 93 | 23 | 0 |
| T1D_GCST90000529 | 619 | 537 | 82 | 32 | 23 |
| T1D_GCST90014023 | 192 | 123 | 69 | 2 | 4 |
| White.blood.cell.count | 1313 | 1118 | 195 | 39 | 48 |
| Trait | Significant DE genes | Up-regulated Genes | Down-regulated Genes | Up-regulated GO pathways | Down-regulated GO pathways |
|---|---|---|---|---|---|
| Basophil.count_quantile | 712 | 685 | 27 | 144 | 16 |
| Basophil.percentage.of.white.cells_quantile | 184 | 64 | 120 | 61 | 89 |
| Celiac_GCST90014442_quantile | 336 | 124 | 212 | 105 | 320 |
| Celiac_GCST90468120_quantile | 212 | 42 | 170 | 94 | 27 |
| Eosinophil.count_quantile | 507 | 197 | 310 | 12 | 157 |
| Eosinophil.percentage.of.white.cells_quantile | 431 | 218 | 213 | 3 | 165 |
| Hematocrit_quantile | 269 | 118 | 151 | 114 | 35 |
| Hemoglobin.concentration_quantile | 160 | 37 | 123 | 36 | 104 |
| High.light.scatter.reticulocyte.count_quantile | 225 | 93 | 132 | 61 | 98 |
| High.light.scatter.reticulocyte.percentage.of.red.cells_quantile | 312 | 141 | 171 | 49 | 158 |
| IBD_GCST90013901_quantile | 272 | 204 | 68 | 147 | 130 |
| IBD_GCST90013951_quantile | 346 | 256 | 90 | 158 | 65 |
| Immature.fraction.of.reticulocytes_quantile | 473 | 128 | 345 | 104 | 149 |
| LUPUS_GCST003156_quantile | 205 | 96 | 109 | 20 | 57 |
| LUPUS_GCST011096_quantile | 493 | 52 | 441 | 0 | 202 |
| Lymphocyte.count_quantile | 1456 | 852 | 604 | 95 | 188 |
| Lymphocyte.percentage.of.white.cells_quantile | 532 | 289 | 243 | 37 | 112 |
| Mean.corpuscular.hemoglobin_quantile | 410 | 23 | 387 | 40 | 89 |
| Mean.corpuscular.hemoglobin.concentration_quantile | 317 | 86 | 231 | 30 | 336 |
| Mean.corpuscular.volume_quantile | 453 | 58 | 395 | 119 | 98 |
| Mean.platelet.volume_quantile | 209 | 85 | 124 | 55 | 26 |
| Mean.reticulocyte.volume_quantile | 406 | 115 | 291 | 106 | 45 |
| Mean.sphered.corpuscular.volume_quantile | 355 | 76 | 279 | 55 | 48 |
| Monocyte.count_quantile | 1175 | 845 | 330 | 109 | 62 |
| Monocyte.percentage.of.white.cells_quantile | 566 | 50 | 516 | 36 | 69 |
| Neutrophil.count_quantile | 806 | 736 | 70 | 105 | 13 |
| Neutrophil.percentage.of.white.cells_quantile | 1451 | 1330 | 121 | 396 | 31 |
| Platelet.count_quantile | 248 | 59 | 189 | 59 | 48 |
| Platelet.crit_quantile | 314 | 100 | 214 | 59 | 103 |
| Platelet.distribution.width_quantile | 469 | 149 | 320 | 39 | 160 |
| Red.blood.cell.count_quantile | 291 | 157 | 134 | 103 | 61 |
| Red.cell.distribution.width_quantile | 399 | 287 | 112 | 45 | 24 |
| Reticulocyte.count_quantile | 170 | 74 | 96 | 34 | 68 |
| Reticulocyte.fraction.of.red.cells_quantile | 517 | 458 | 59 | 191 | 1 |
| T1D_GCST90000529_quantile | 1559 | 1438 | 121 | 268 | 0 |
| T1D_GCST90014023_quantile | 575 | 504 | 71 | 87 | 18 |
| White.blood.cell.count_quantile | 1216 | 1082 | 134 | 156 | 118 |
| Trait | Significant DE genes | Up-regulated Genes | Down-regulated Genes | Up-regulated GO pathways | Down-regulated GO pathways |
|---|---|---|---|---|---|
| Basophil.count_quantile | 707 | 682 | 25 | 148 | 32 |
| Basophil.percentage.of.white.cells_quantile | 174 | 62 | 112 | 70 | 89 |
| Celiac_GCST90014442_quantile | 335 | 124 | 211 | 106 | 306 |
| Celiac_GCST90468120_quantile | 208 | 43 | 165 | 86 | 28 |
| Eosinophil.count_quantile | 498 | 191 | 307 | 13 | 158 |
| Eosinophil.percentage.of.white.cells_quantile | 426 | 219 | 207 | 3 | 160 |
| Hematocrit_quantile | 235 | 108 | 127 | 114 | 37 |
| Hemoglobin.concentration_quantile | 156 | 35 | 121 | 0 | 117 |
| High.light.scatter.reticulocyte.count_quantile | 218 | 94 | 124 | 62 | 82 |
| High.light.scatter.reticulocyte.percentage.of.red.cells_quantile | 316 | 149 | 167 | 42 | 151 |
| IBD_GCST90013901_quantile | 269 | 200 | 69 | 142 | 129 |
| IBD_GCST90013951_quantile | 337 | 250 | 87 | 168 | 66 |
| Immature.fraction.of.reticulocytes_quantile | 456 | 127 | 329 | 103 | 146 |
| LUPUS_GCST003156_quantile | 202 | 97 | 105 | 17 | 61 |
| LUPUS_GCST011096_quantile | 471 | 47 | 424 | 0 | 173 |
| Lymphocyte.count_quantile | 1603 | 907 | 696 | 109 | 228 |
| Lymphocyte.percentage.of.white.cells_quantile | 511 | 285 | 226 | 37 | 117 |
| Mean.corpuscular.hemoglobin_quantile | 390 | 25 | 365 | 41 | 100 |
| Mean.corpuscular.hemoglobin.concentration_quantile | 316 | 91 | 225 | 48 | 331 |
| Mean.corpuscular.volume_quantile | 425 | 55 | 370 | 110 | 106 |
| Mean.platelet.volume_quantile | 205 | 85 | 120 | 106 | 28 |
| Mean.reticulocyte.volume_quantile | 403 | 111 | 292 | 115 | 34 |
| Mean.sphered.corpuscular.volume_quantile | 366 | 78 | 288 | 62 | 45 |
| Monocyte.count_quantile | 1186 | 844 | 342 | 104 | 49 |
| Monocyte.percentage.of.white.cells_quantile | 589 | 50 | 539 | 18 | 82 |
| Neutrophil.count_quantile | 808 | 736 | 72 | 121 | 13 |
| Neutrophil.percentage.of.white.cells_quantile | 1467 | 1335 | 132 | 416 | 21 |
| Platelet.count_quantile | 235 | 53 | 182 | 49 | 61 |
| Platelet.crit_quantile | 309 | 93 | 216 | 68 | 92 |
| Platelet.distribution.width_quantile | 450 | 147 | 303 | 32 | 150 |
| Red.blood.cell.count_quantile | 264 | 143 | 121 | 116 | 81 |
| Red.cell.distribution.width_quantile | 405 | 293 | 112 | 44 | 23 |
| Reticulocyte.count_quantile | 169 | 74 | 95 | 36 | 66 |
| Reticulocyte.fraction.of.red.cells_quantile | 516 | 461 | 55 | 225 | 3 |
| T1D_GCST90000529_quantile | 1543 | 1417 | 126 | 243 | 0 |
| T1D_GCST90014023_quantile | 554 | 482 | 72 | 91 | 16 |
| White.blood.cell.count_quantile | 1173 | 1042 | 131 | 181 | 120 |
Interpretation:
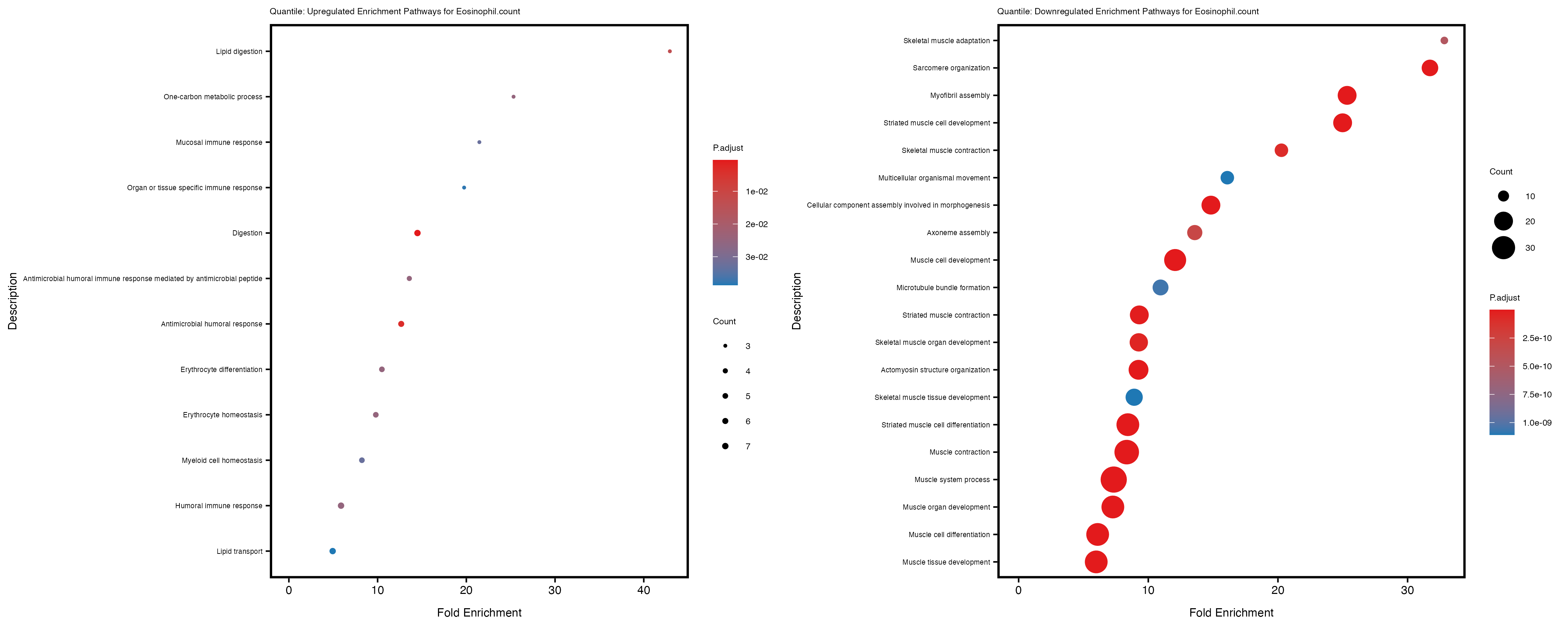
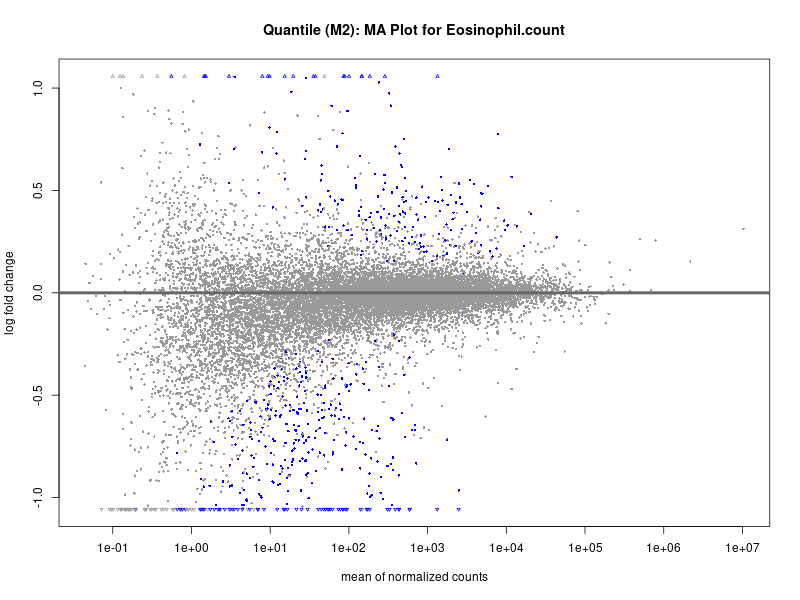
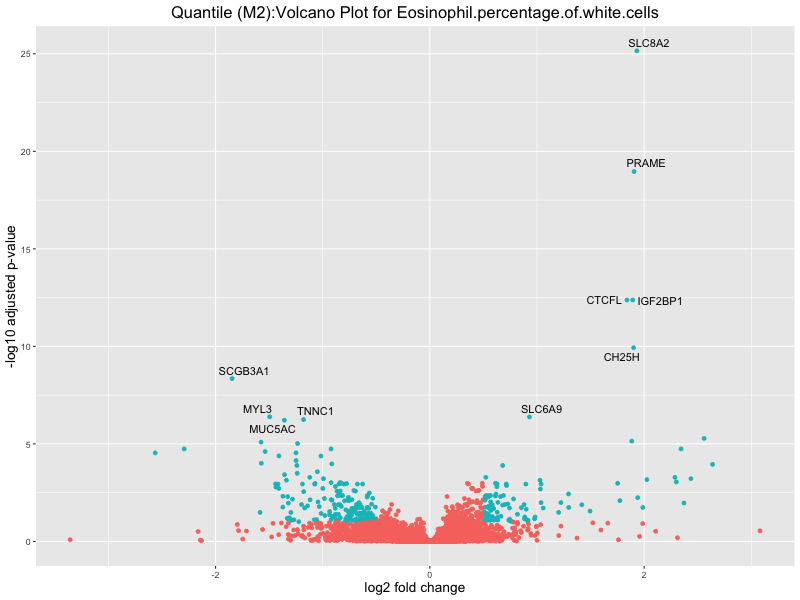
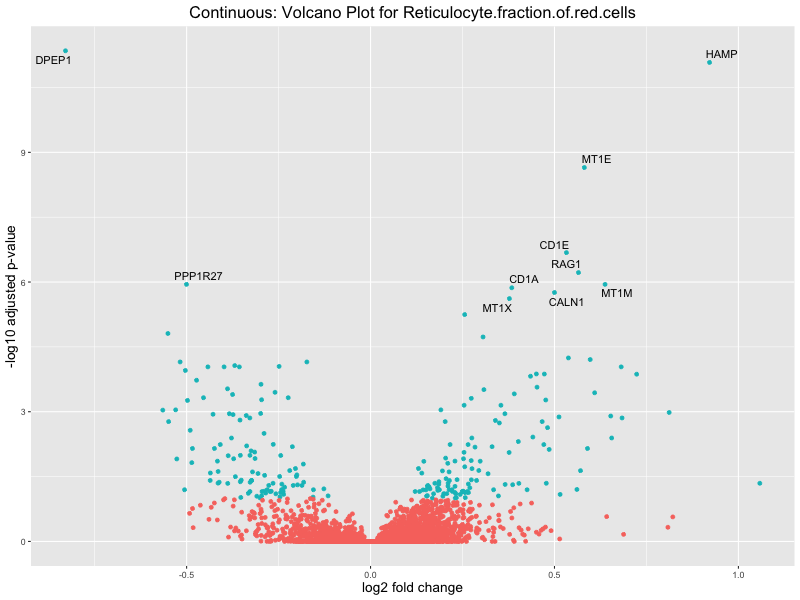
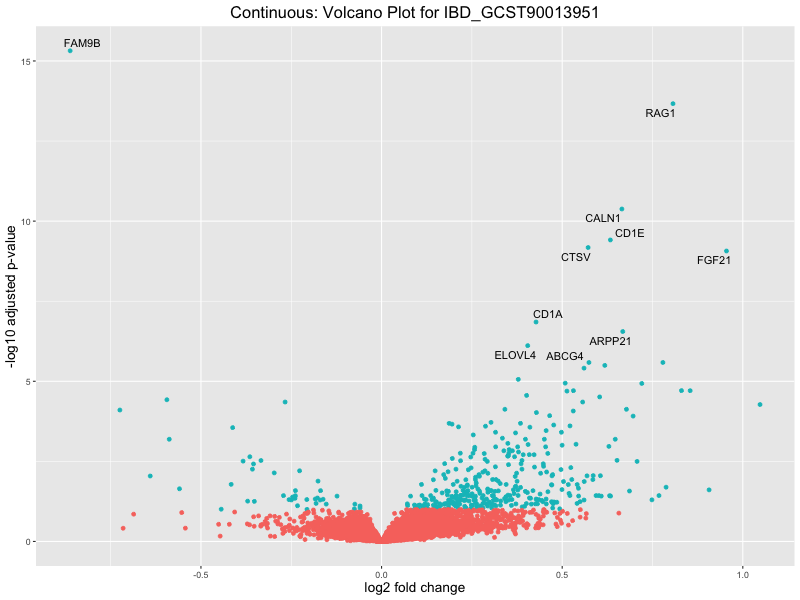
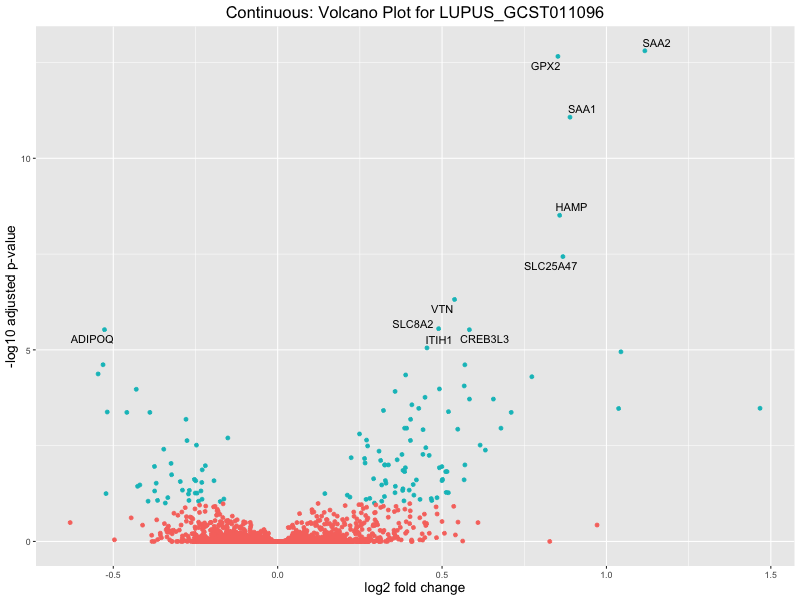
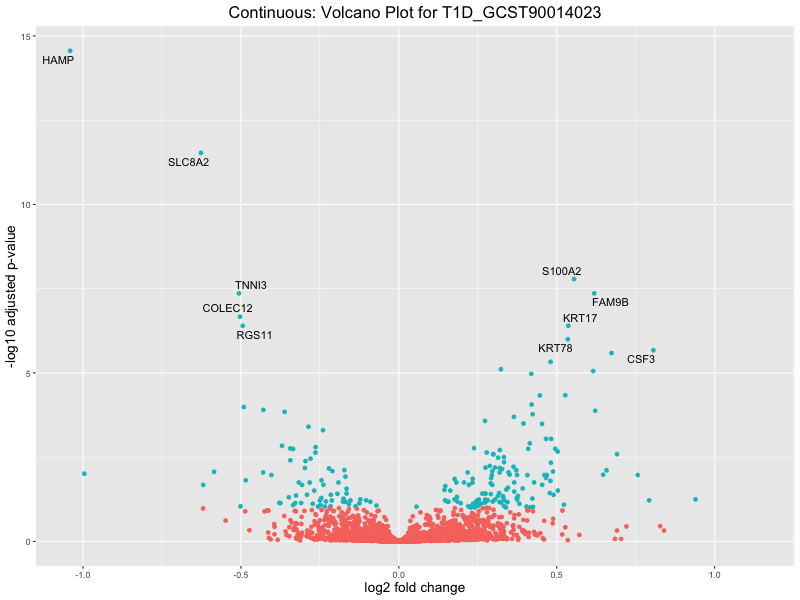
In the quantile analysis, the number of significant differential expressed genes is generally higher compared to the continuous analysis, which suggests that stratifying by PRS (top 25%) may identify more genes with stronger associations to the traits. We observed that certain genes were identified in both the continuous and quantile PRS analyses, though not all genes overlapped. Specifically, for the Eosinophil count trait, ACTN3, HAMP, SCGB3A1, CH25H, and NPIPA8 appeared among the top 10 DEGs across both analyses, suggesting that these genes might play a key role in the trait regardless of the analysis approach. For example:
HAMP encodes Hepcidin, which is involved in the maintenance of iron homeostasis.
SCGB3A1 predicts to enable cytokin activity.
CH25H encodes for cholesterol 25-hydroxylase, involved in lipid metabolism and modulating immune responses.
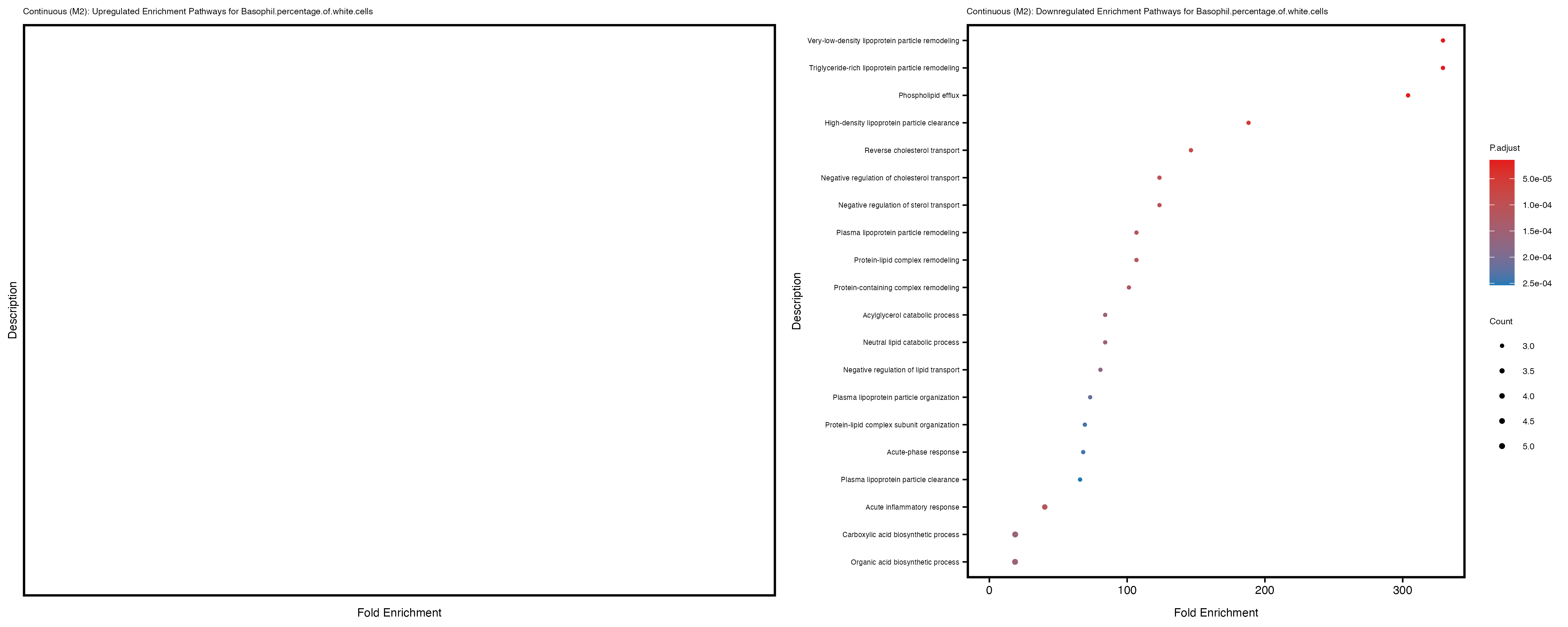
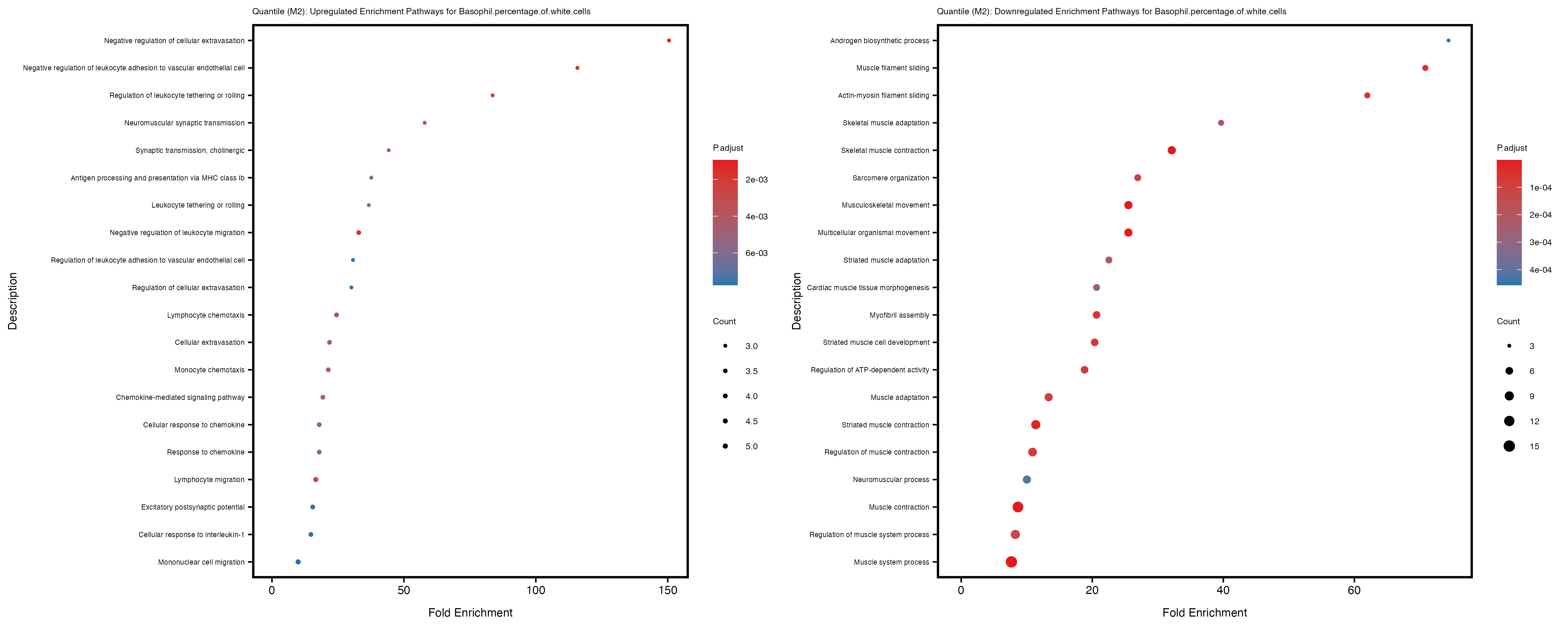
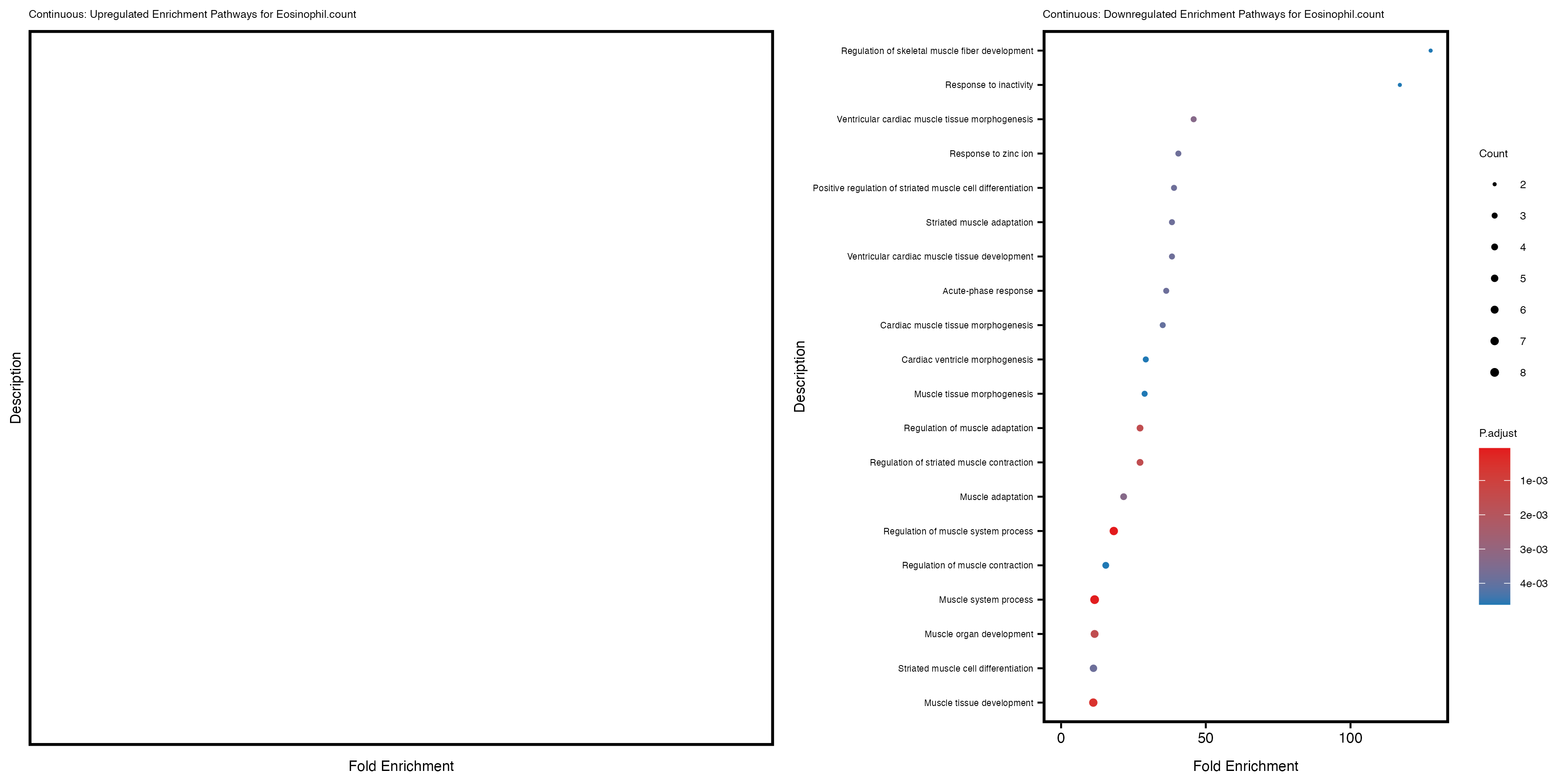
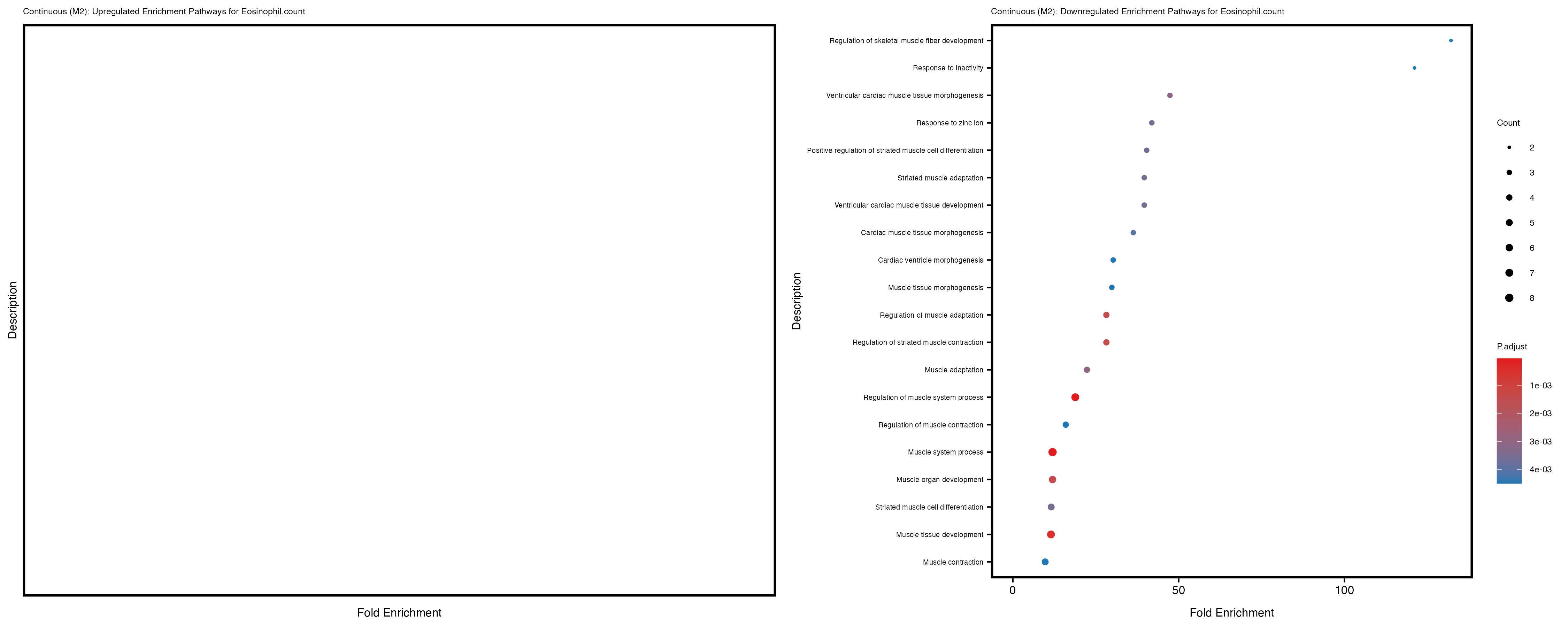
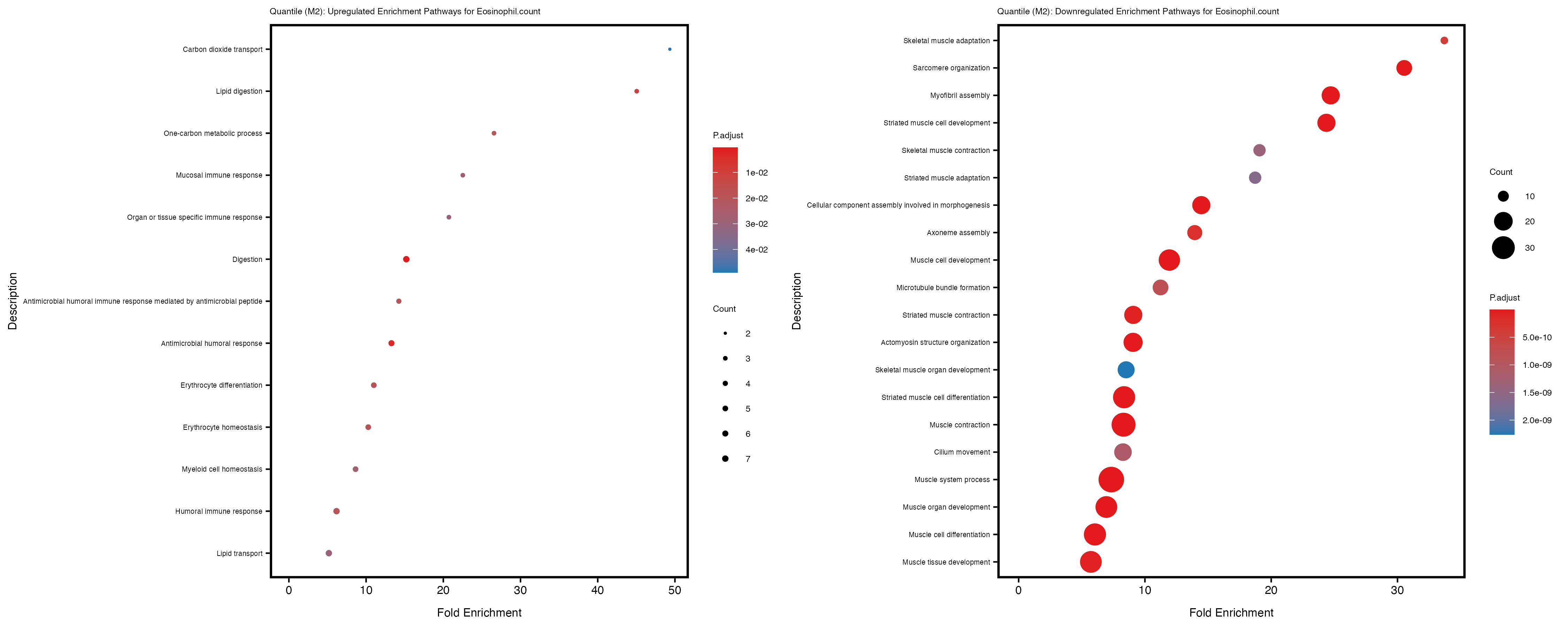
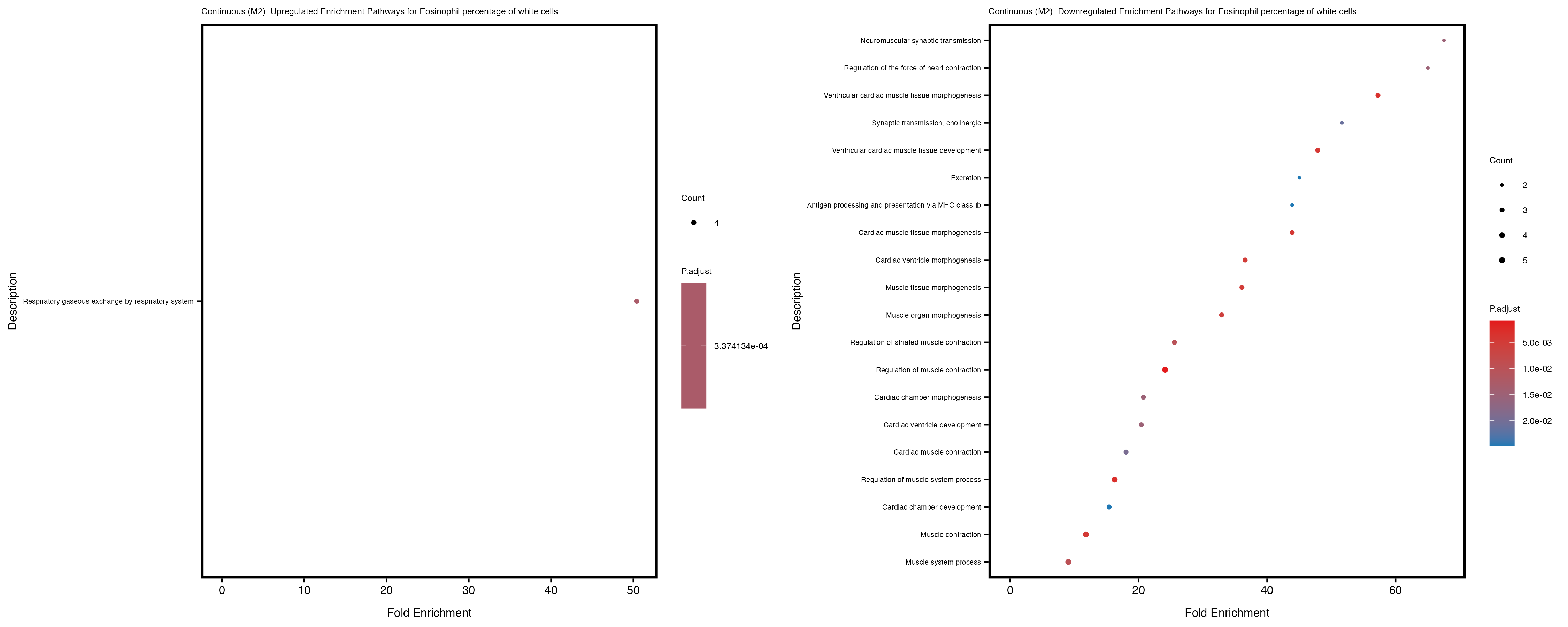
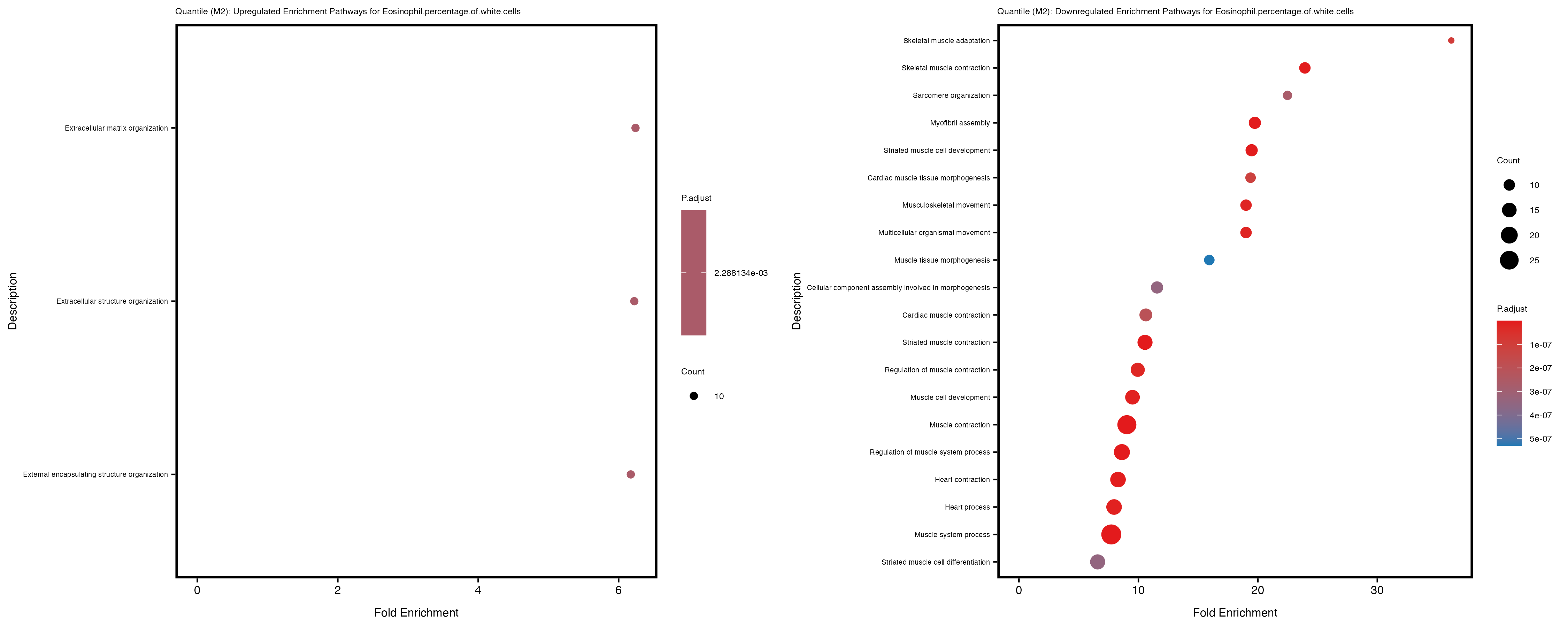
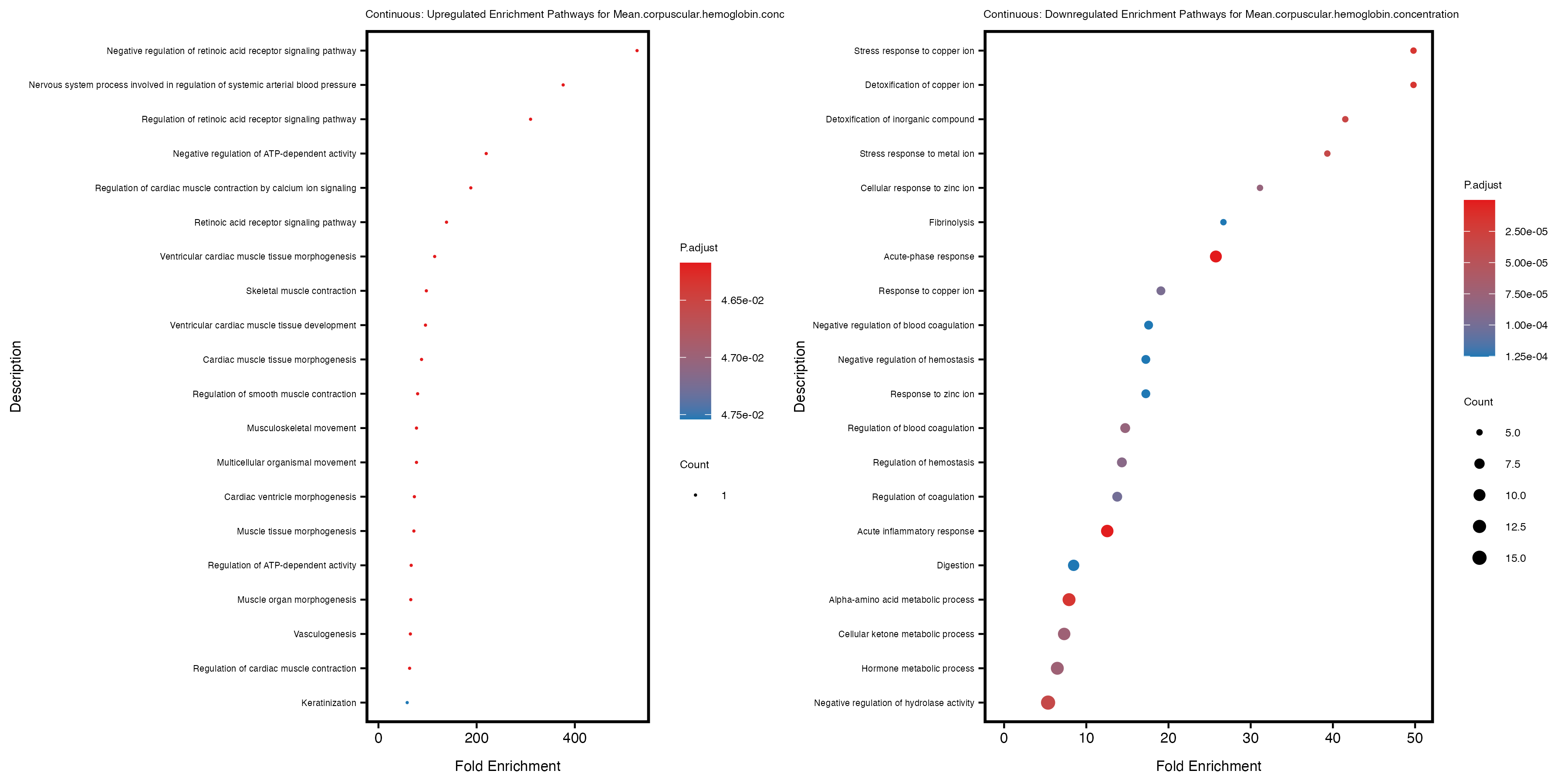
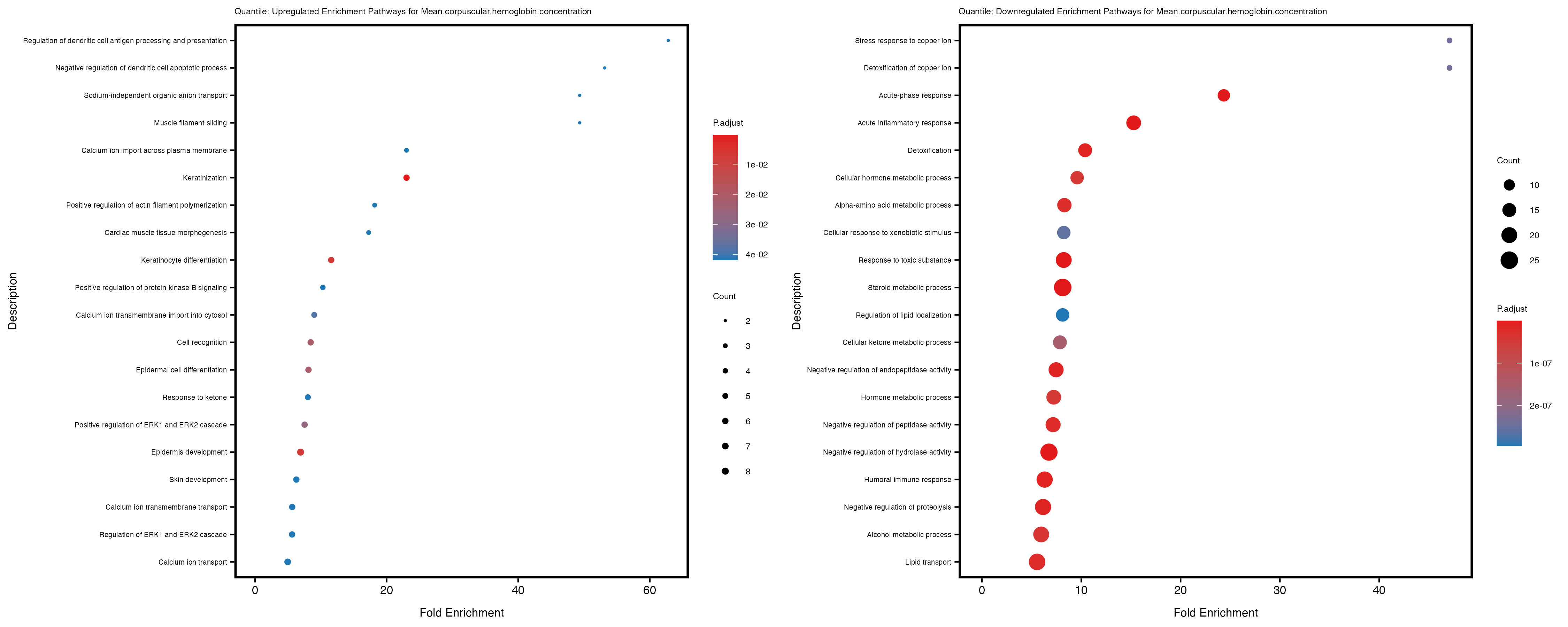
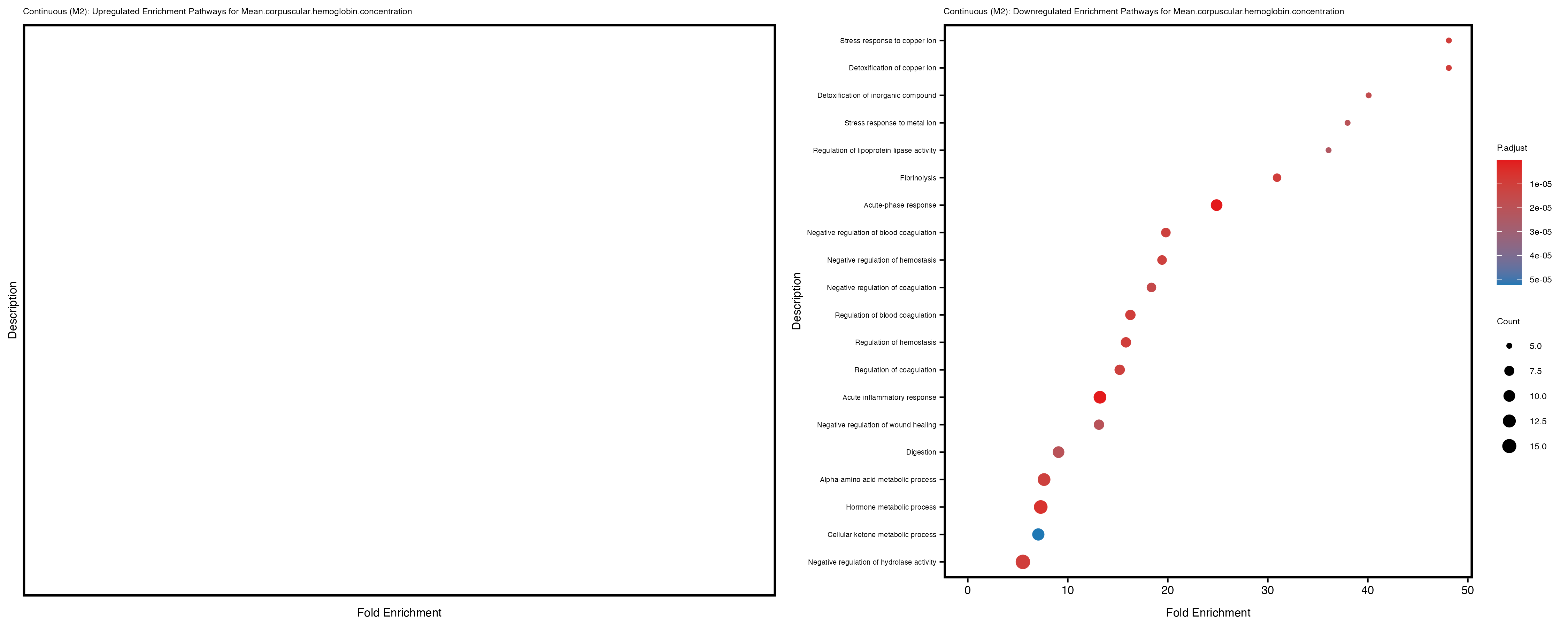
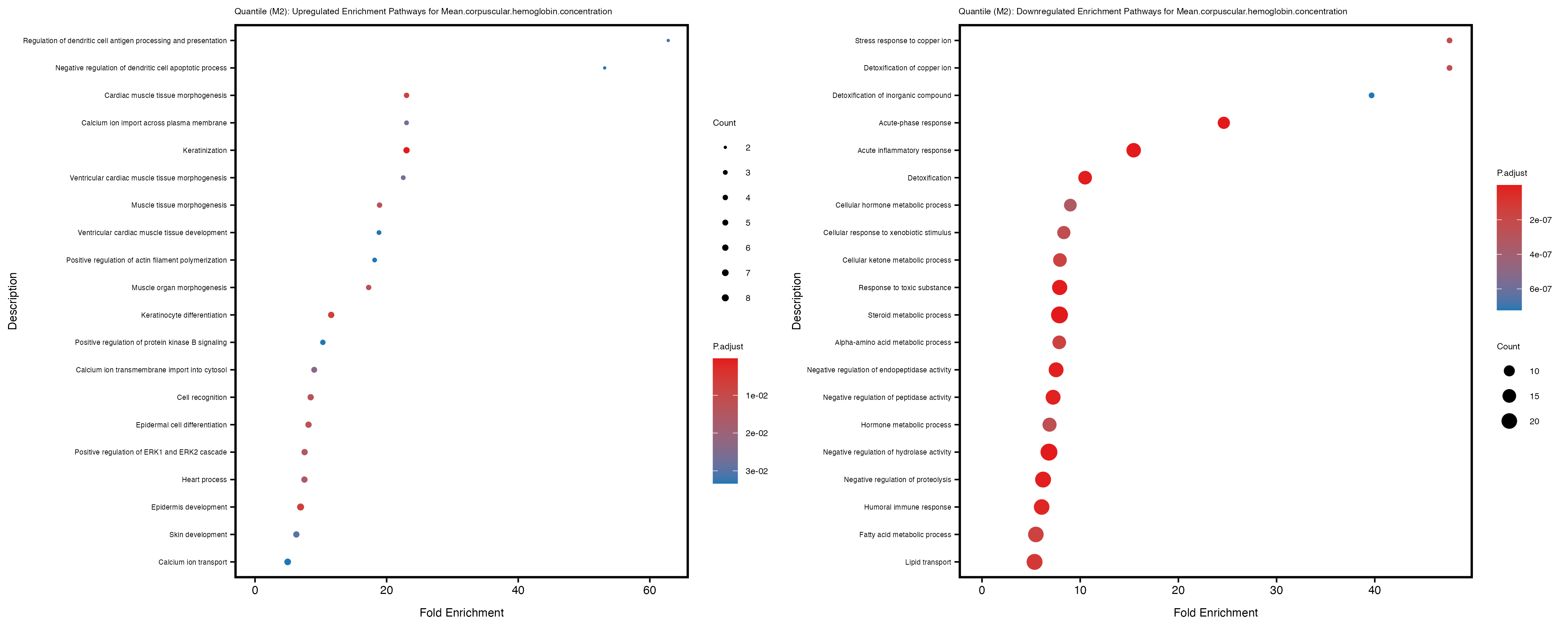
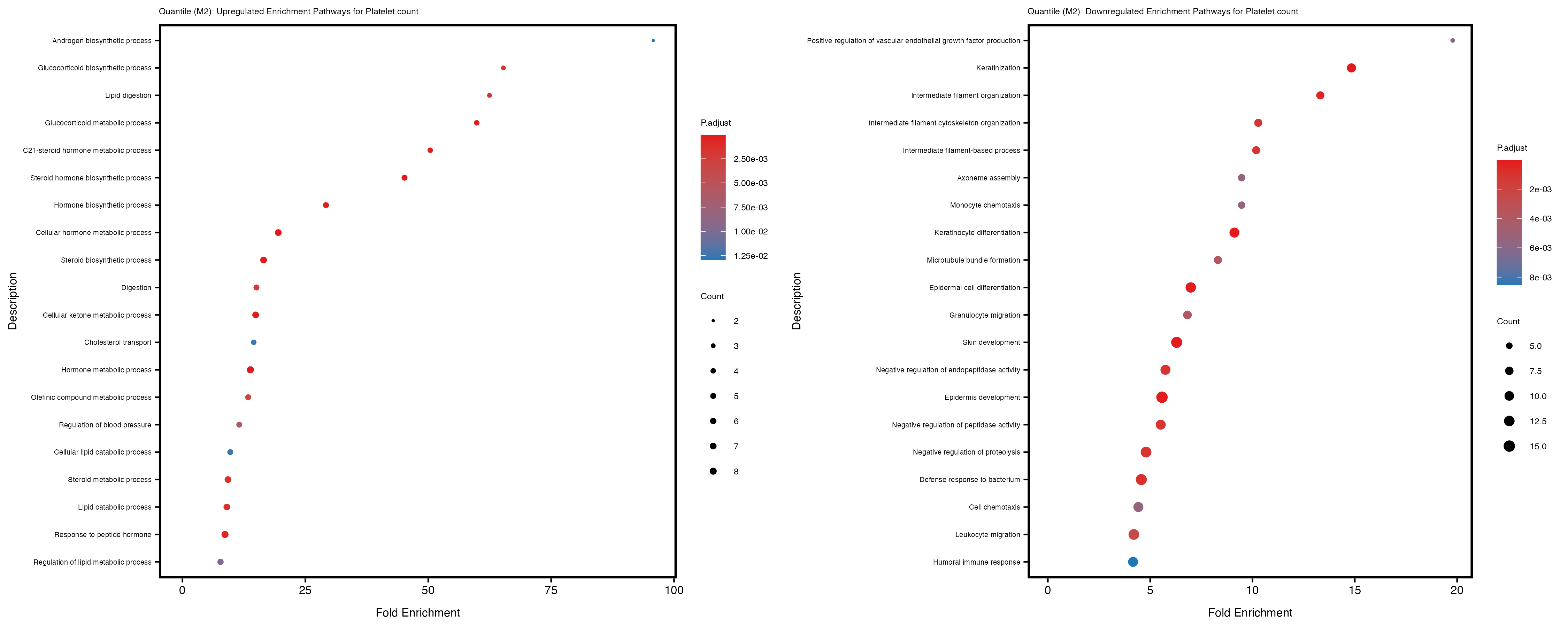
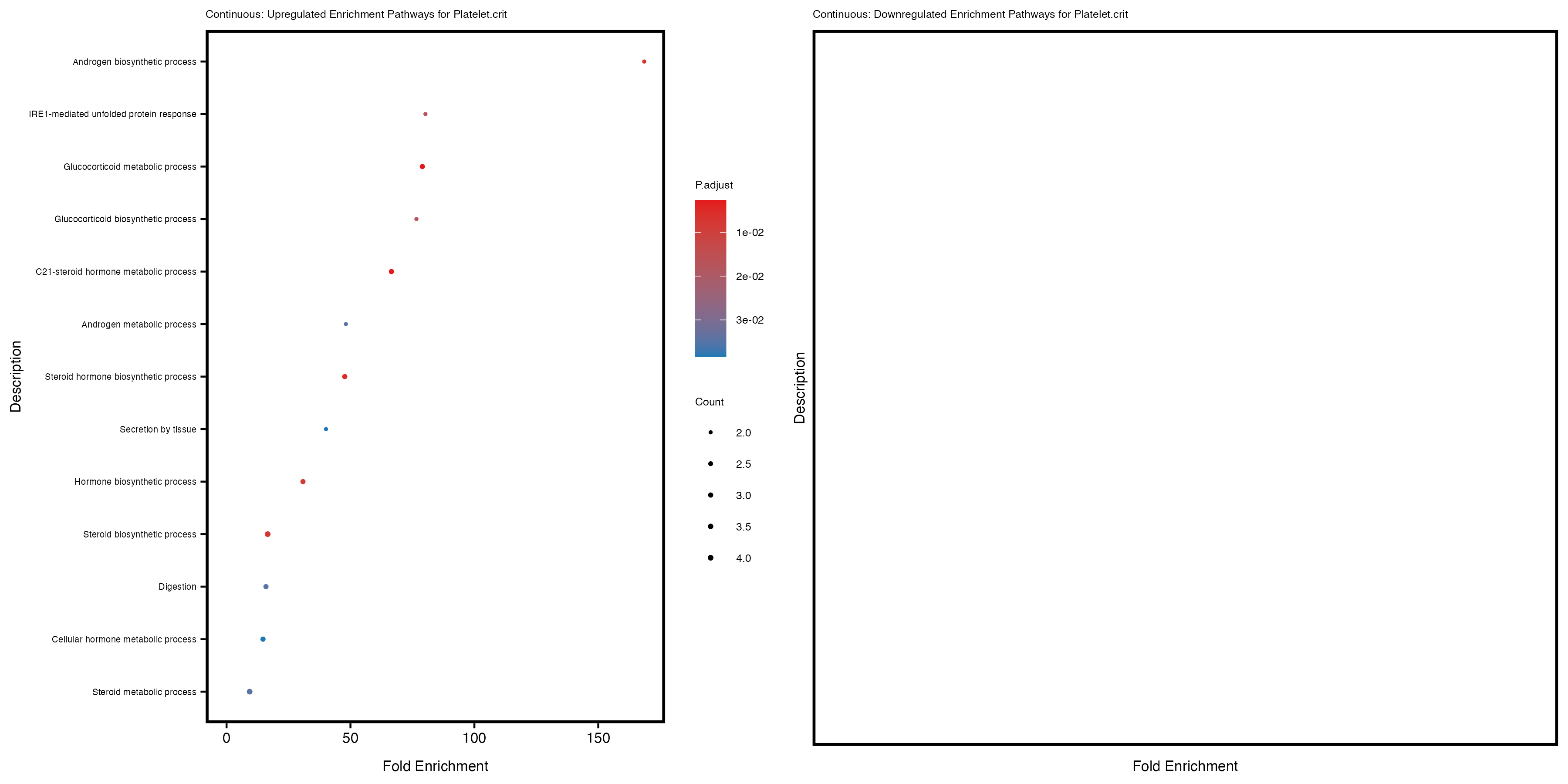
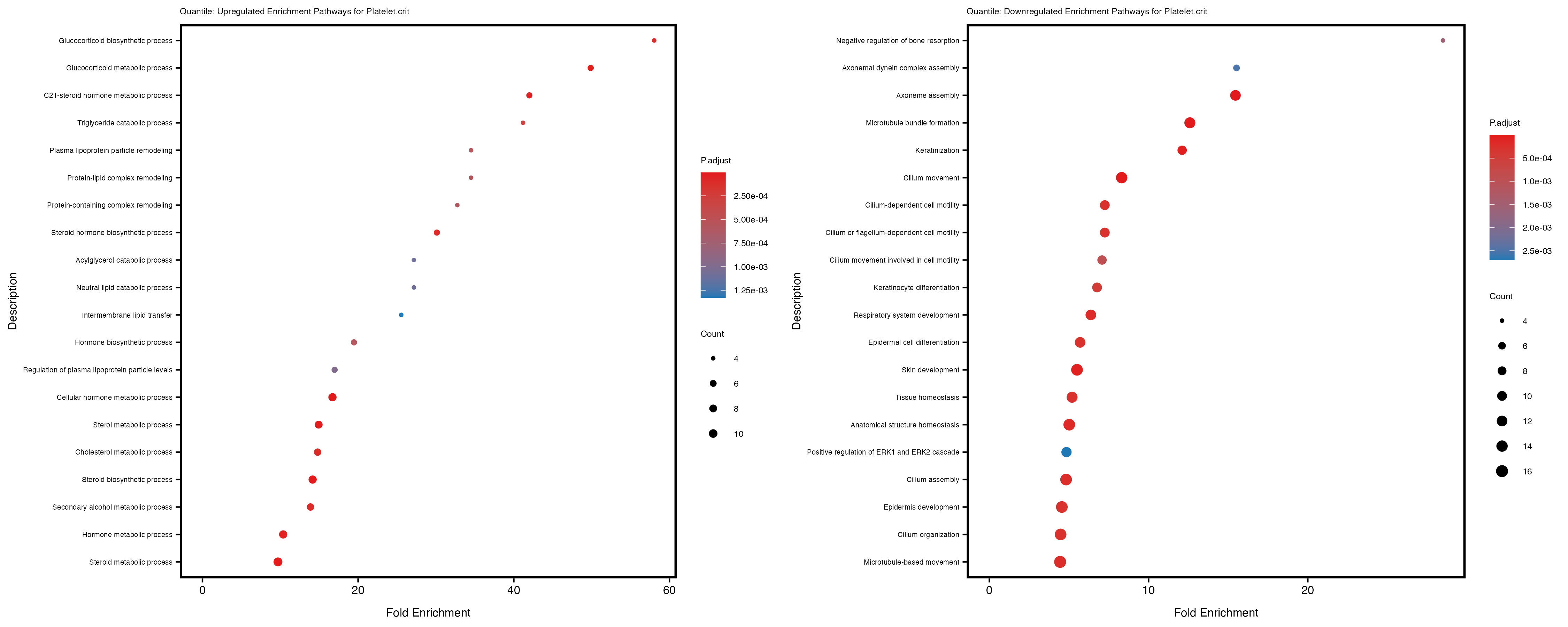
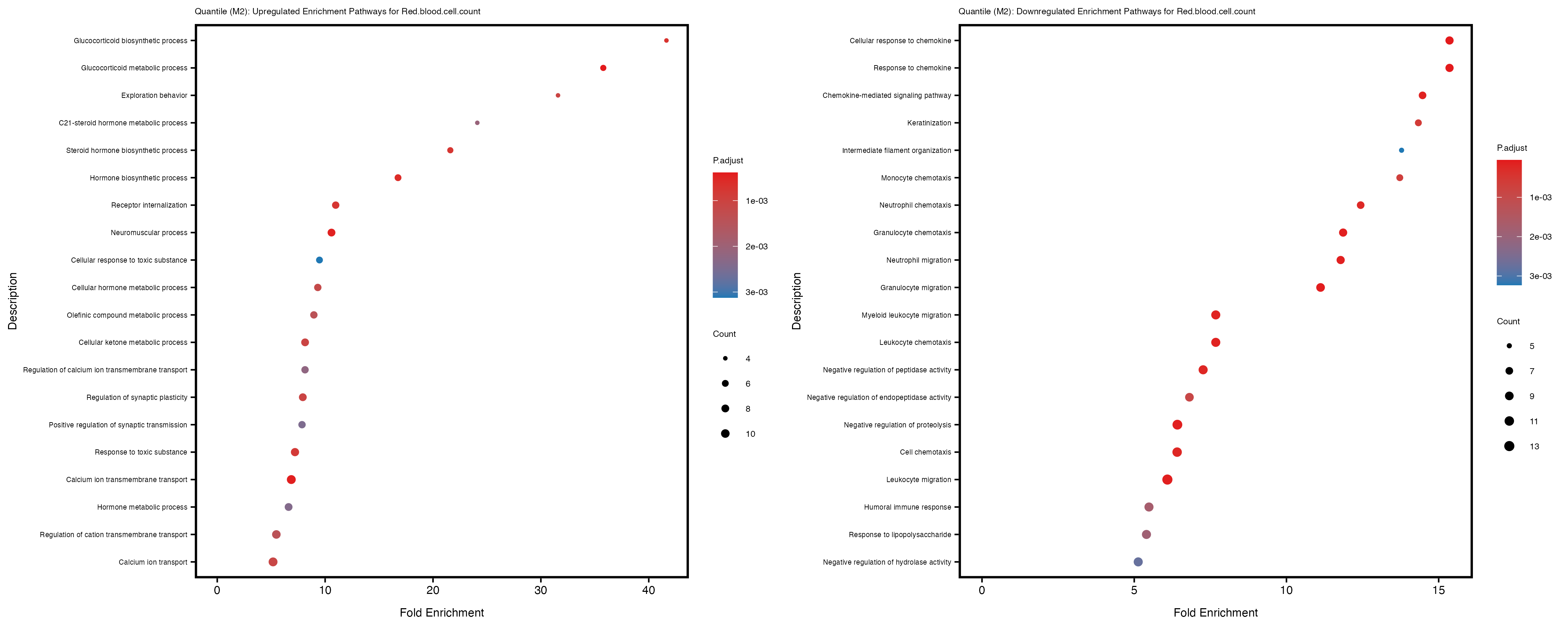
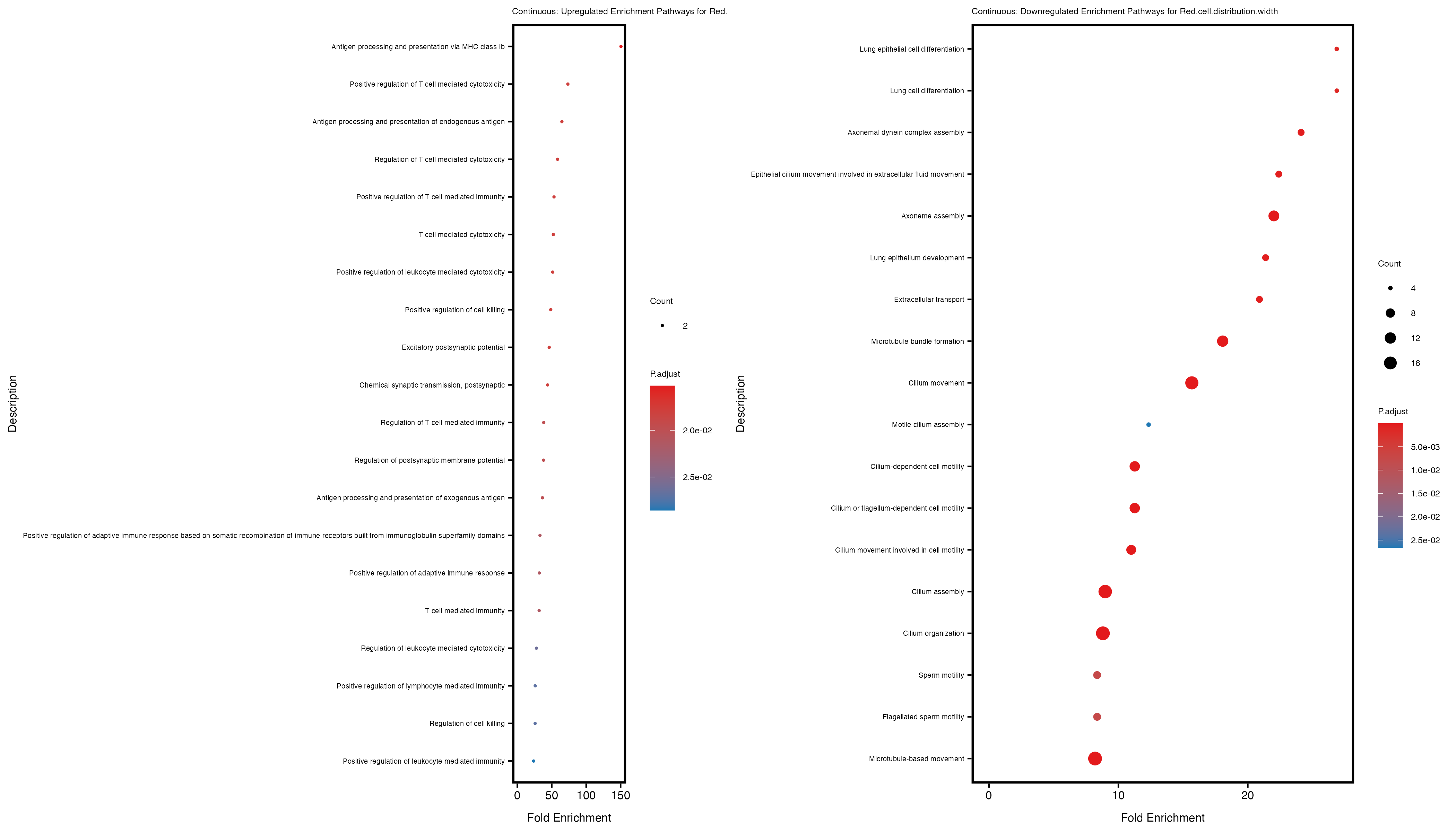
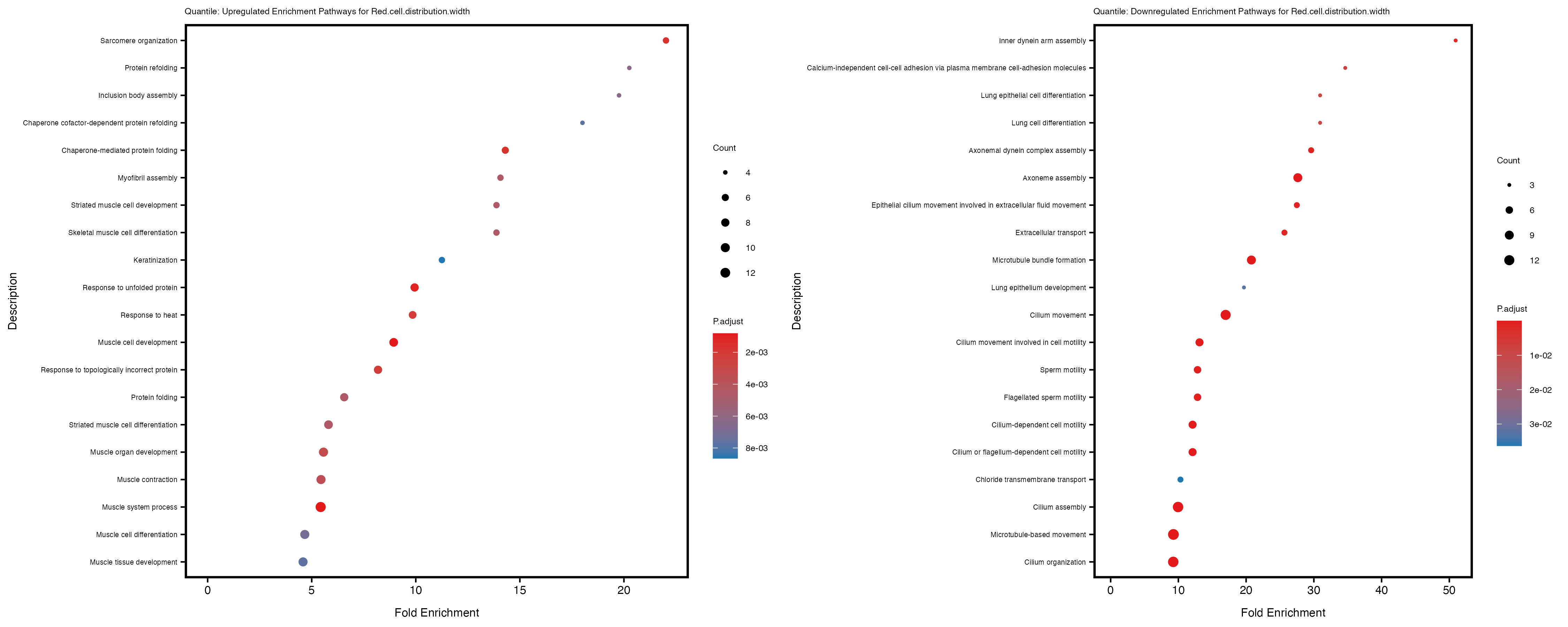
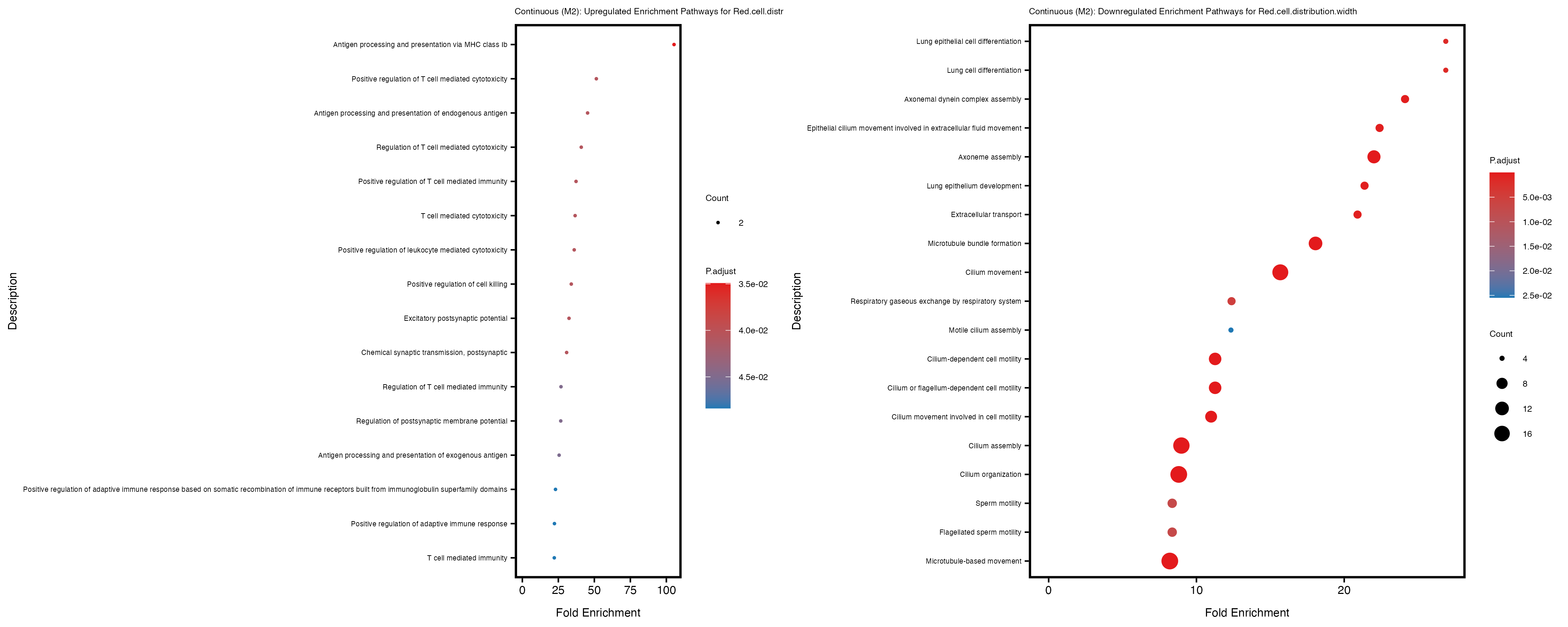
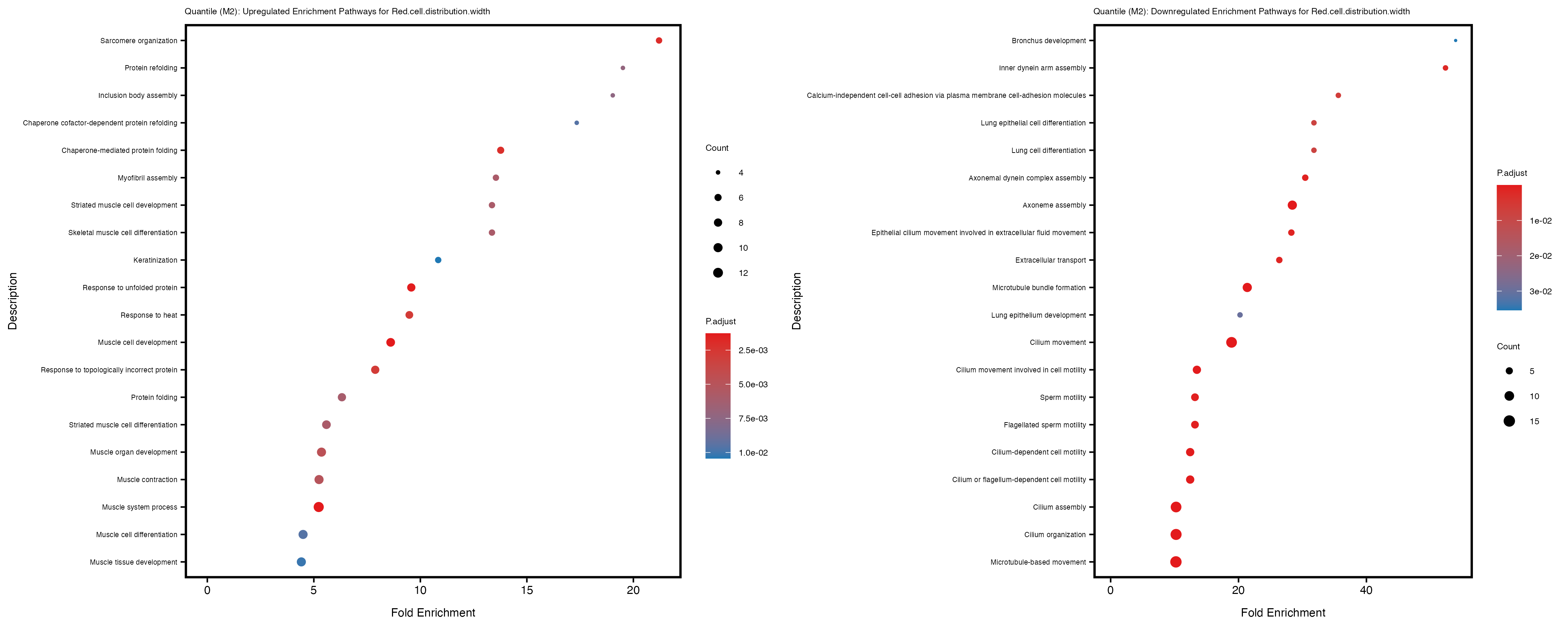
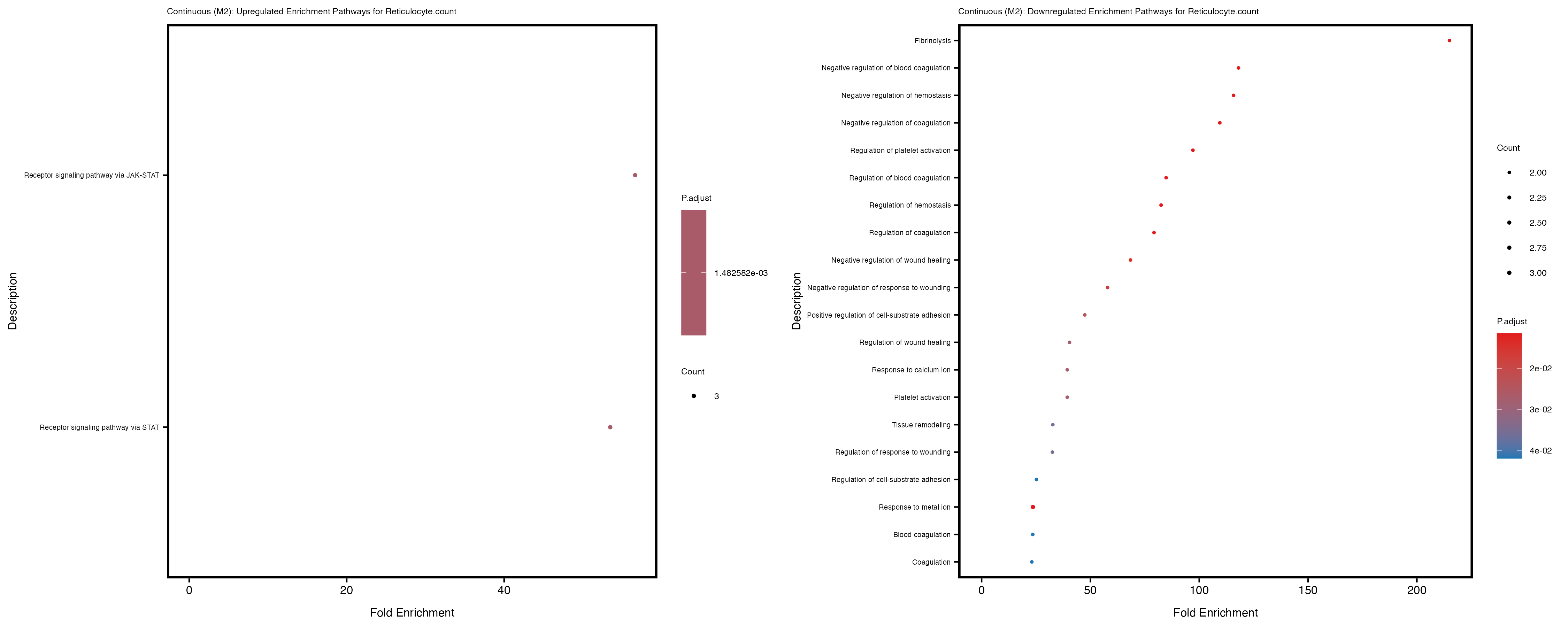
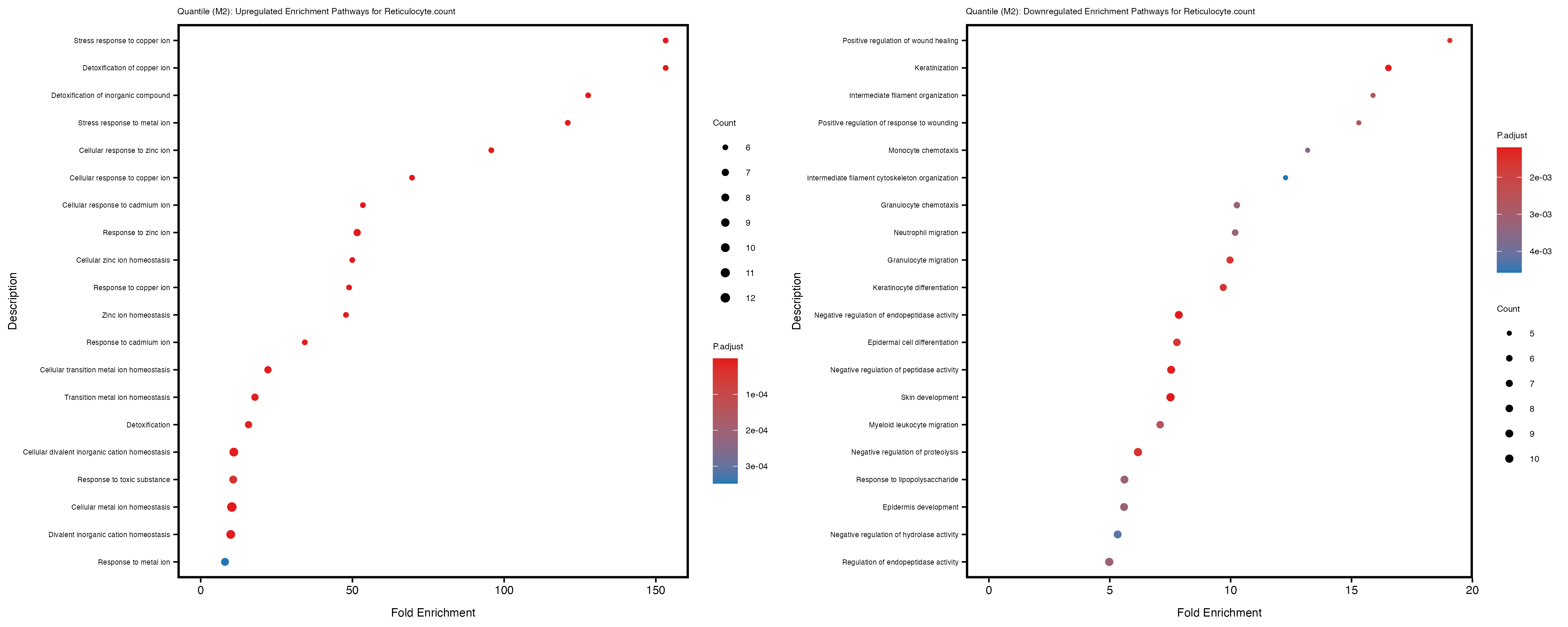
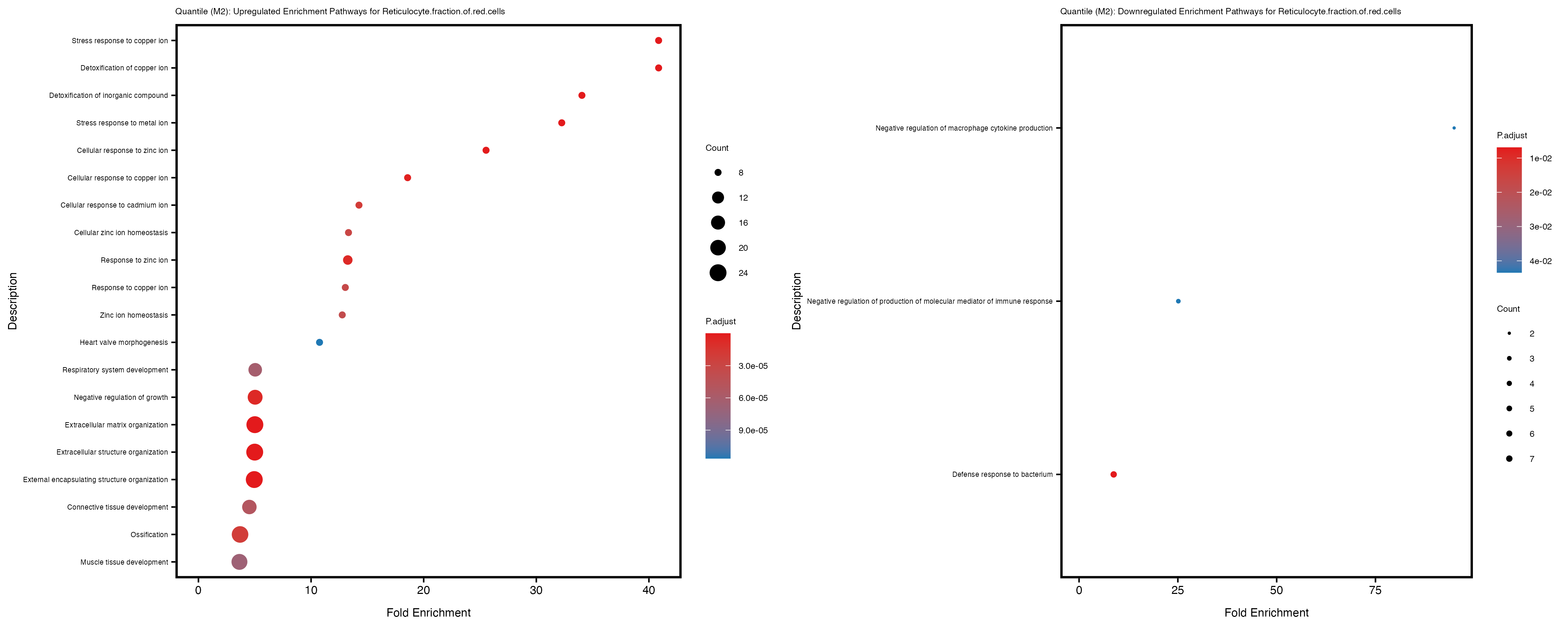
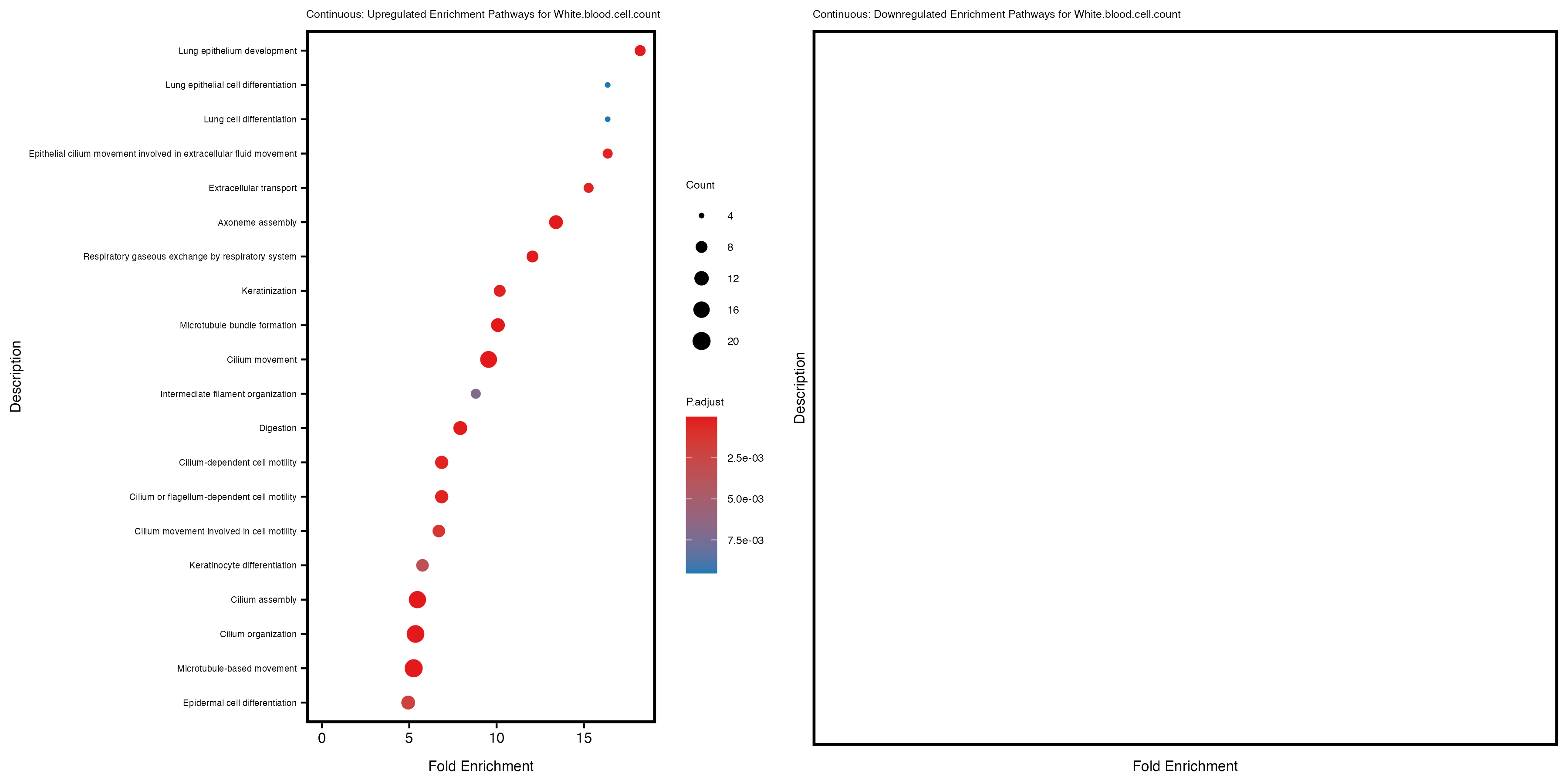
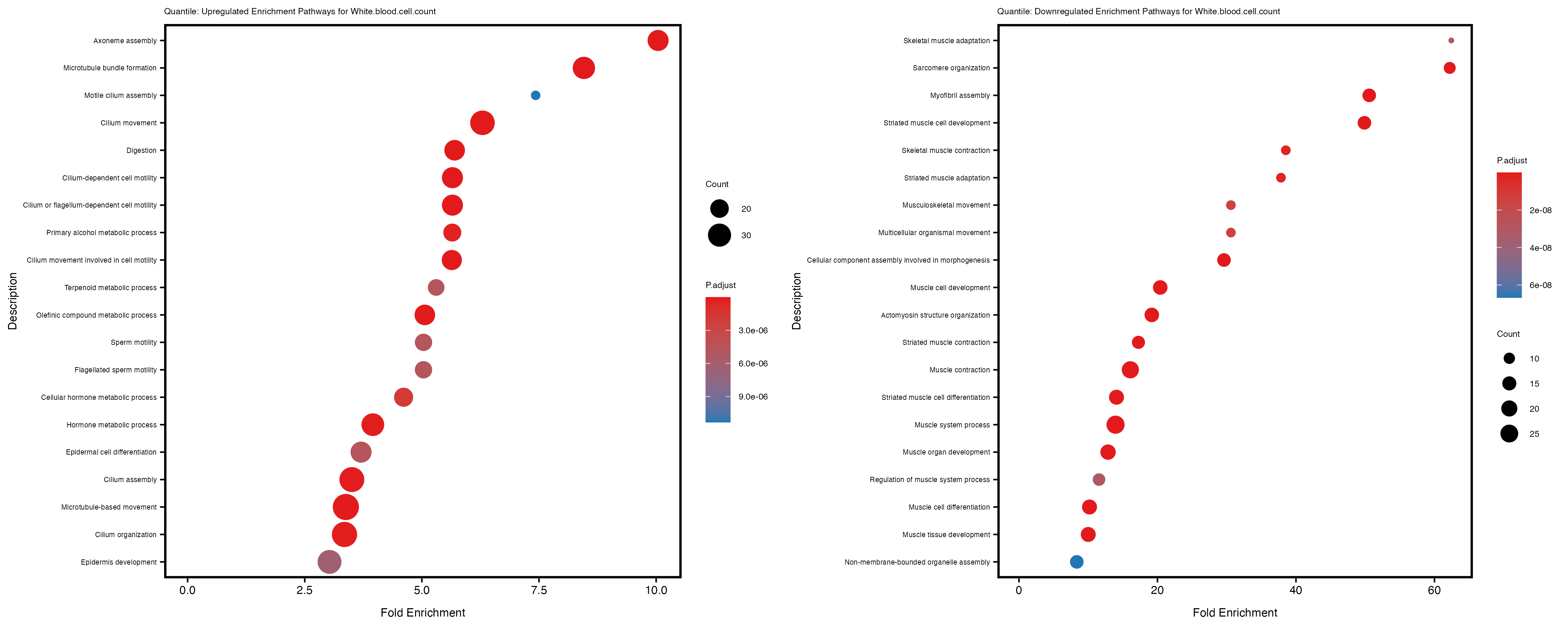
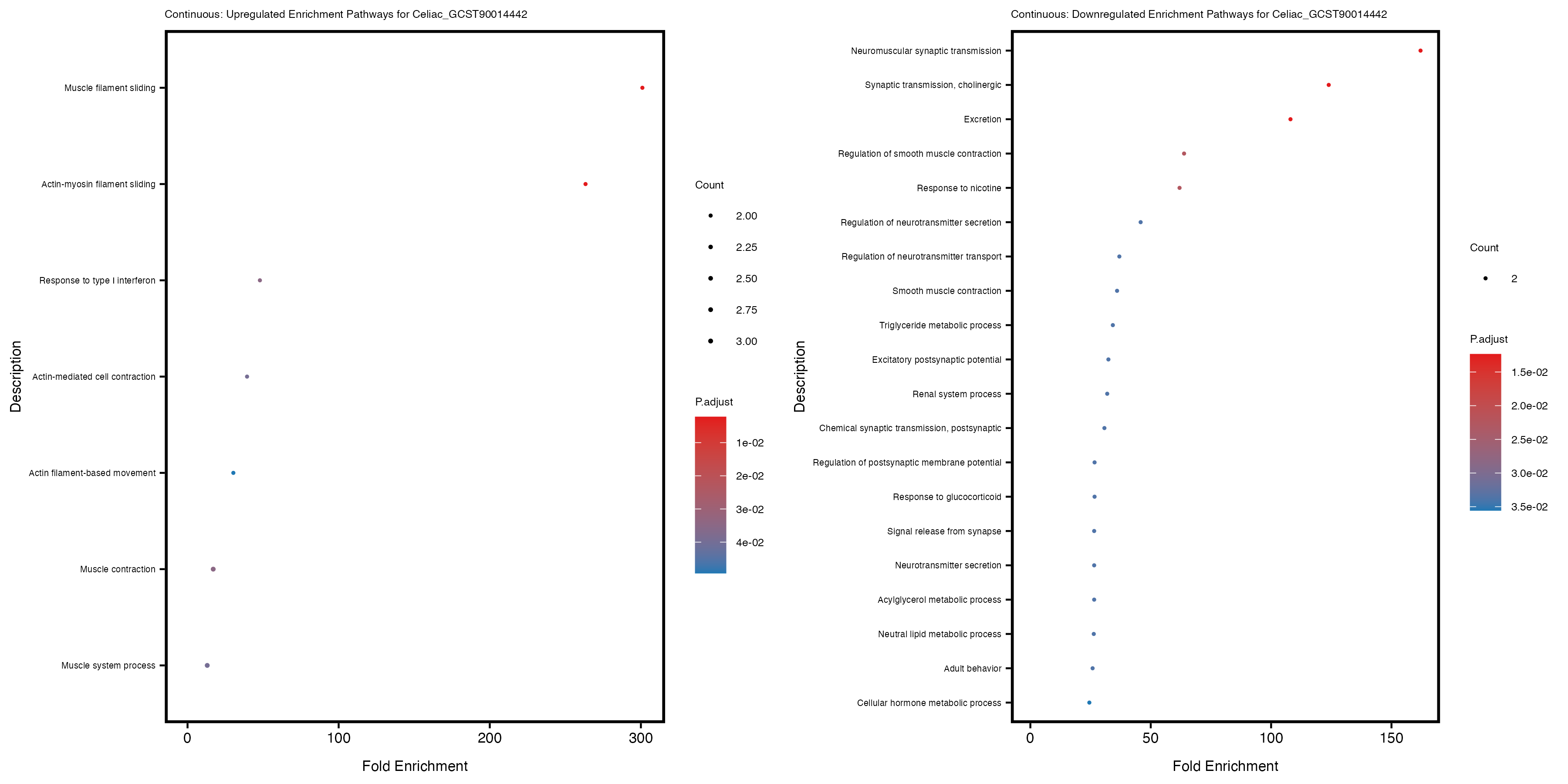
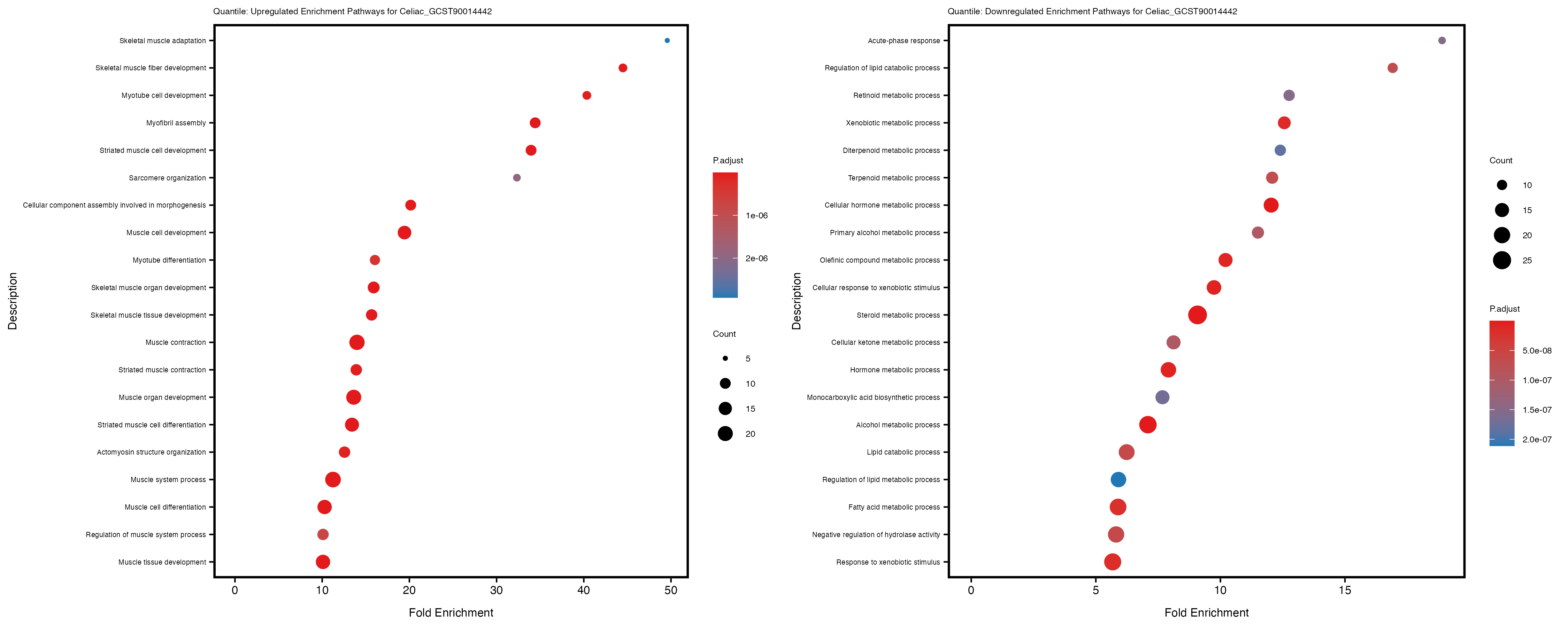
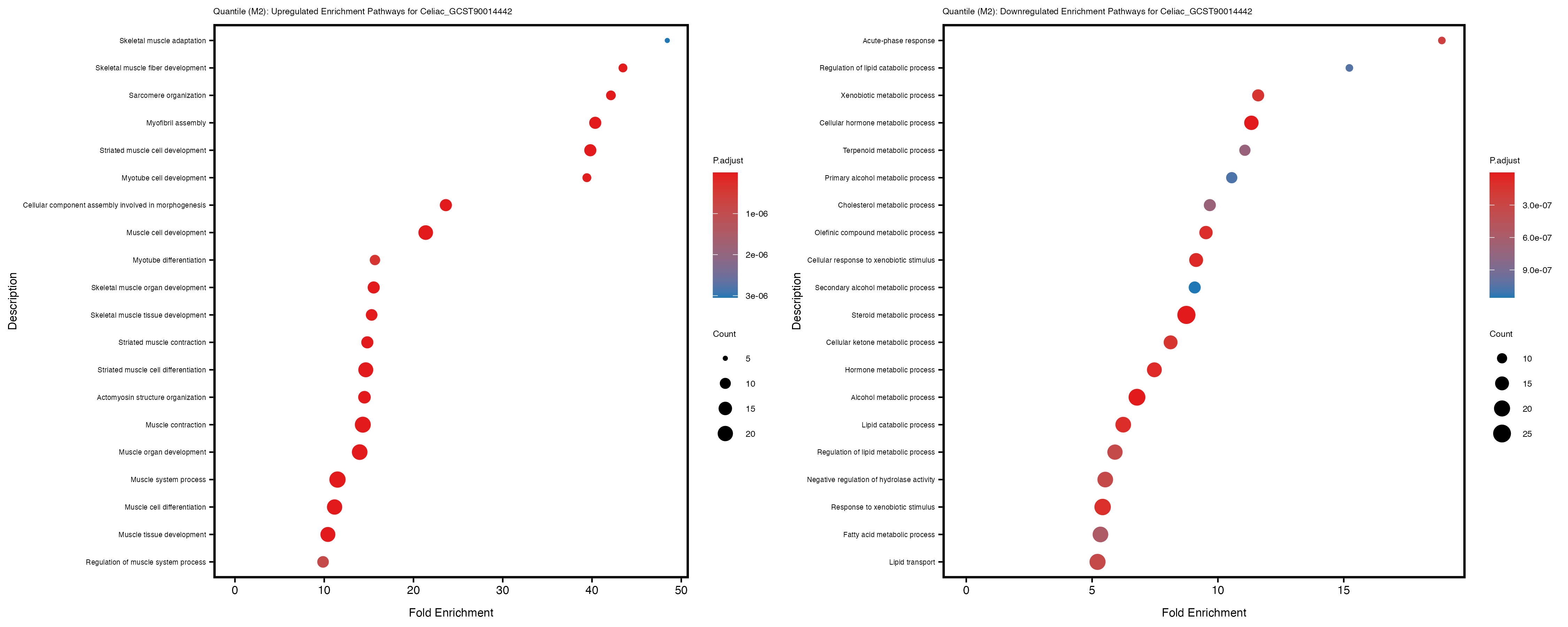
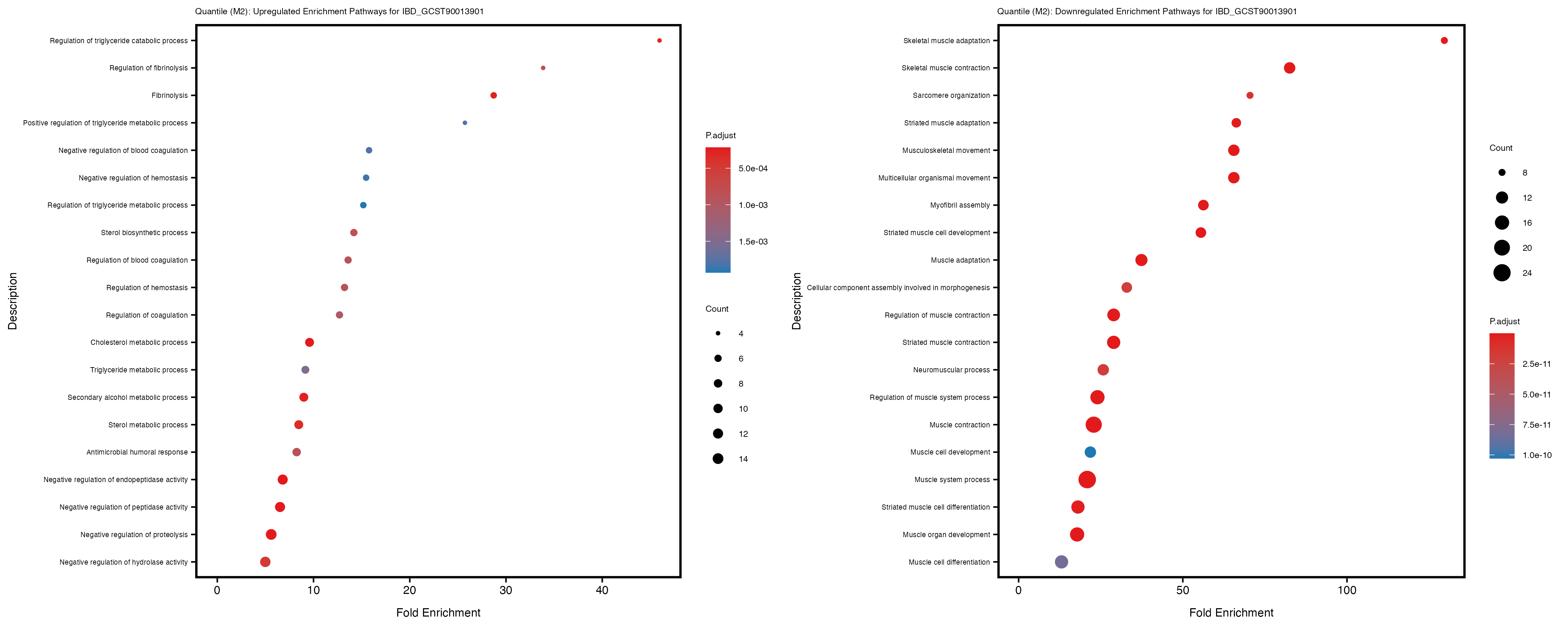
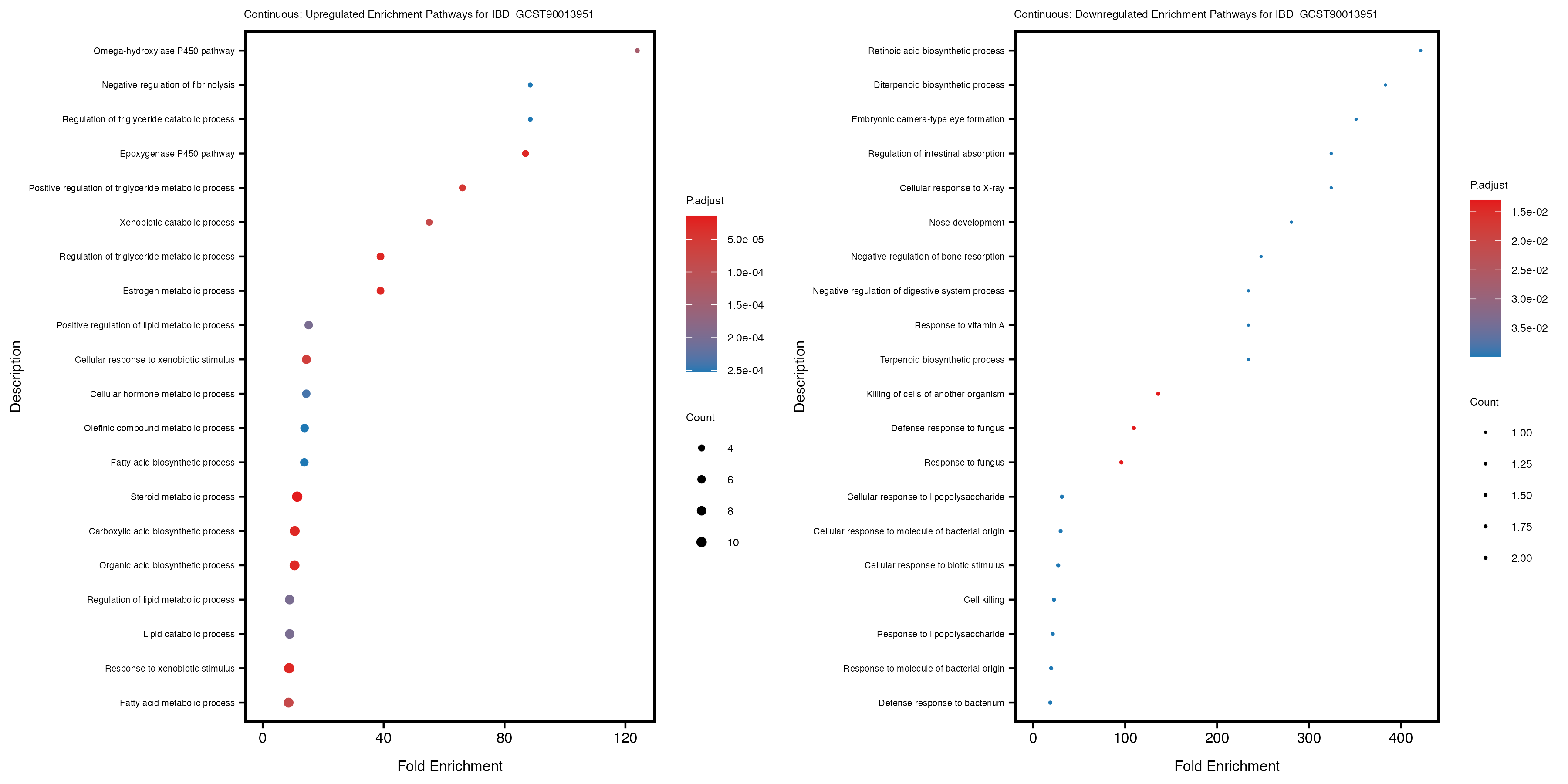
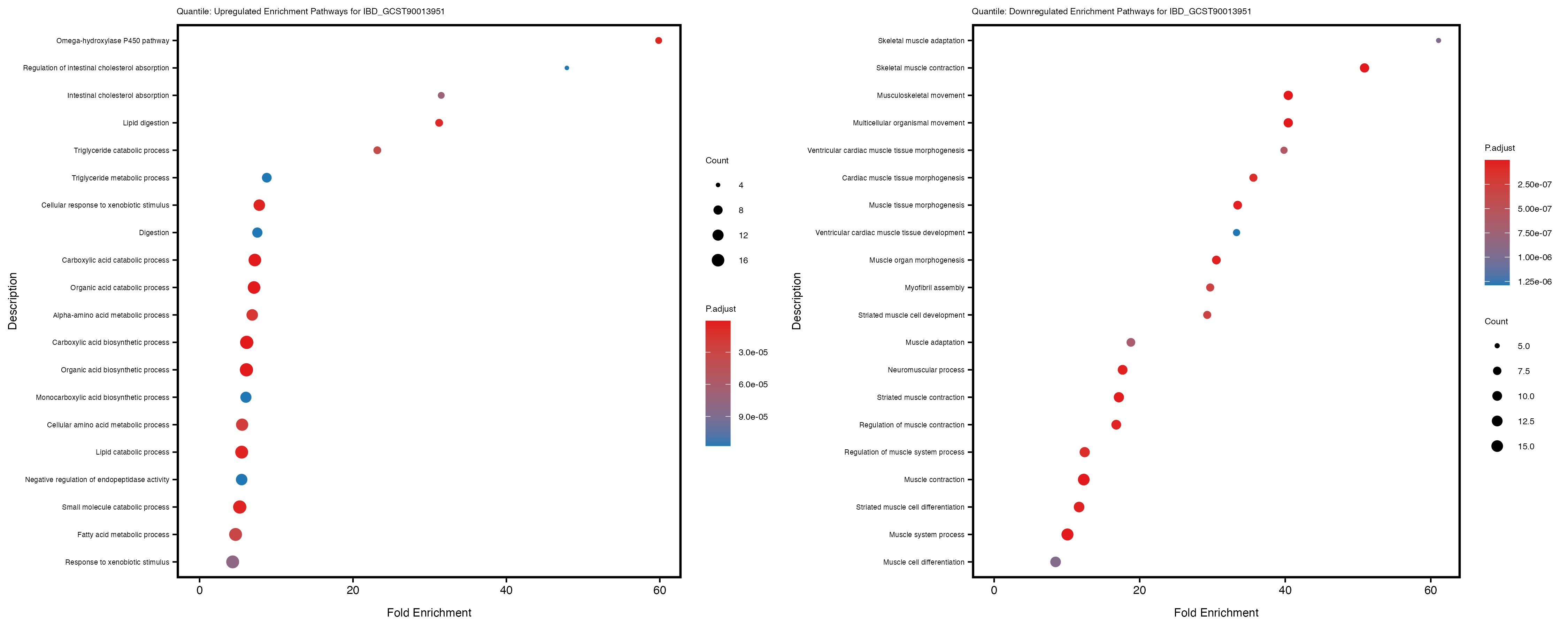
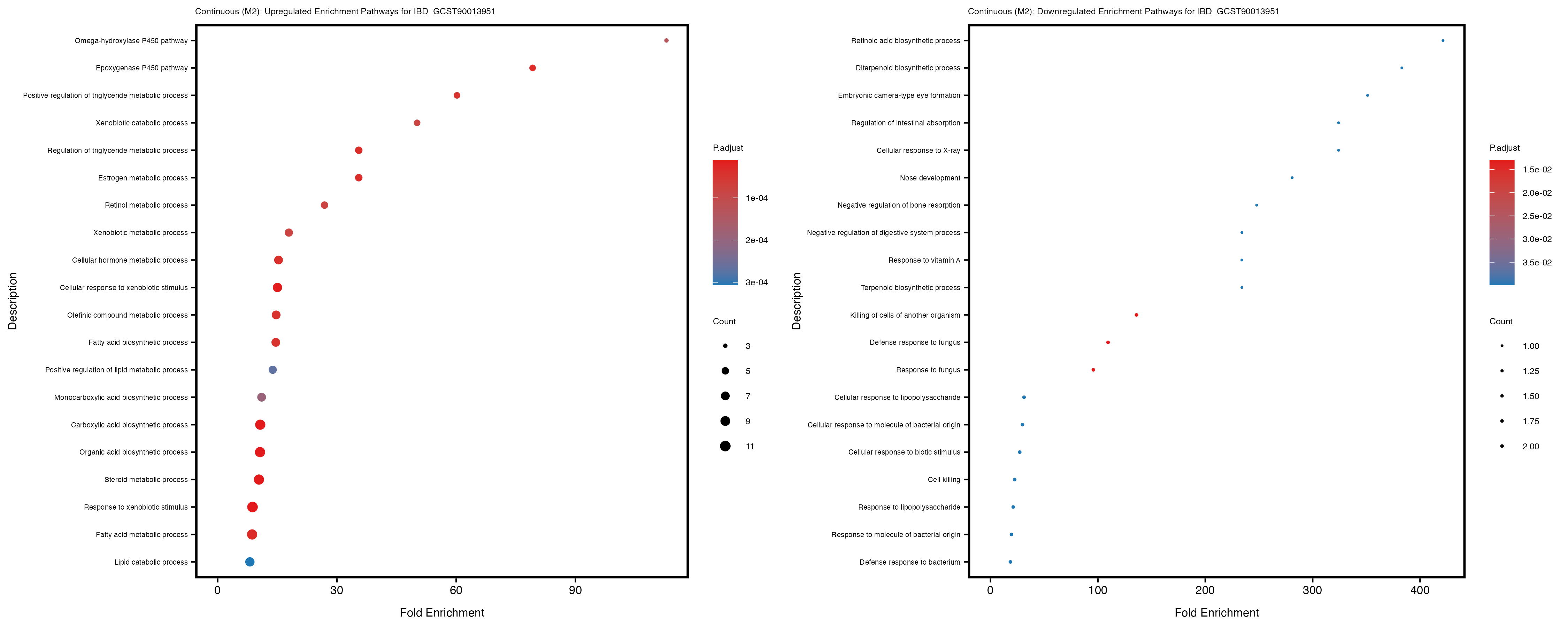
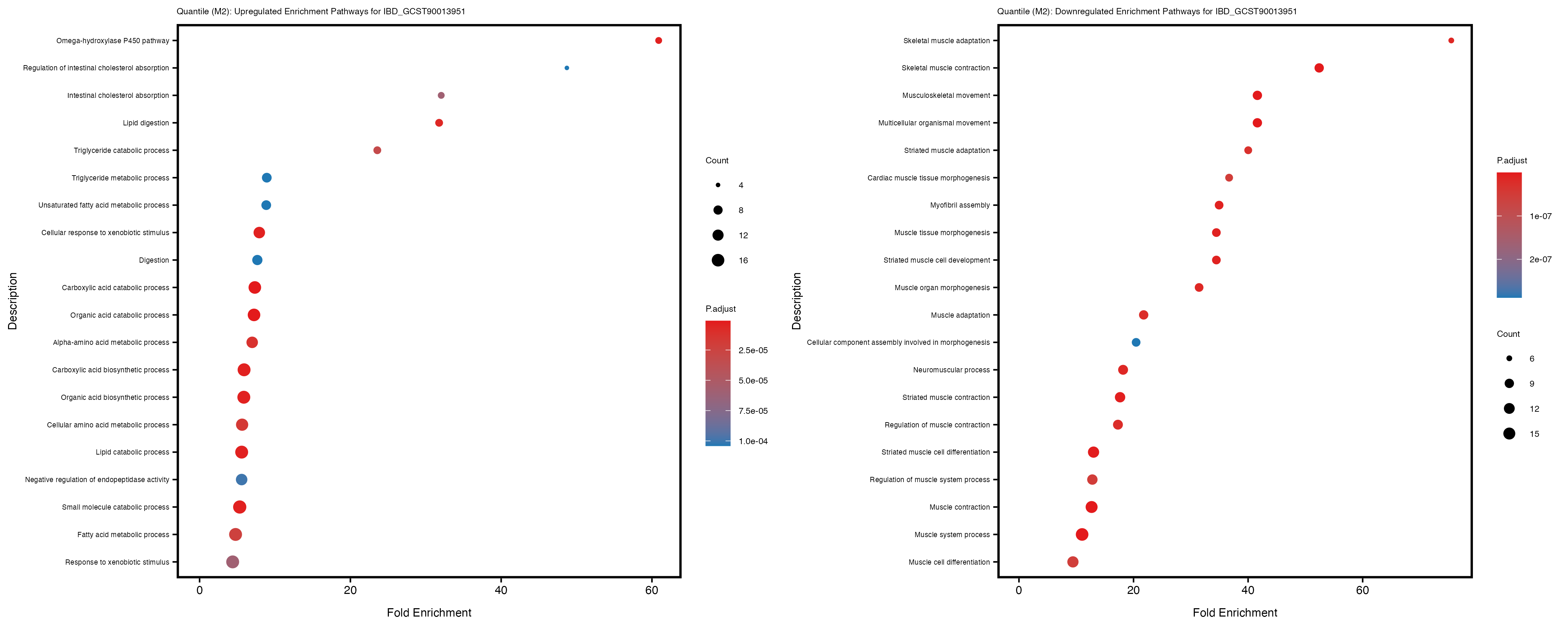
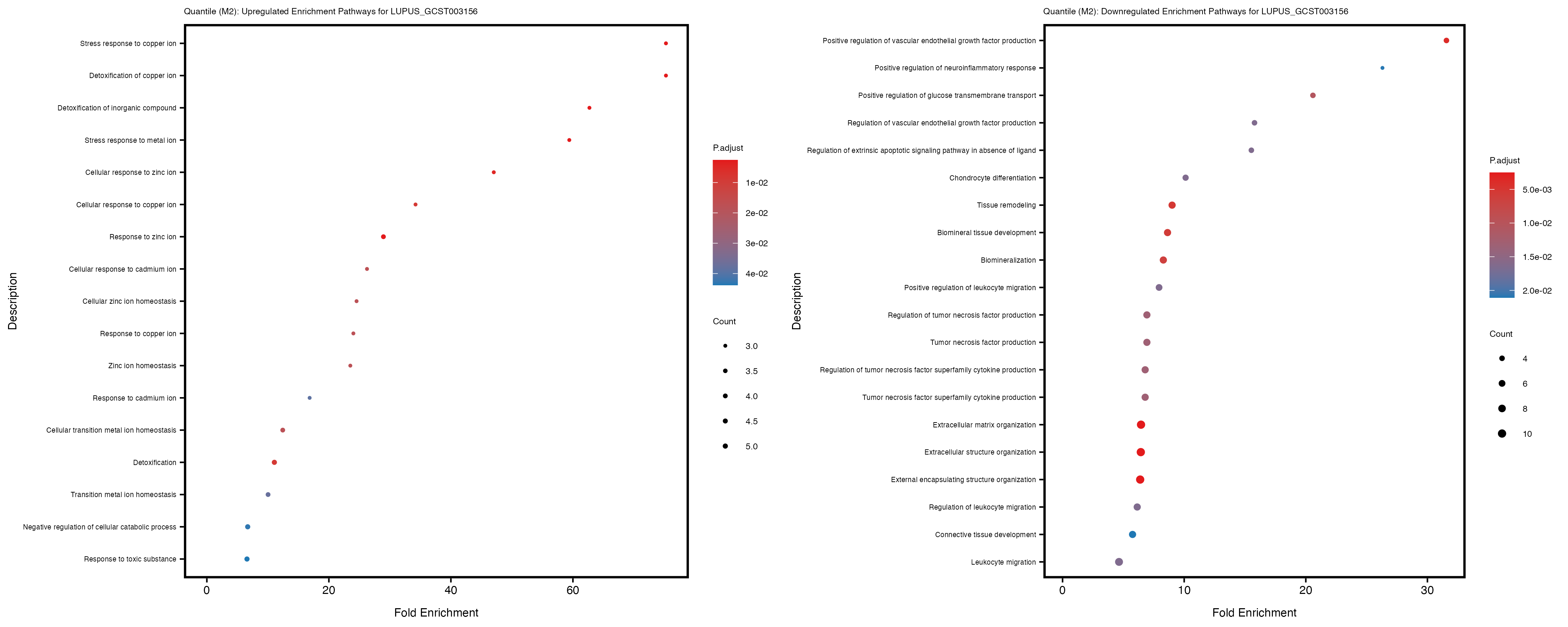
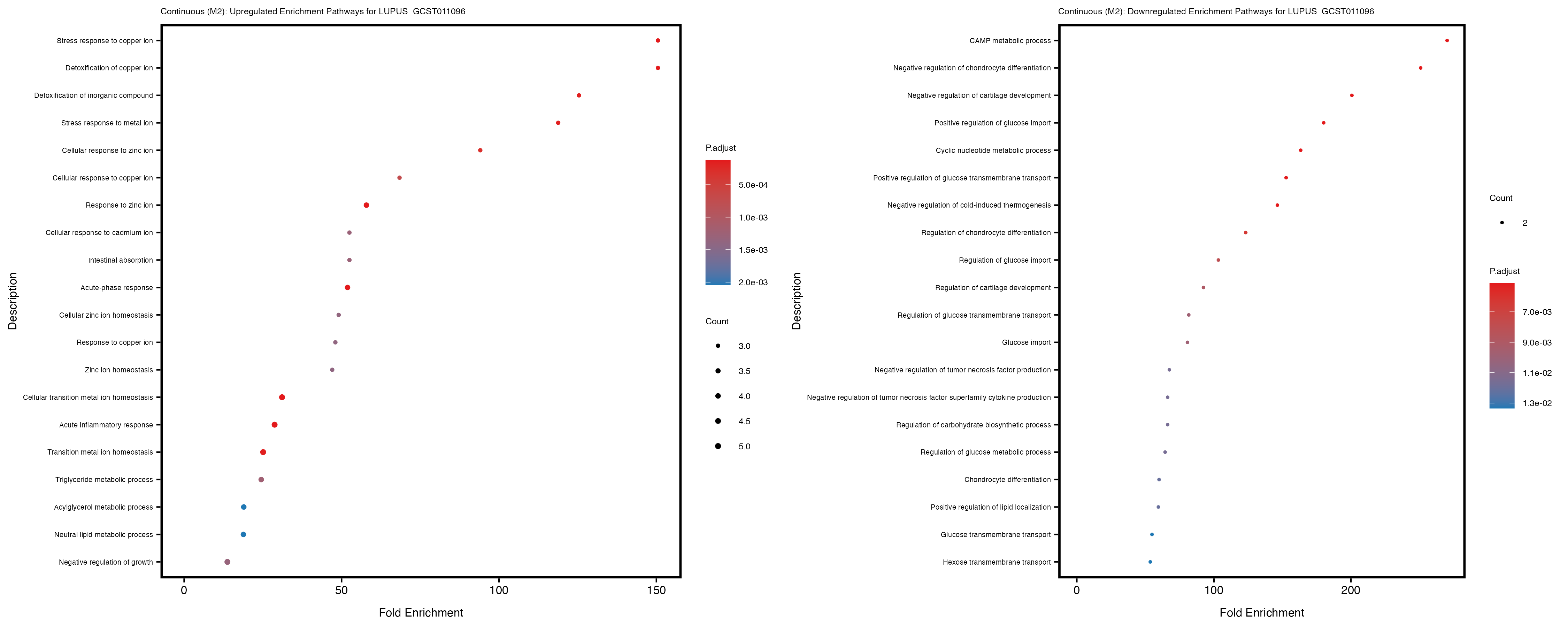
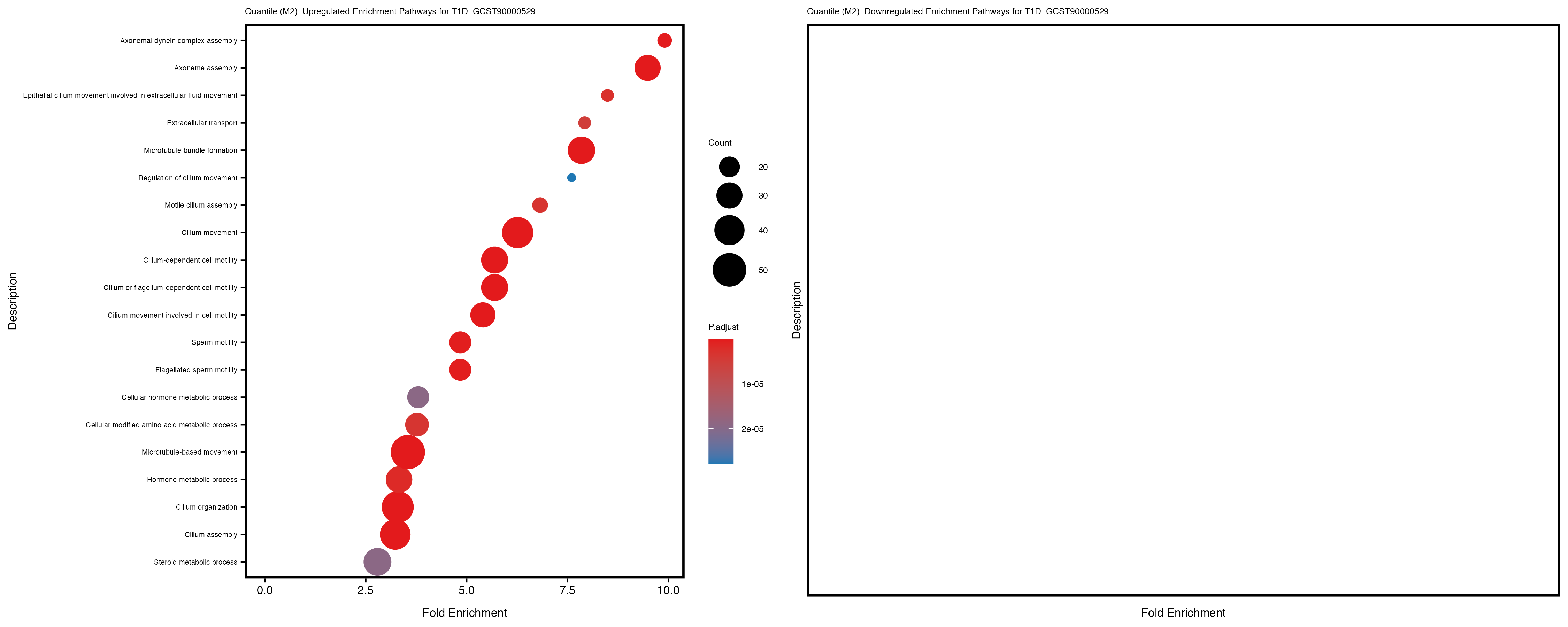
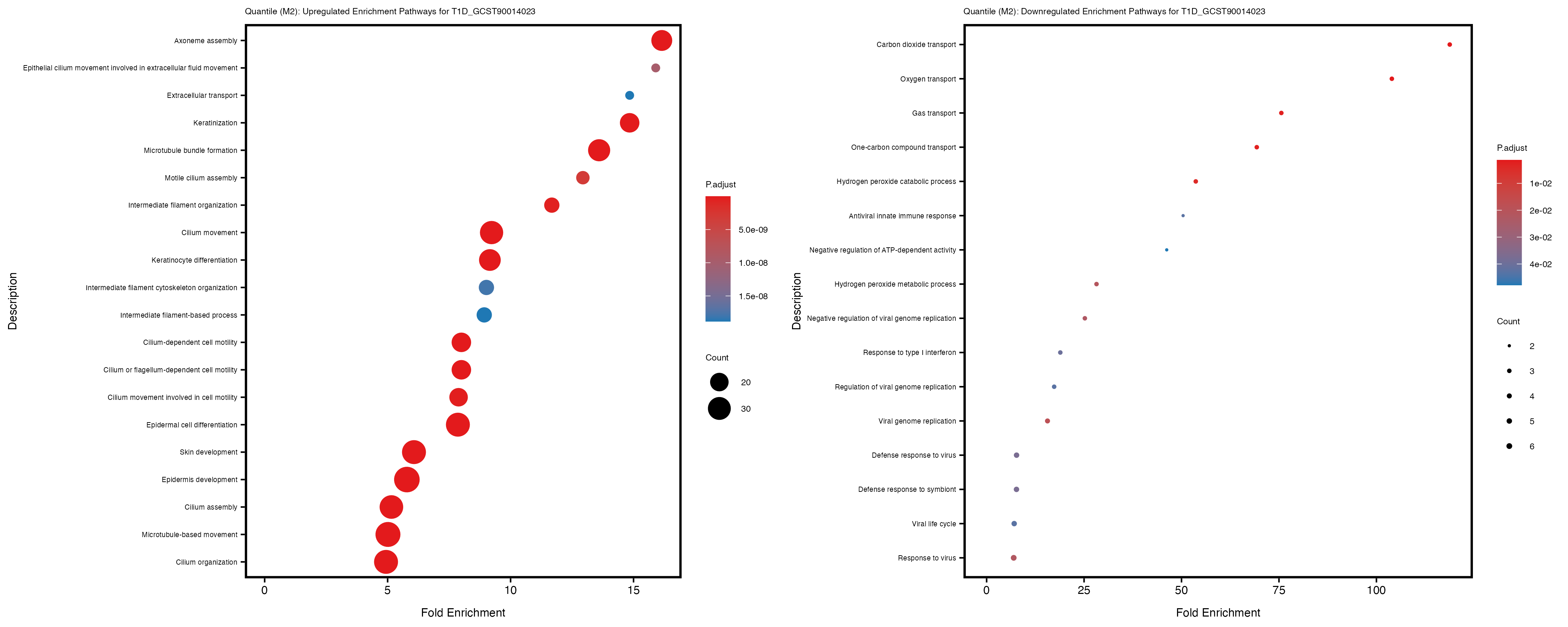
Additionally, the GO enrichment analysis provides insights into the biological pathways and processes that are potentially involved in these traits. BgRatio is the ratio of the total number of genes in a specific pathway (GO term) to the total number of genes. GeneRatio is the ratio of the number of genes enriched in specific pathway to the number of genes that were tested in the analysis. FoldEnrichment is the ratio of GeneRatio to BgRatio, showing how much more enriched a pathway is in your genes of interest compared to the total gene set. Higher FoldEnrichment values indicate that a particular GO term is overrepresented in your gene set compared to the general gene pool, suggesting it may be biologically relevant for the trait. Some traits show higher pathway enrichment in the quantile analysis, indicating that stratification by PRS may provide more biologically meaningful results.
For highly correlated traits, they tend to share similar GO enriched pathways. For example, mean corpuscular hemoglobin and mean corpucular volume are both involved in read blood cell biology, so it’s expected that their GO enrichment would point to similar biological processes, such as hemoglobin metabolism, iron homeostasis and erythropoiesis.
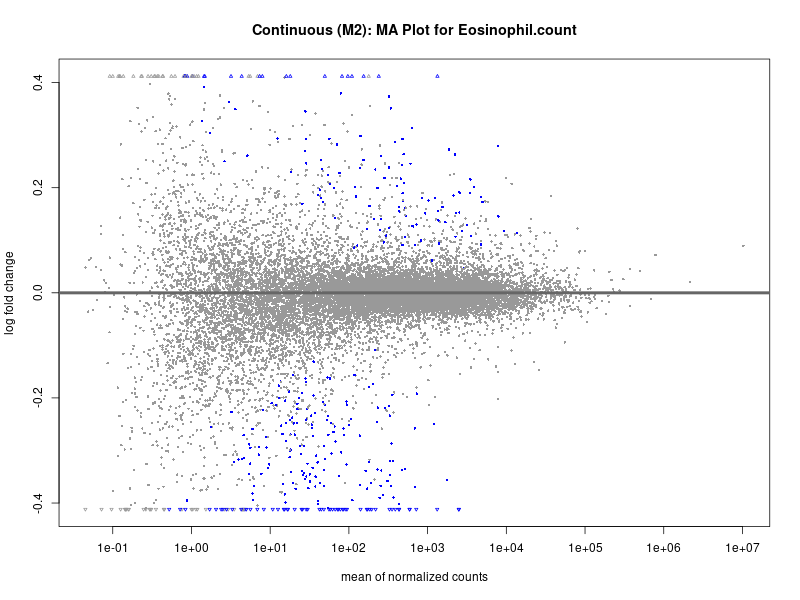
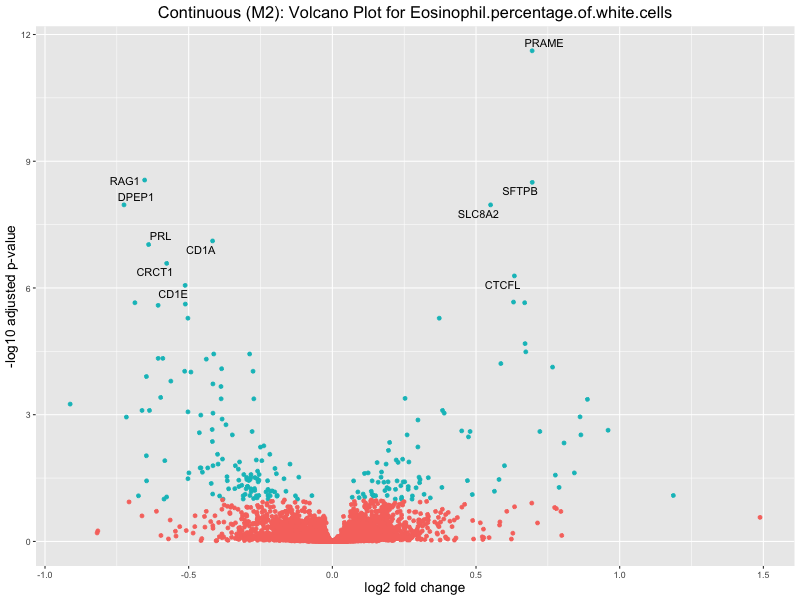
To account for population stratification, we adjusted the model by including the first two genotype principal components (PCs), as they explain most of the genetic variation in the data. For Eosinophil count, we observed that genes were consistently identified as top differentially expressed with and without adjustment. The only difference is that instead of SLC6A9, WFDC2 was detected as top 10 DEGs when adjusting for the first two genotype PCs. WFBC2 is a protease inhibitor with a potential role in innate immune defense (L Bingle, 2006). Although same genes are identified as top DEGs for continuous analysis, additional pathways such as response to inactivity were highlighted when adjusting for the first two genotype PCs.
Basophil count
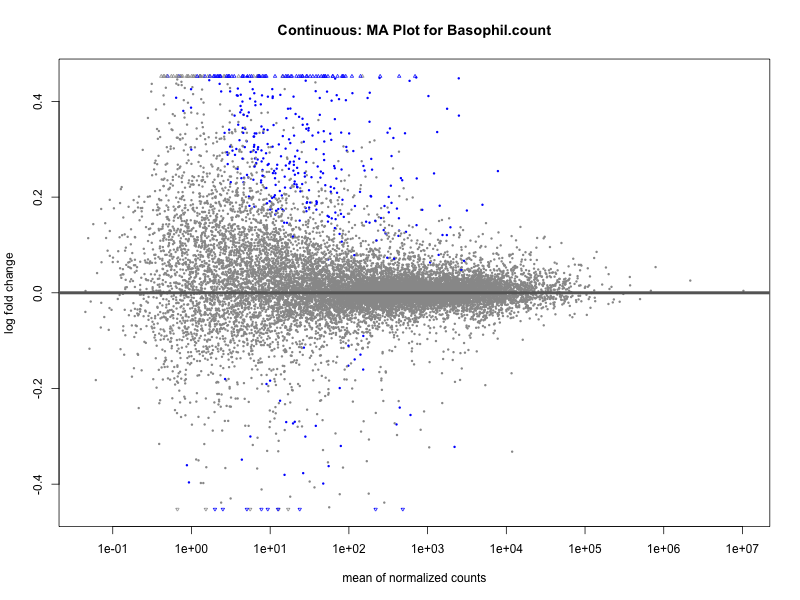
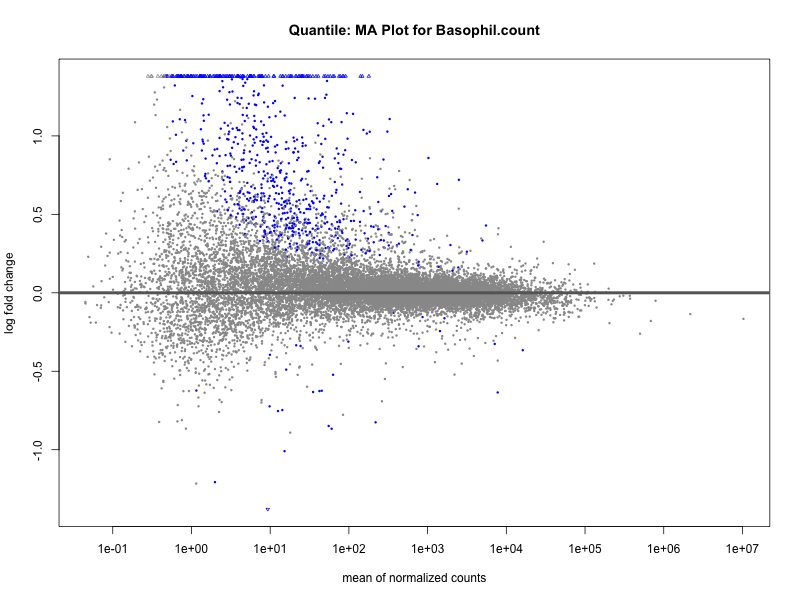
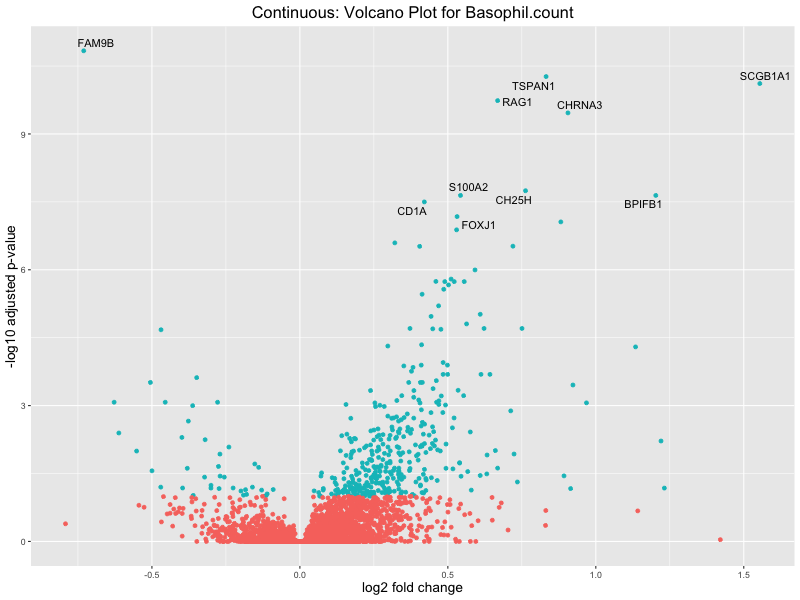
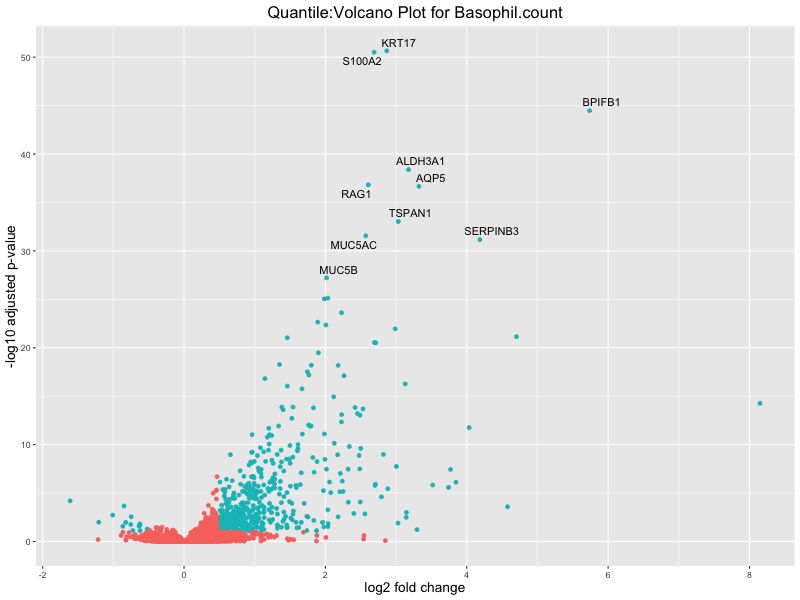
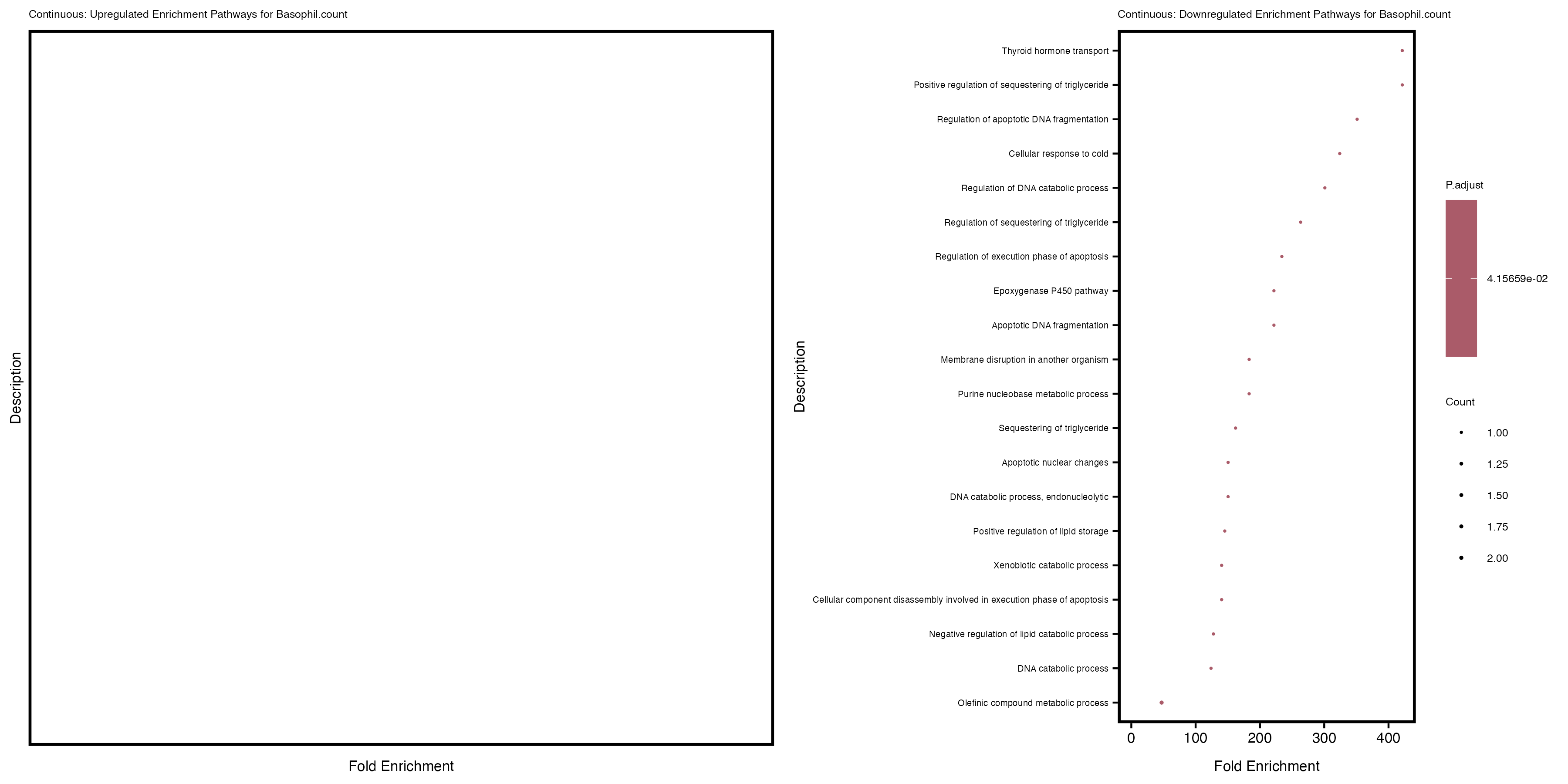
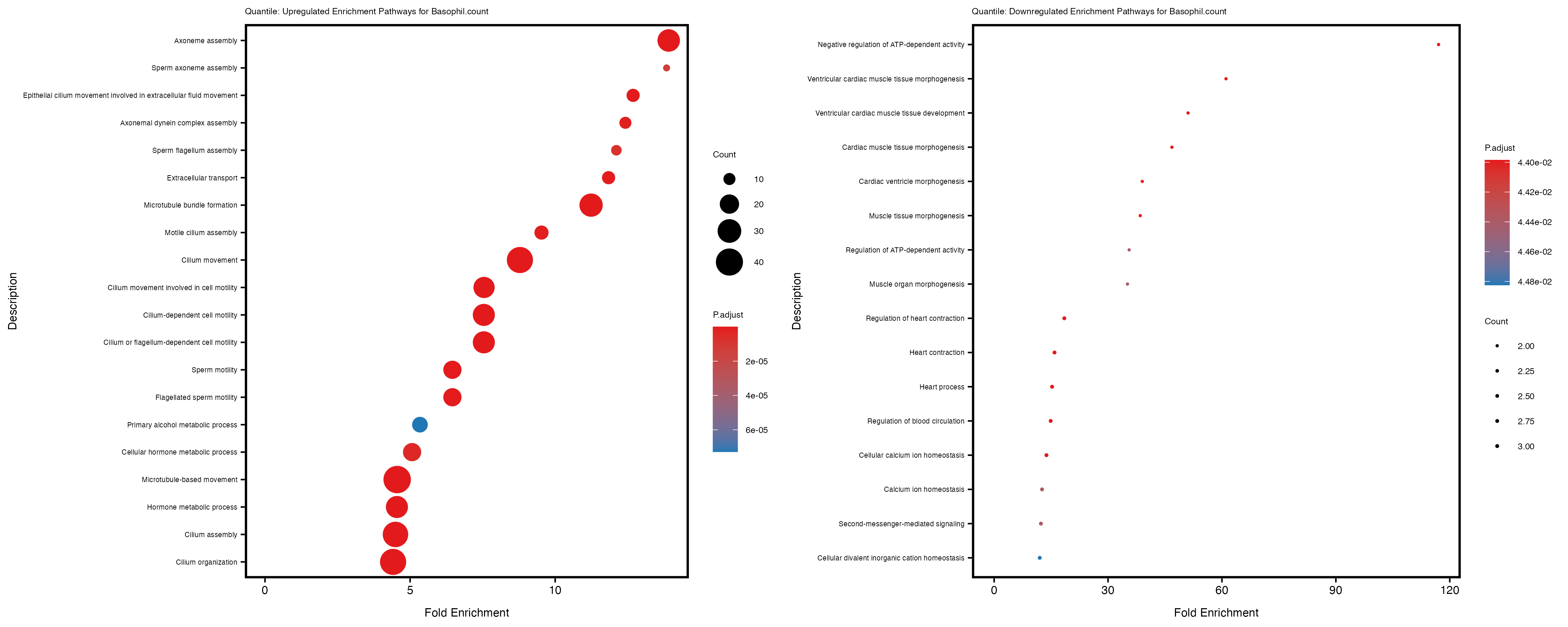
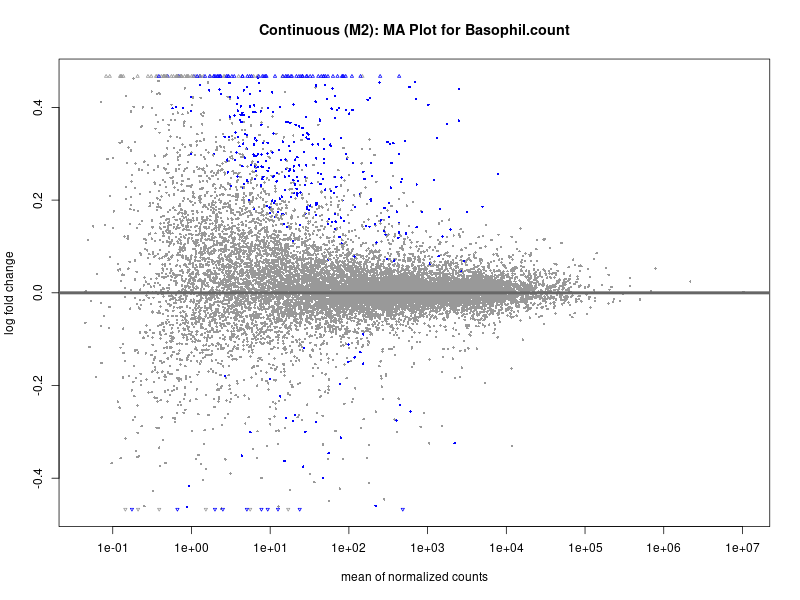
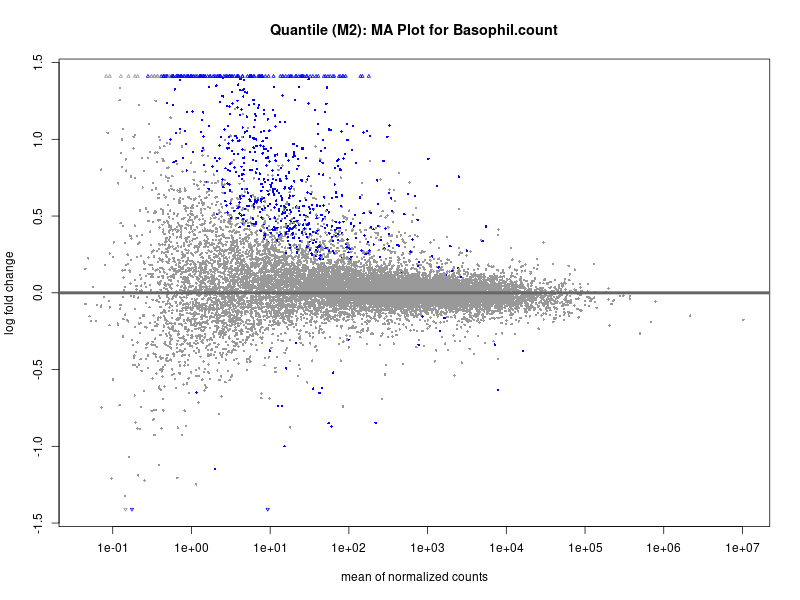
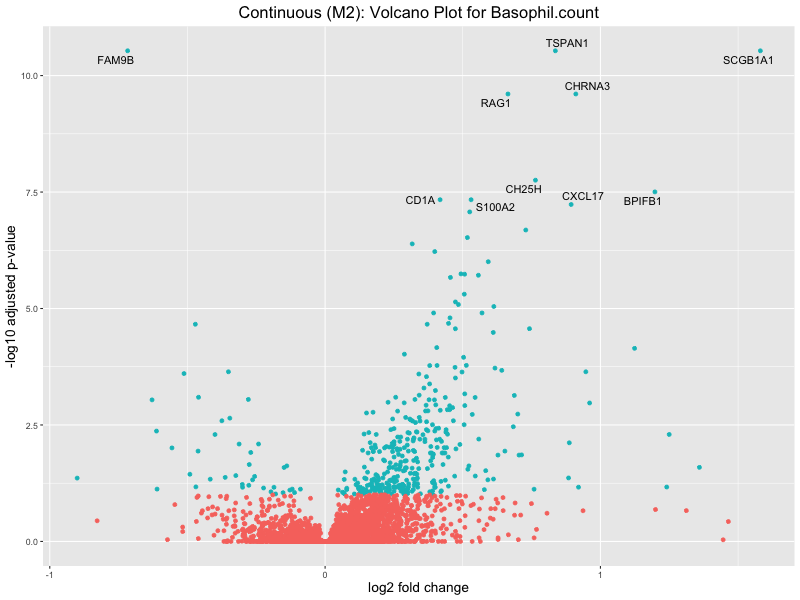
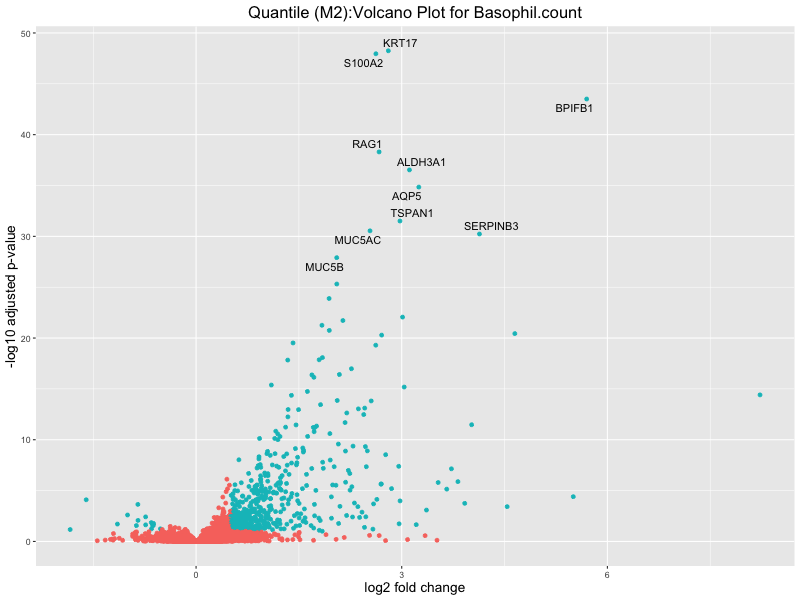
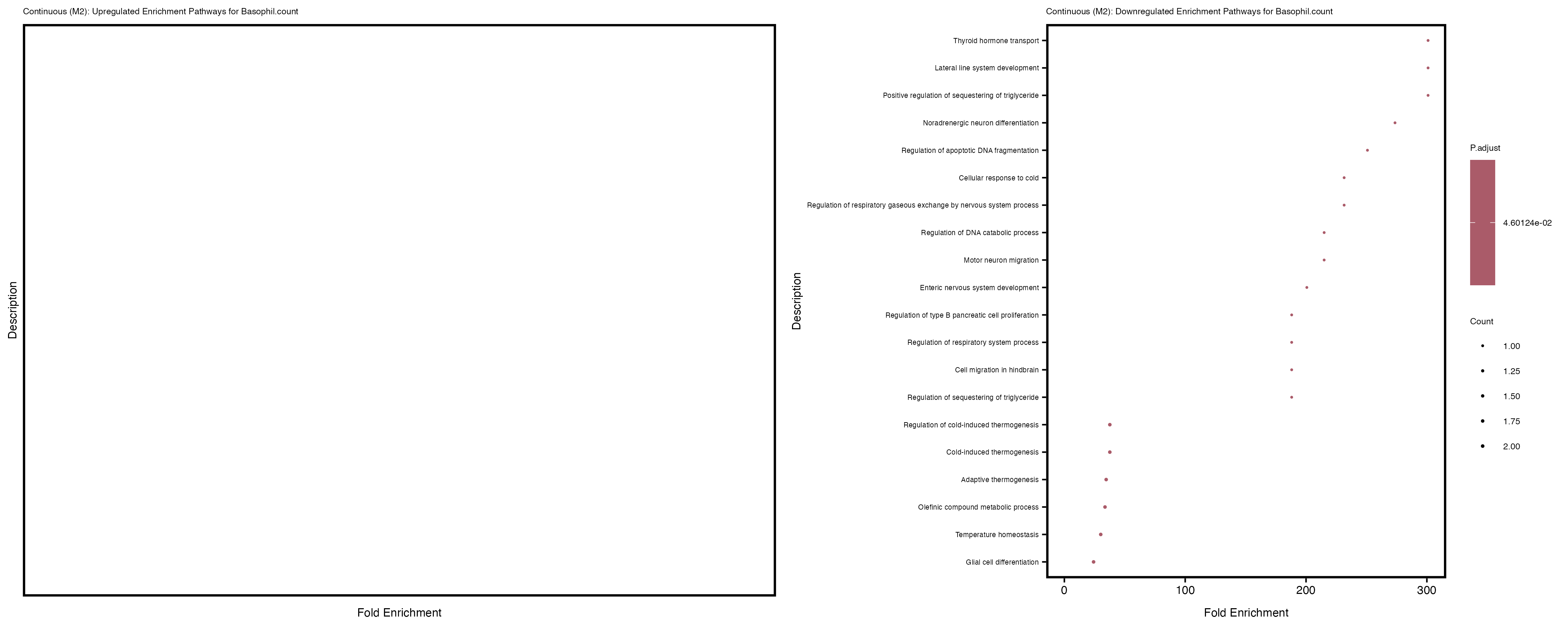
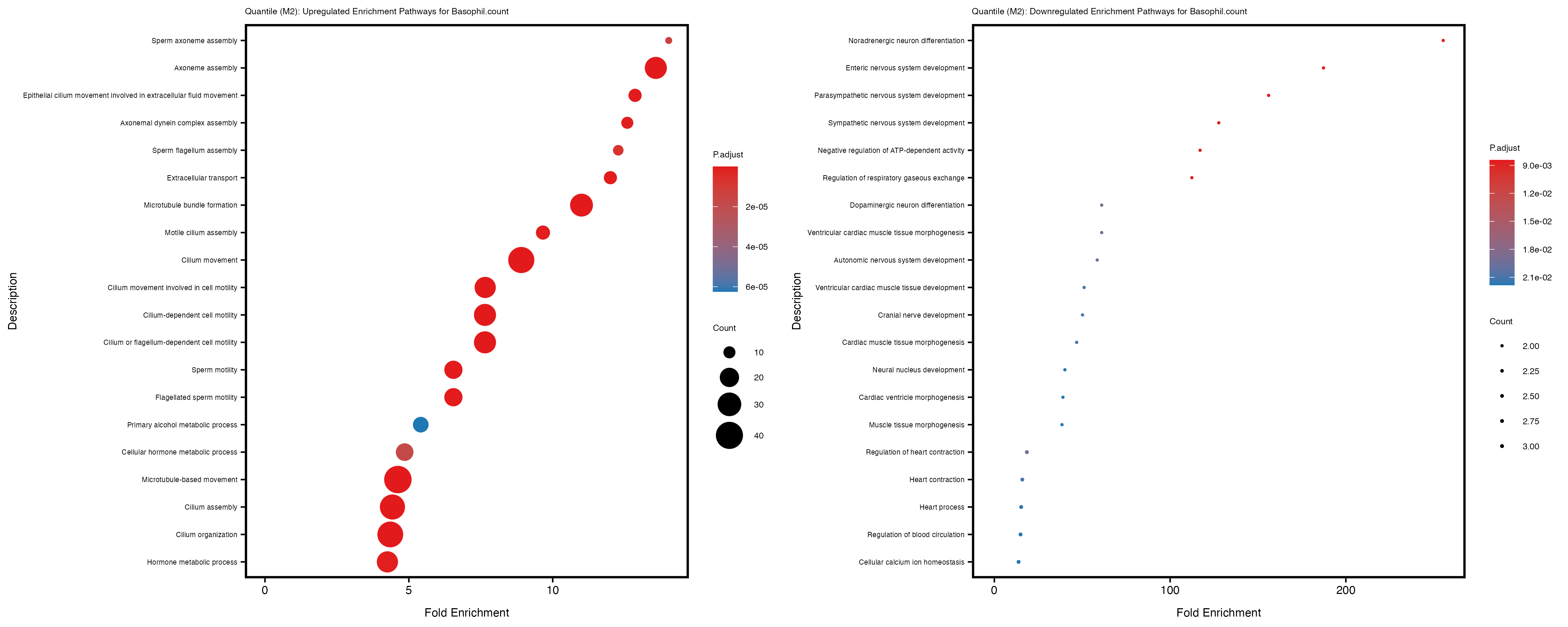
Basophil.percentage.of.white.cells
Eosinophil.count
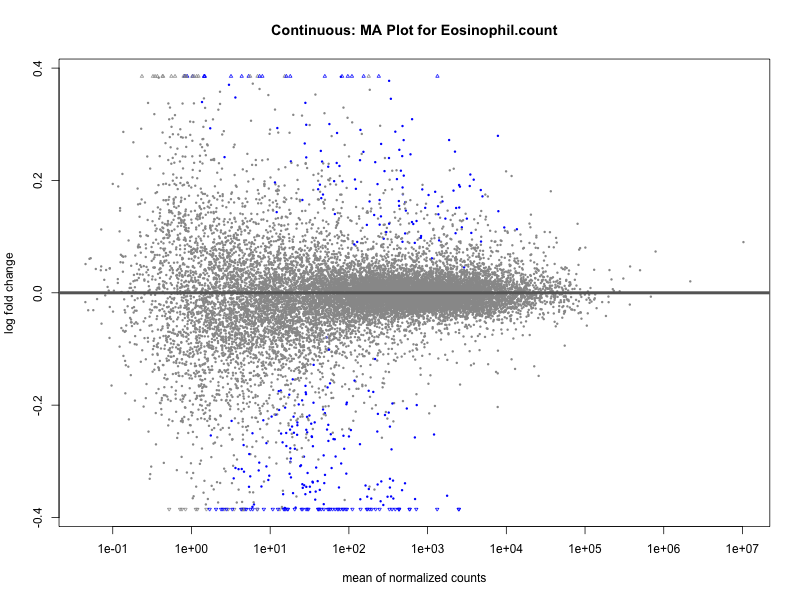
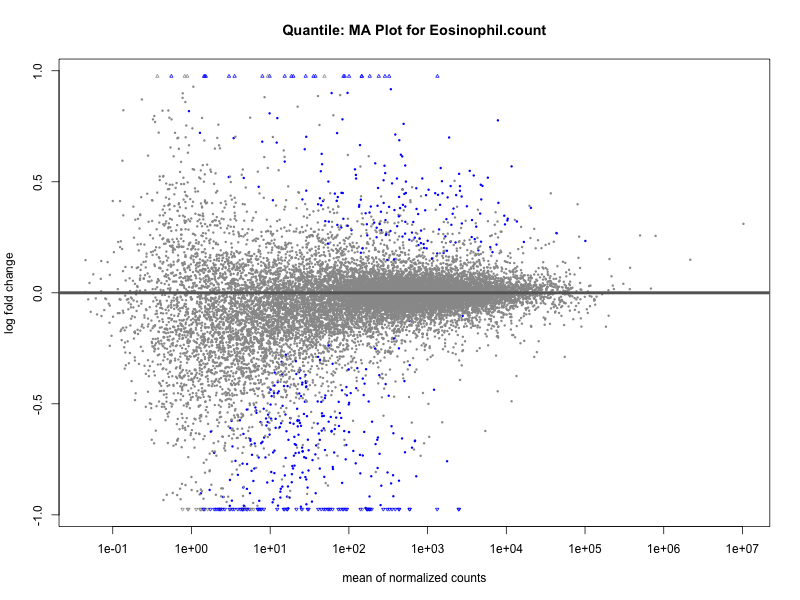
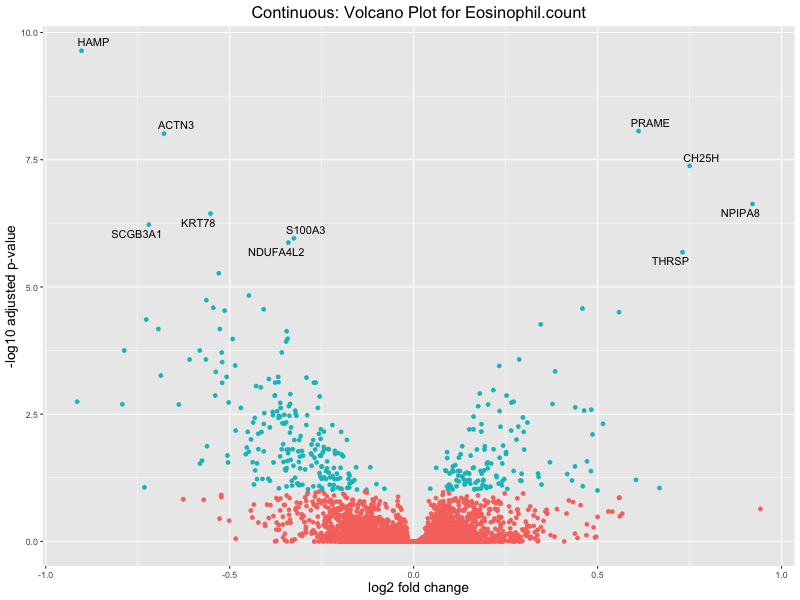
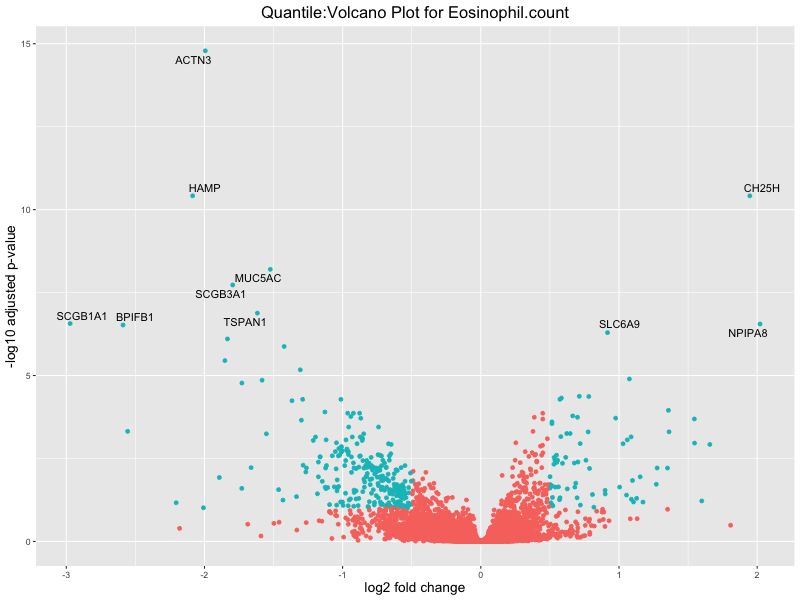
Eosinophil.percentage.of.white.cells
Hematocrit
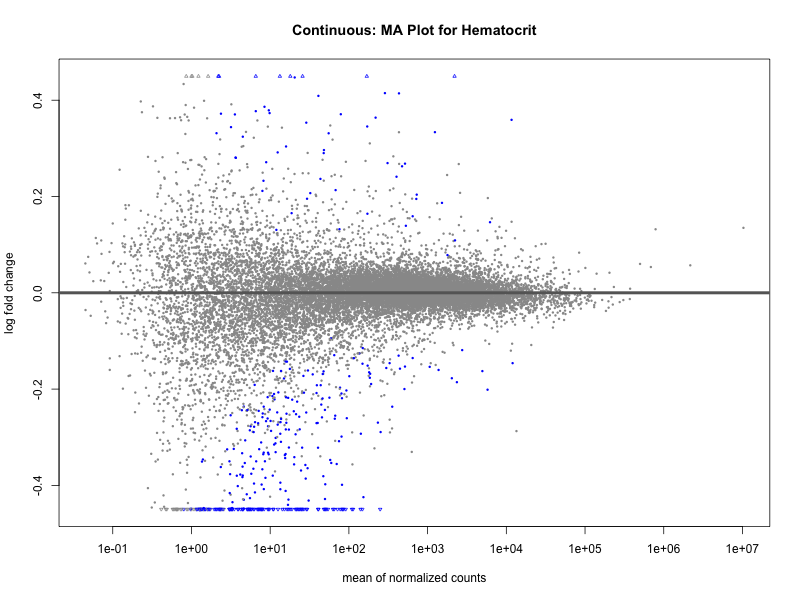
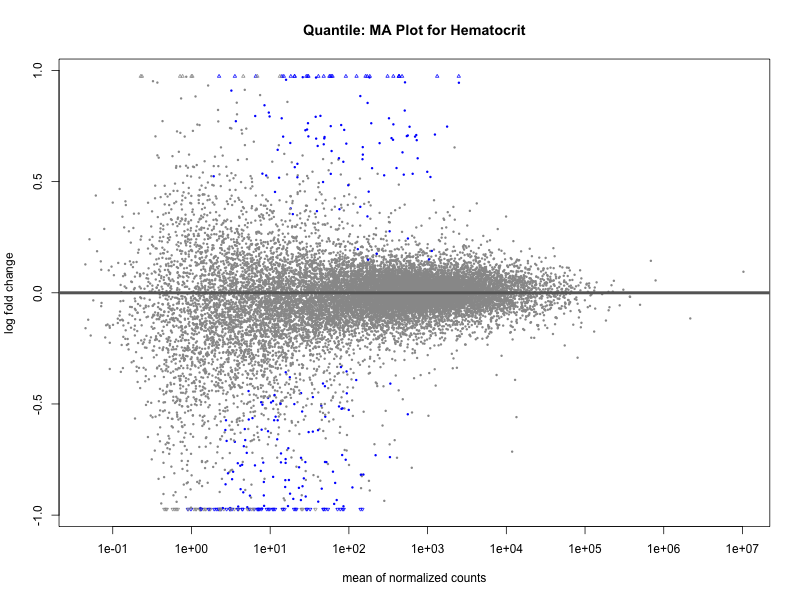
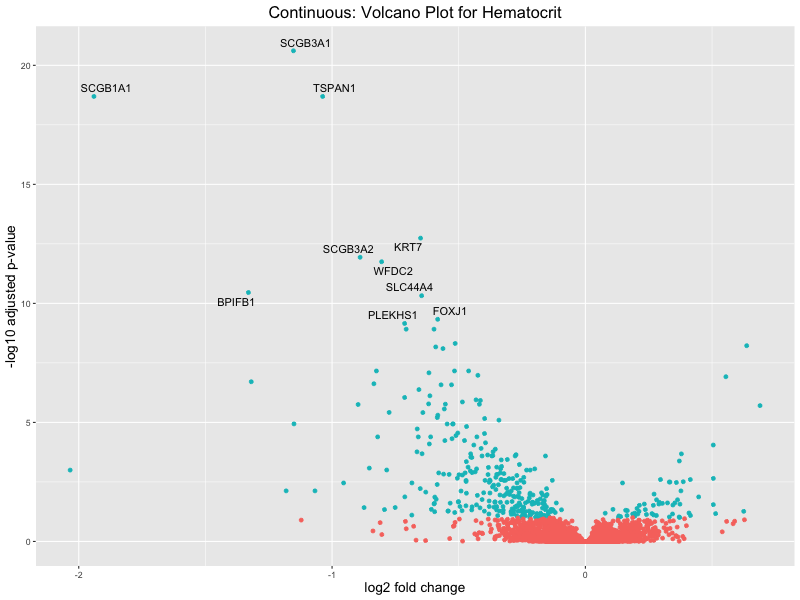
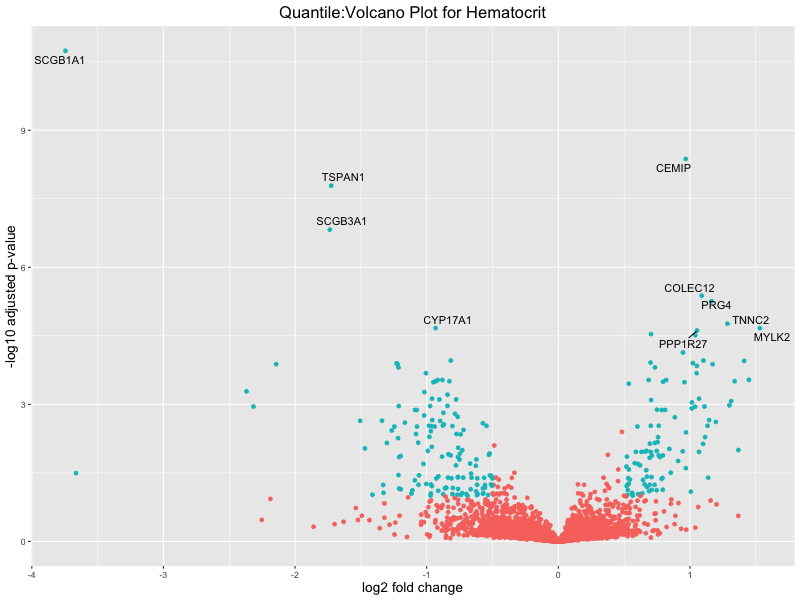
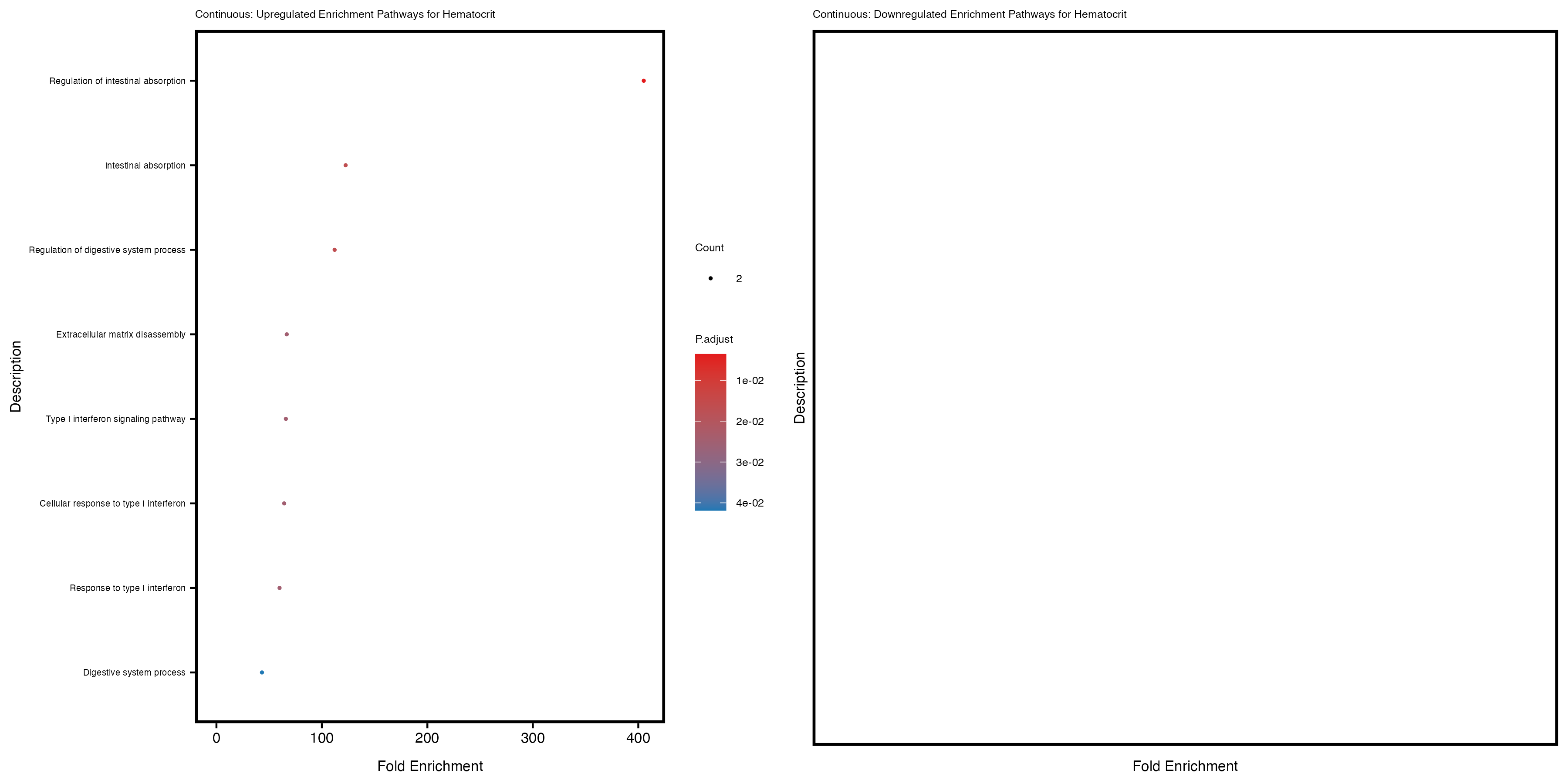
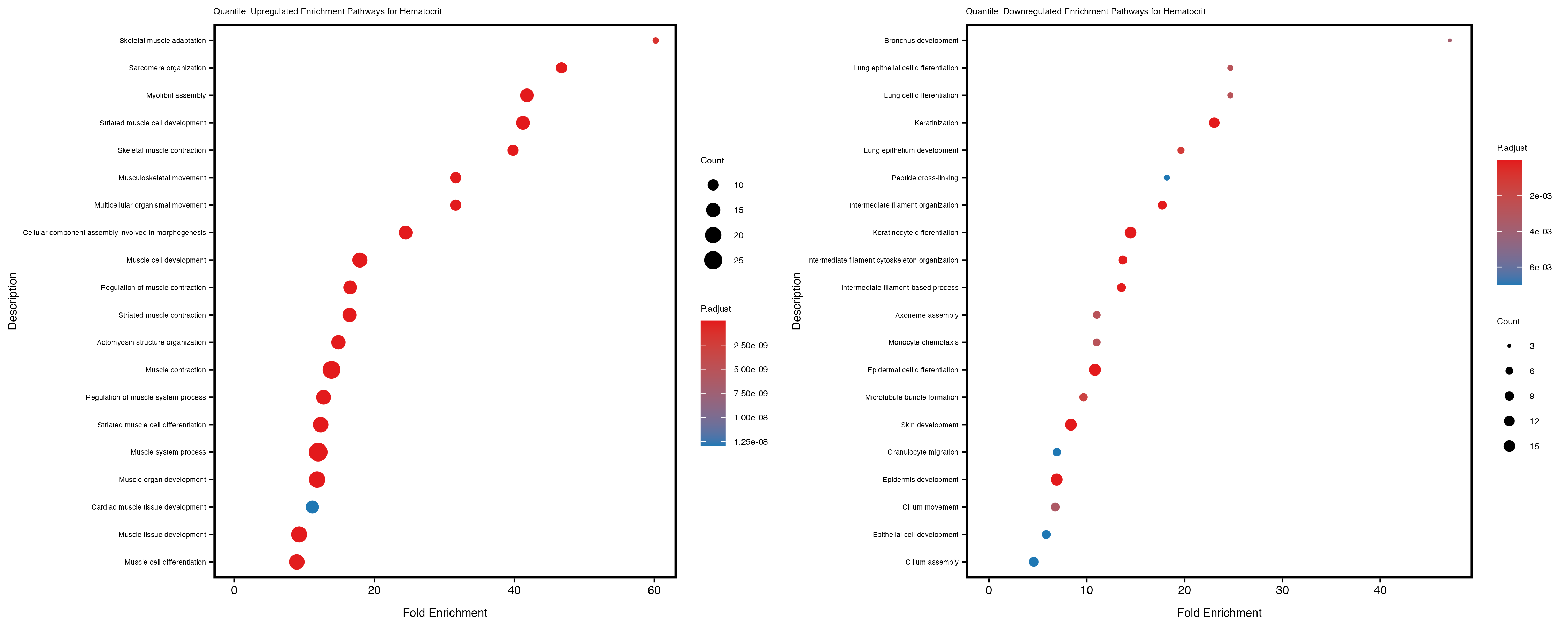
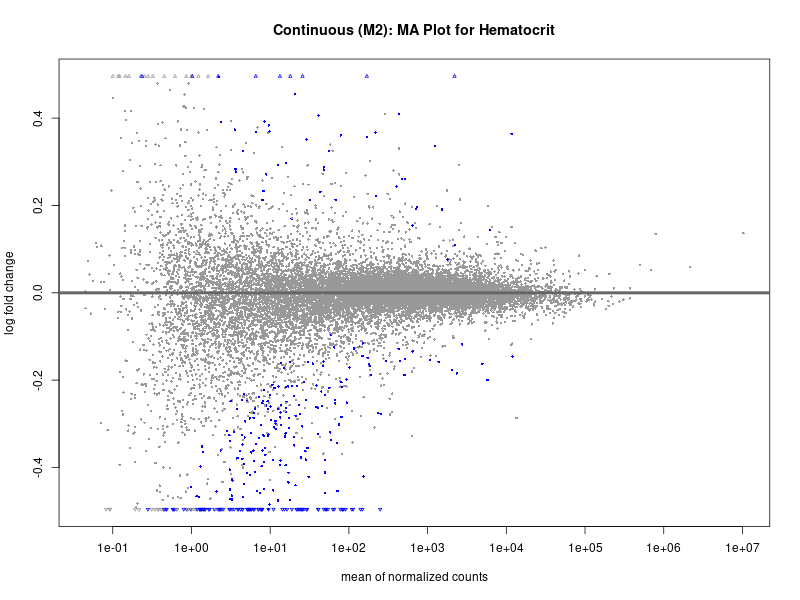
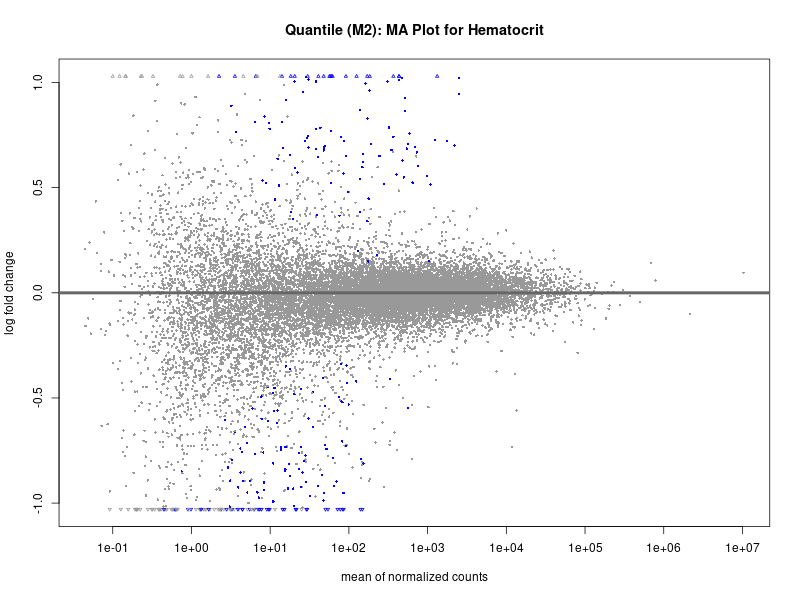
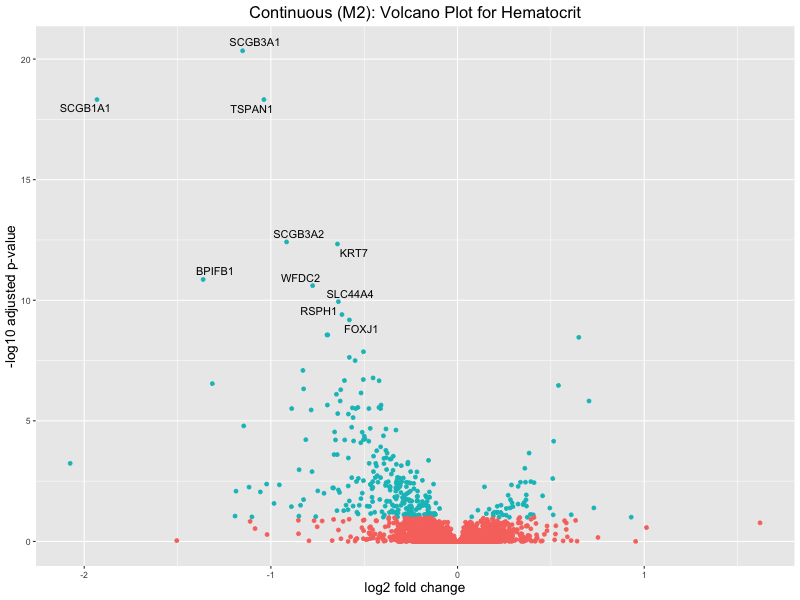
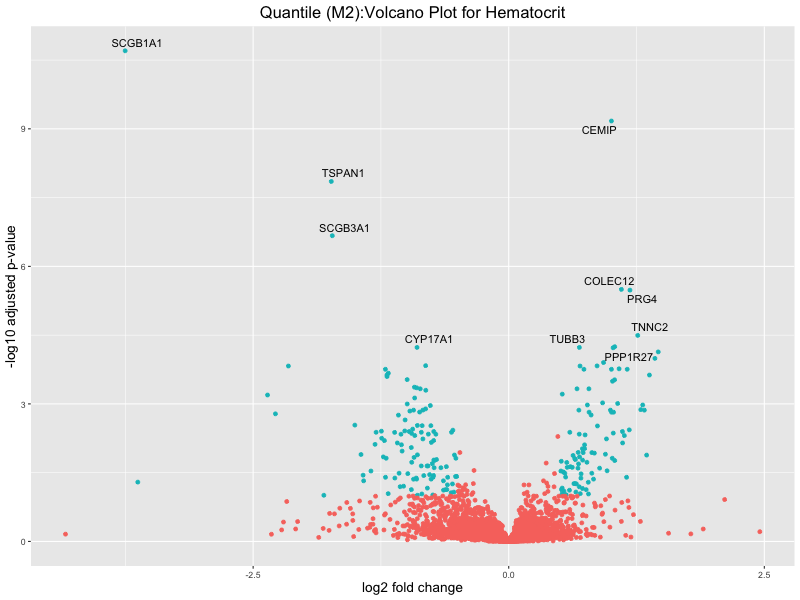
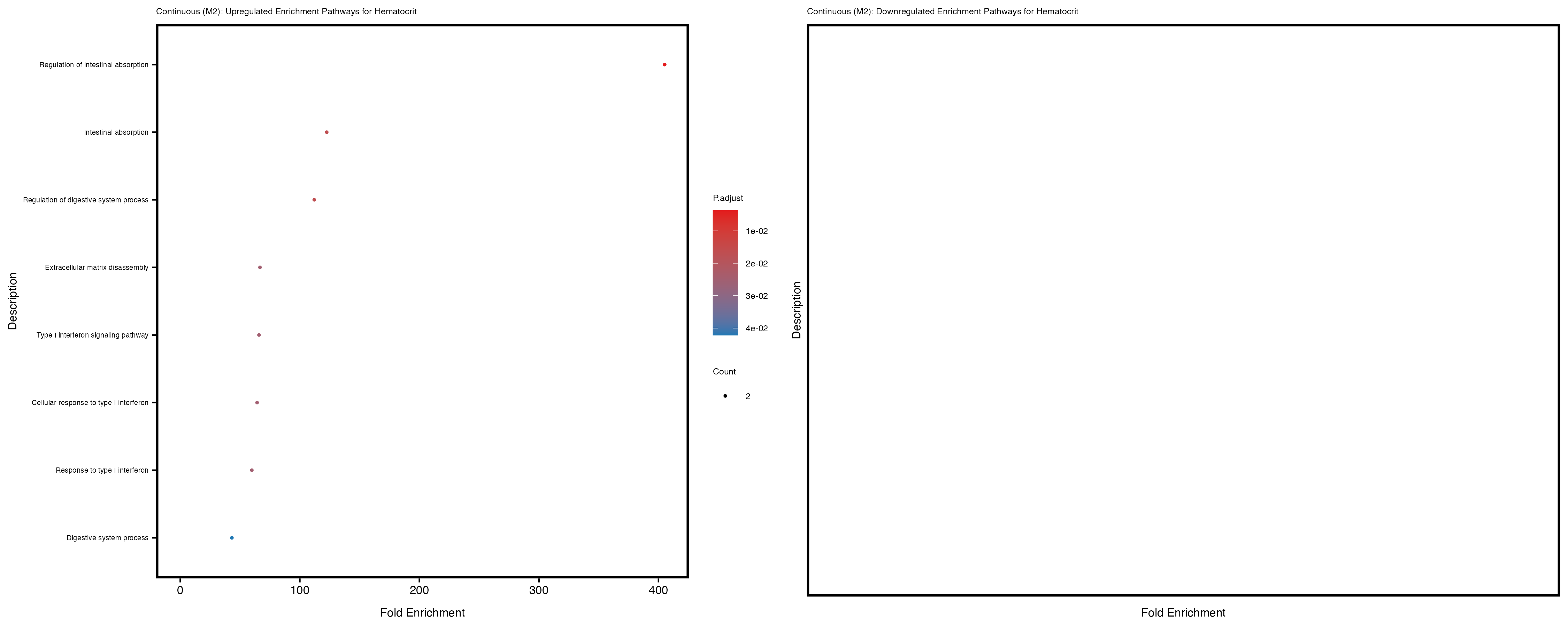
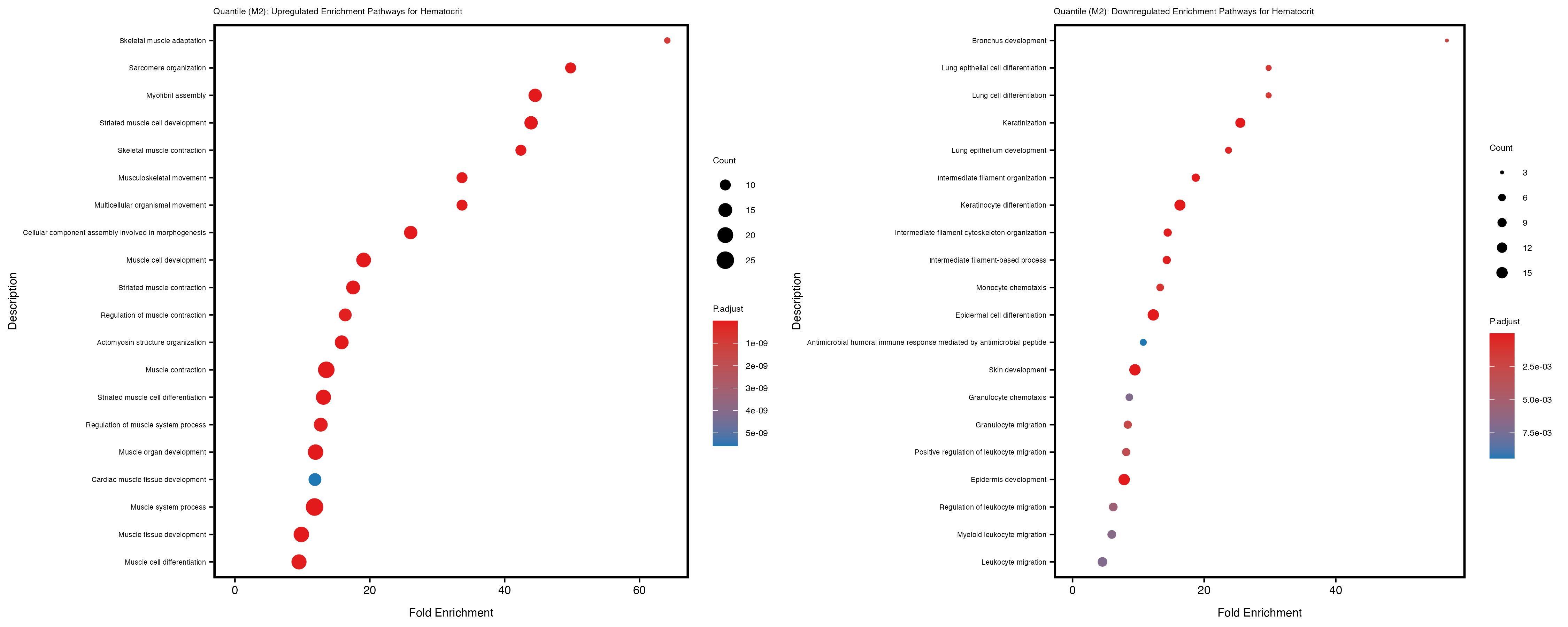
Hemoglobin.concentration
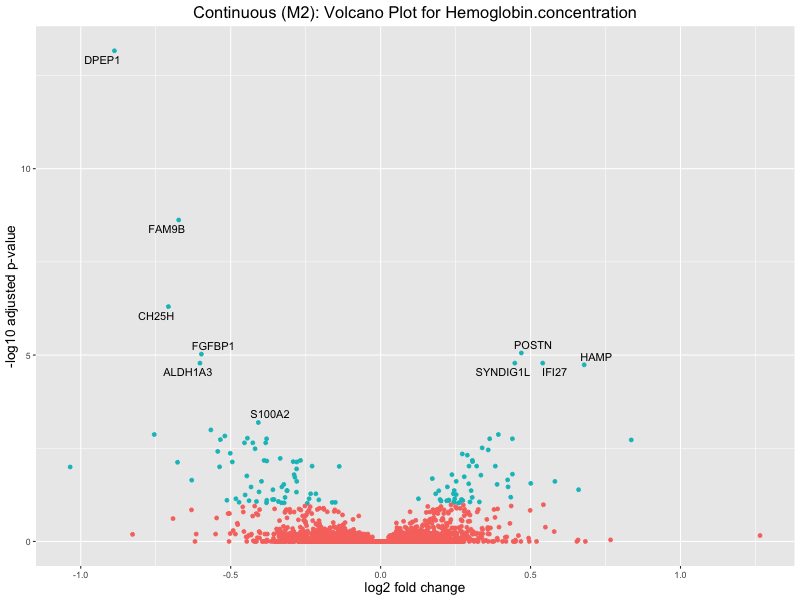
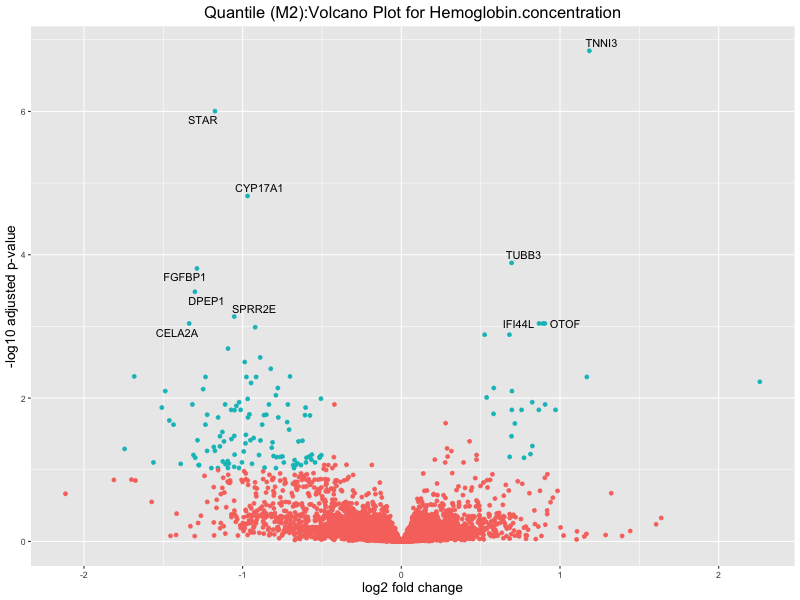


High.light.scatter.reticulocyte.count
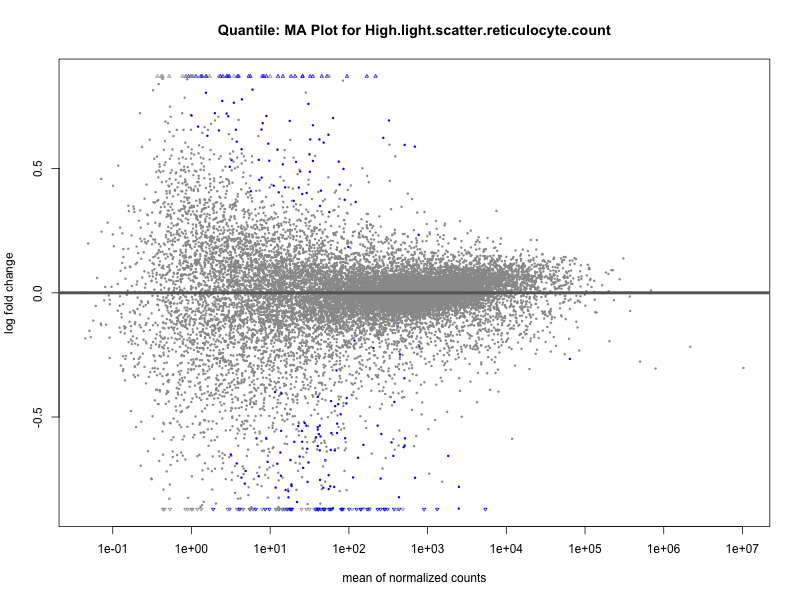
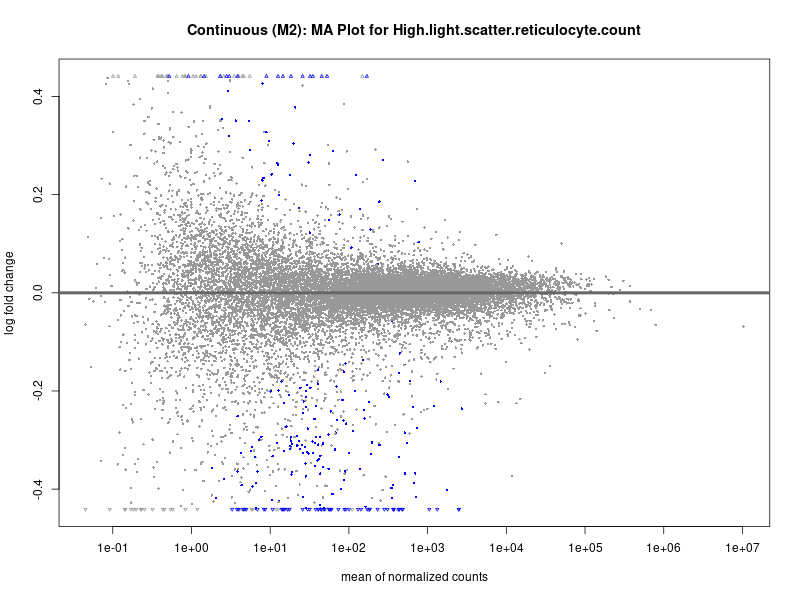
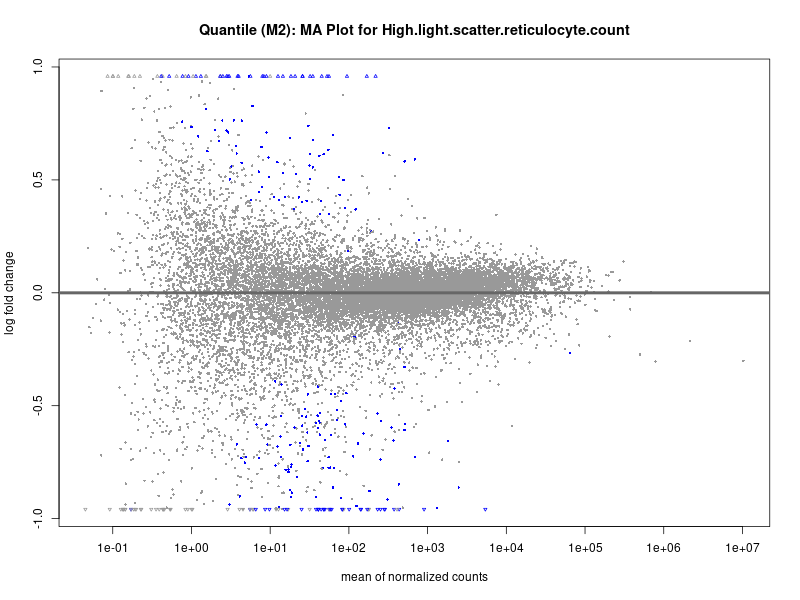
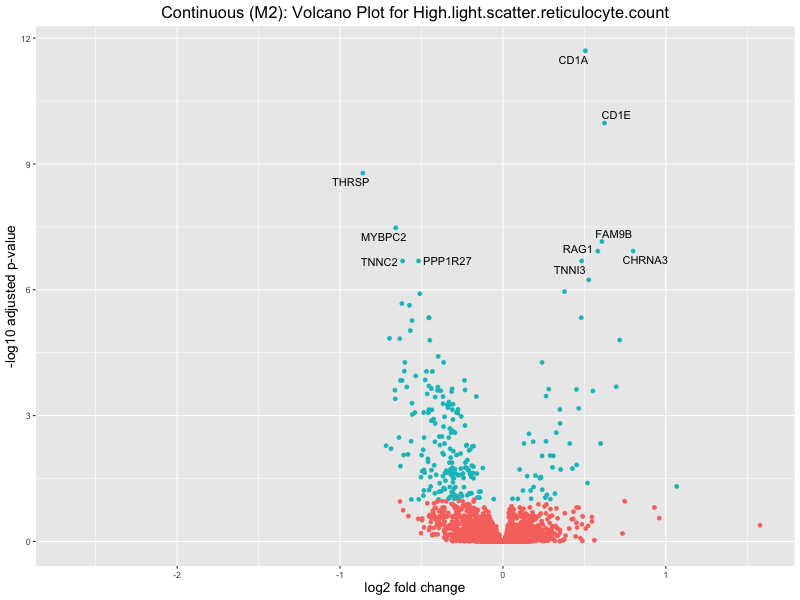
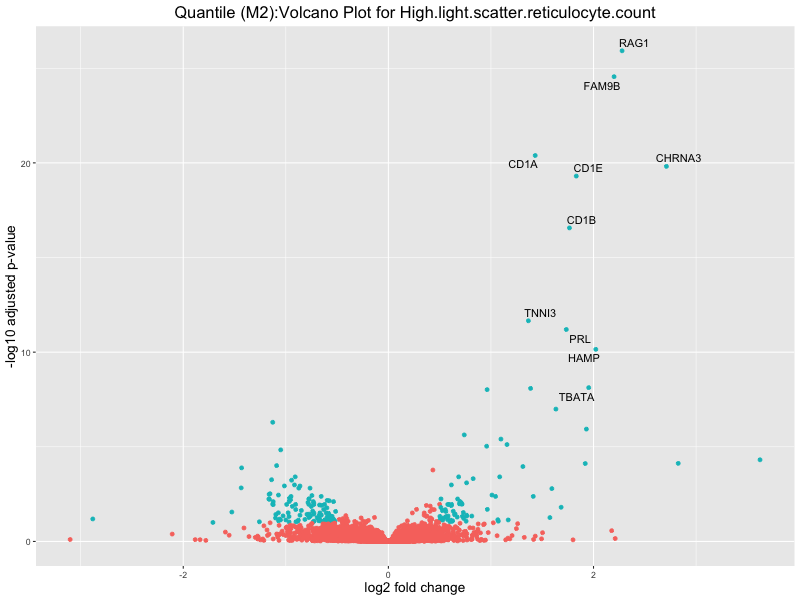
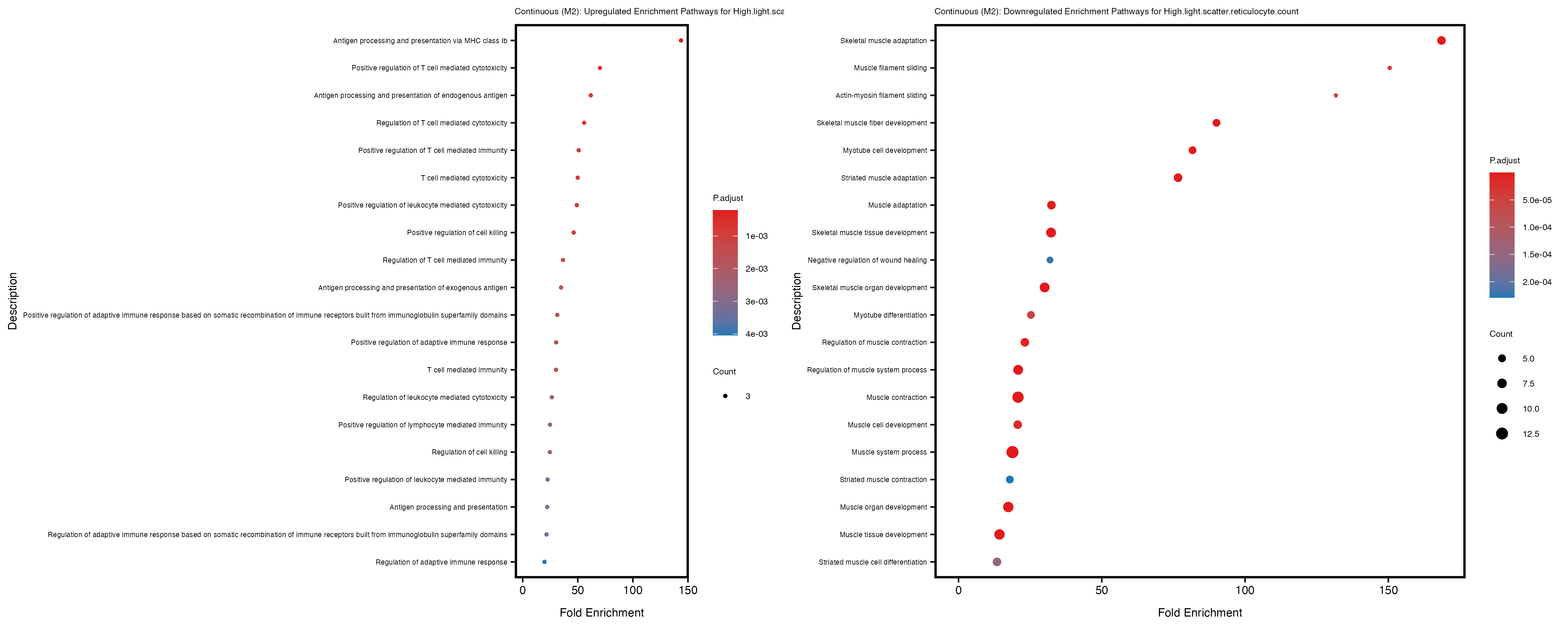
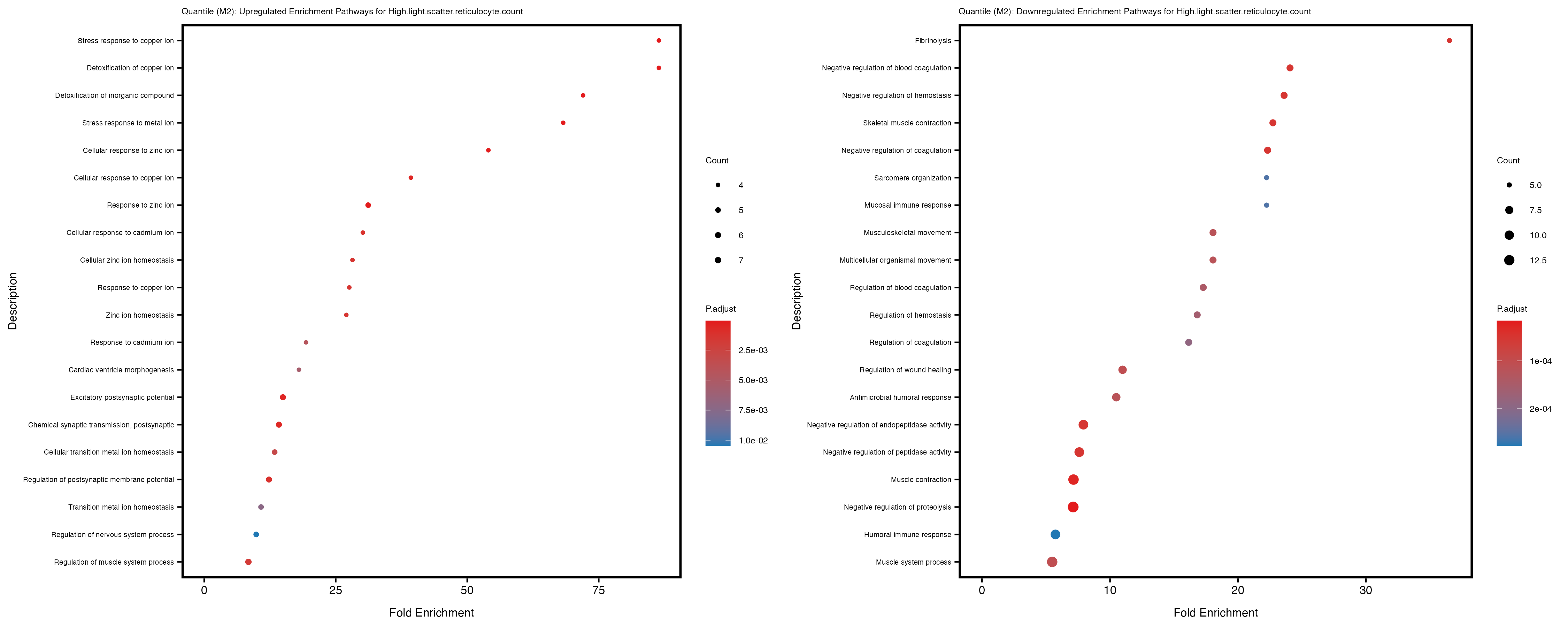
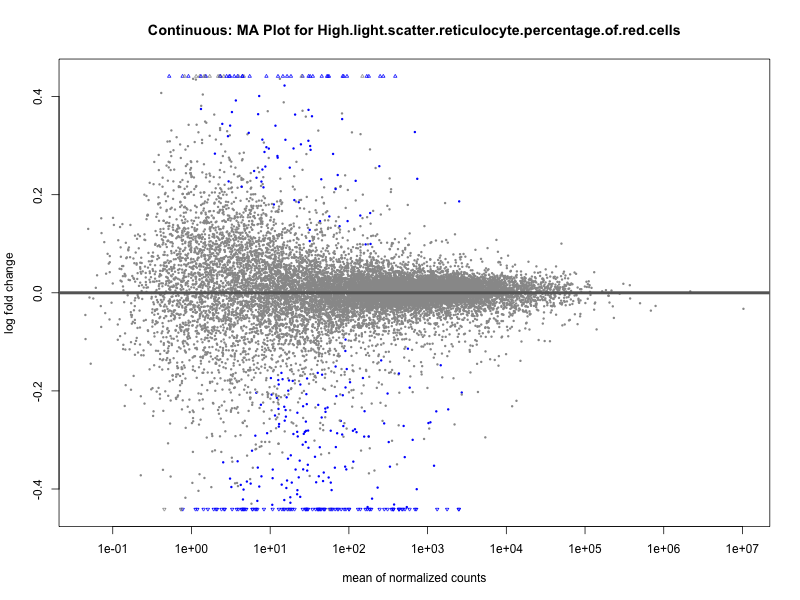
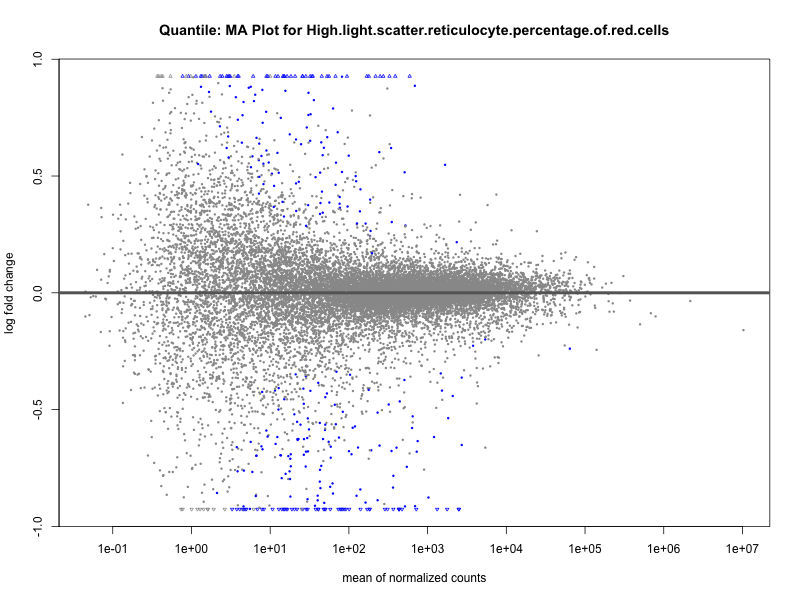
High.light.scatter.reticulocyte.percentage.of.red.cells
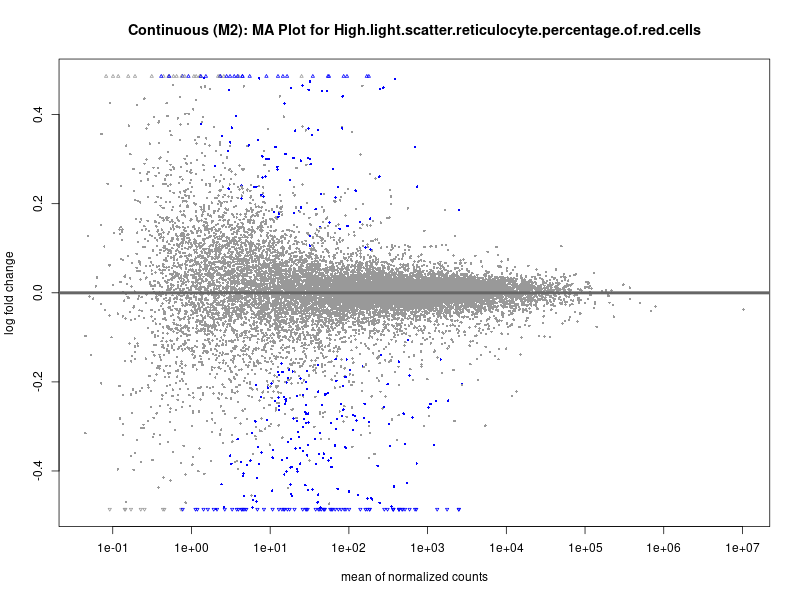
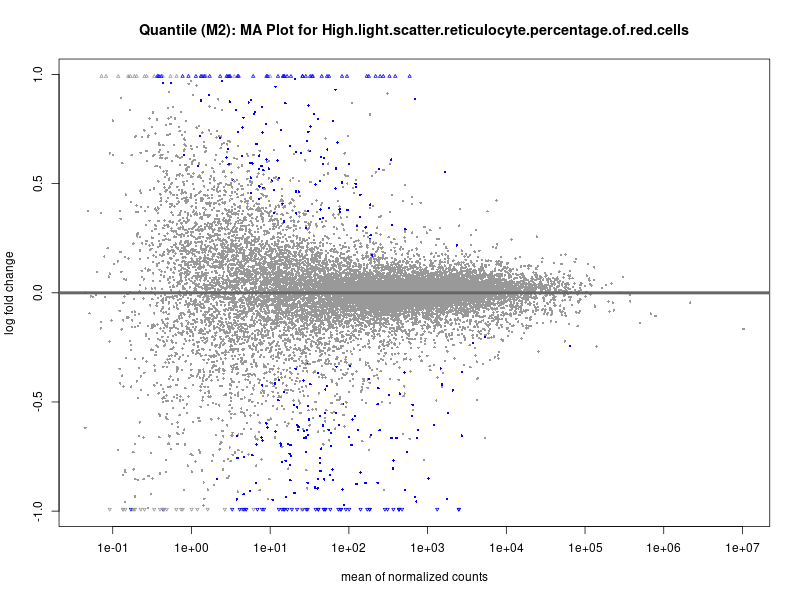
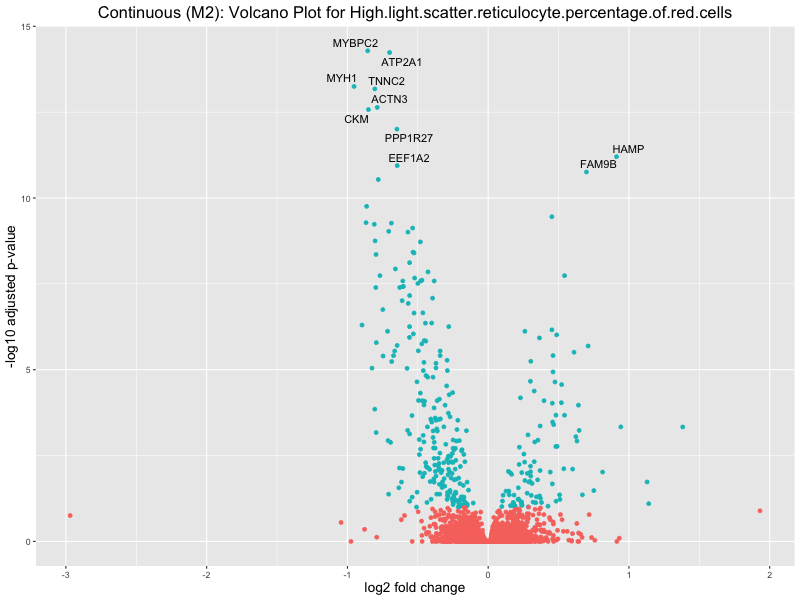
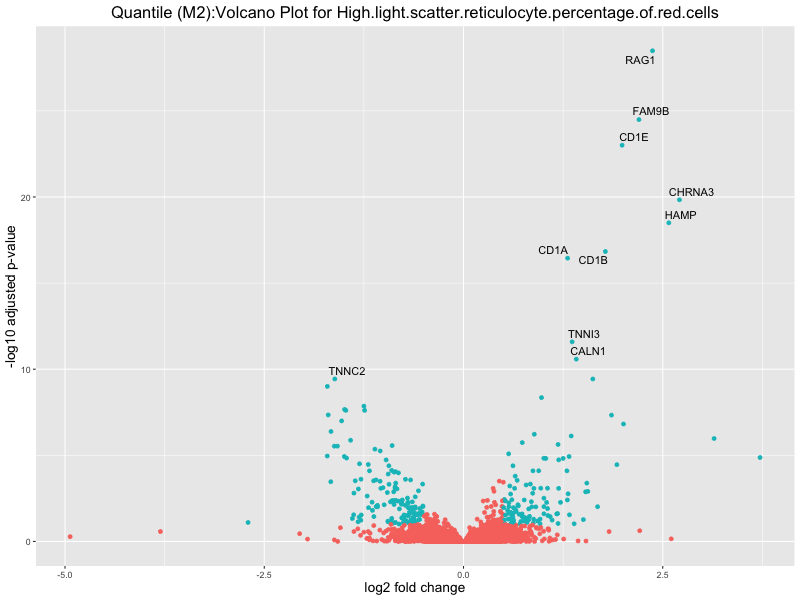
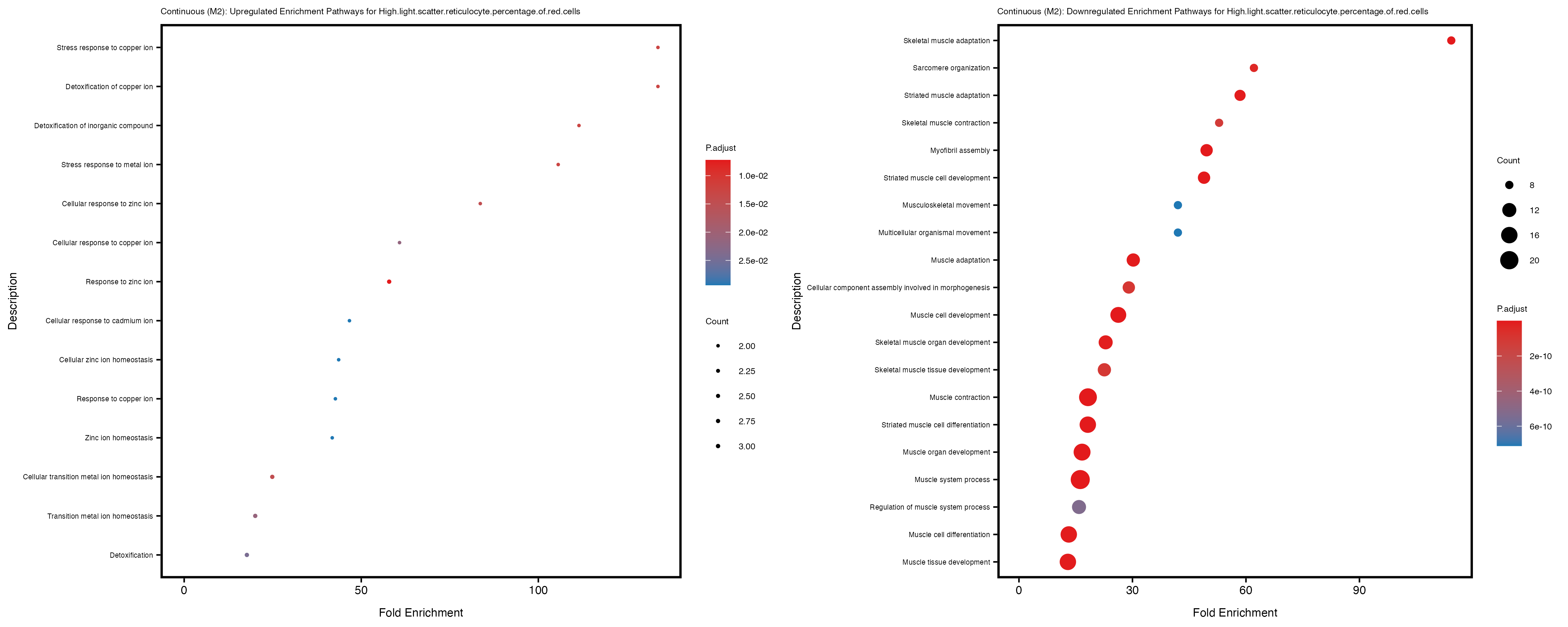
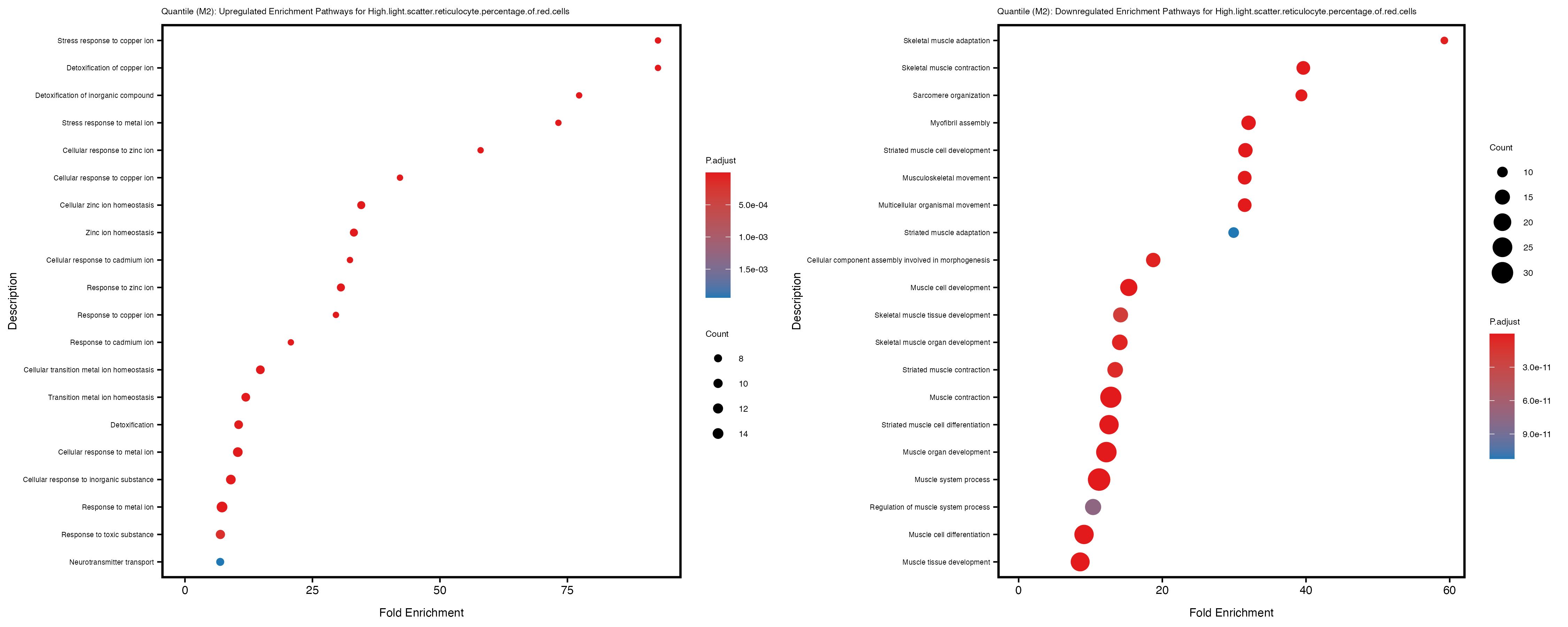
Immature.fraction.of.reticulocytes
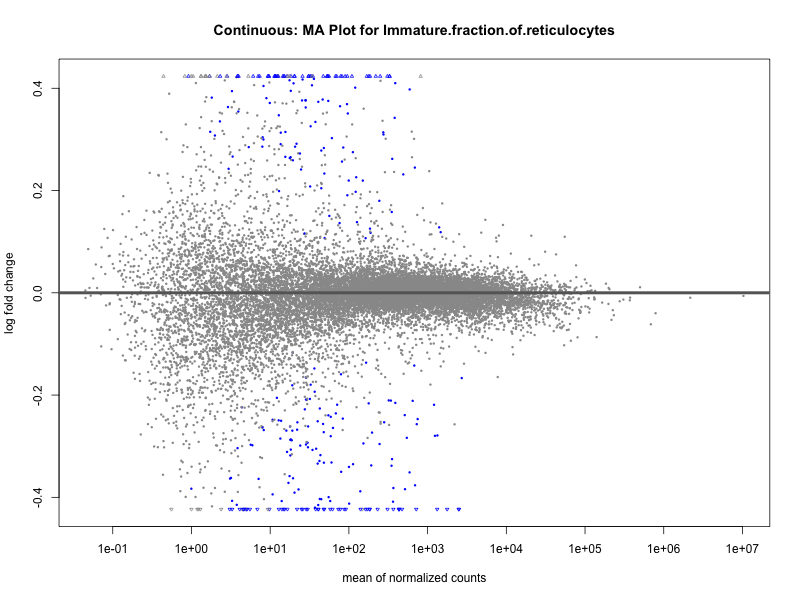
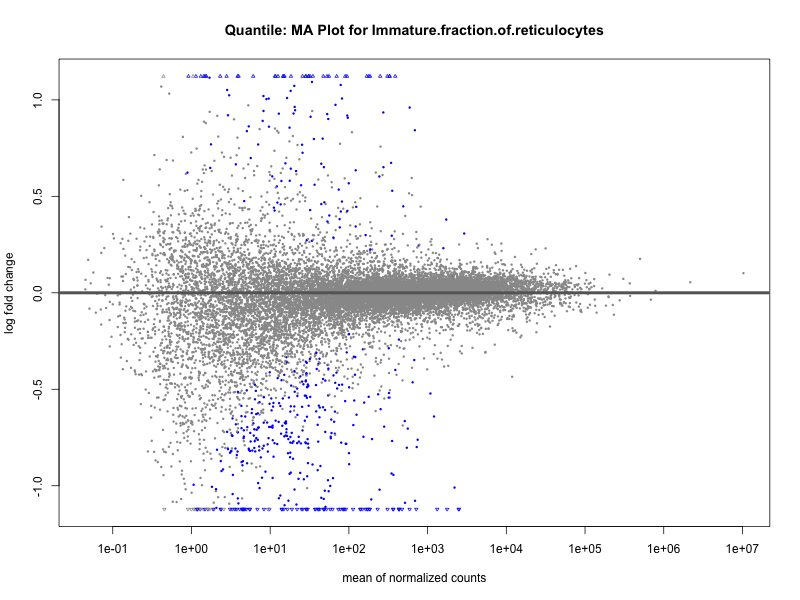
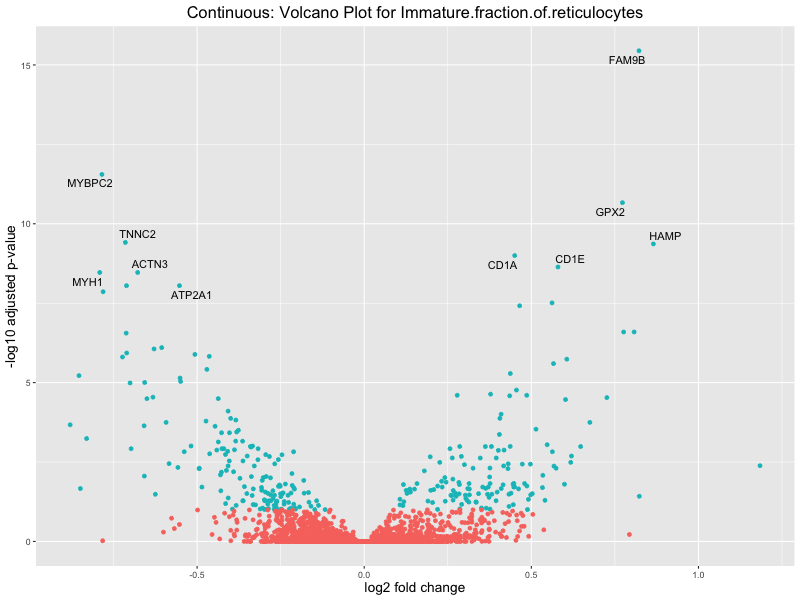
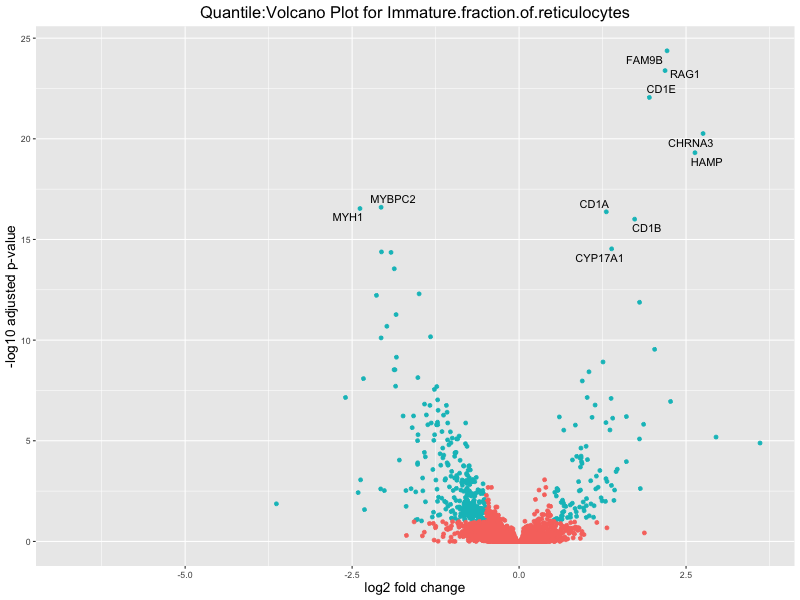
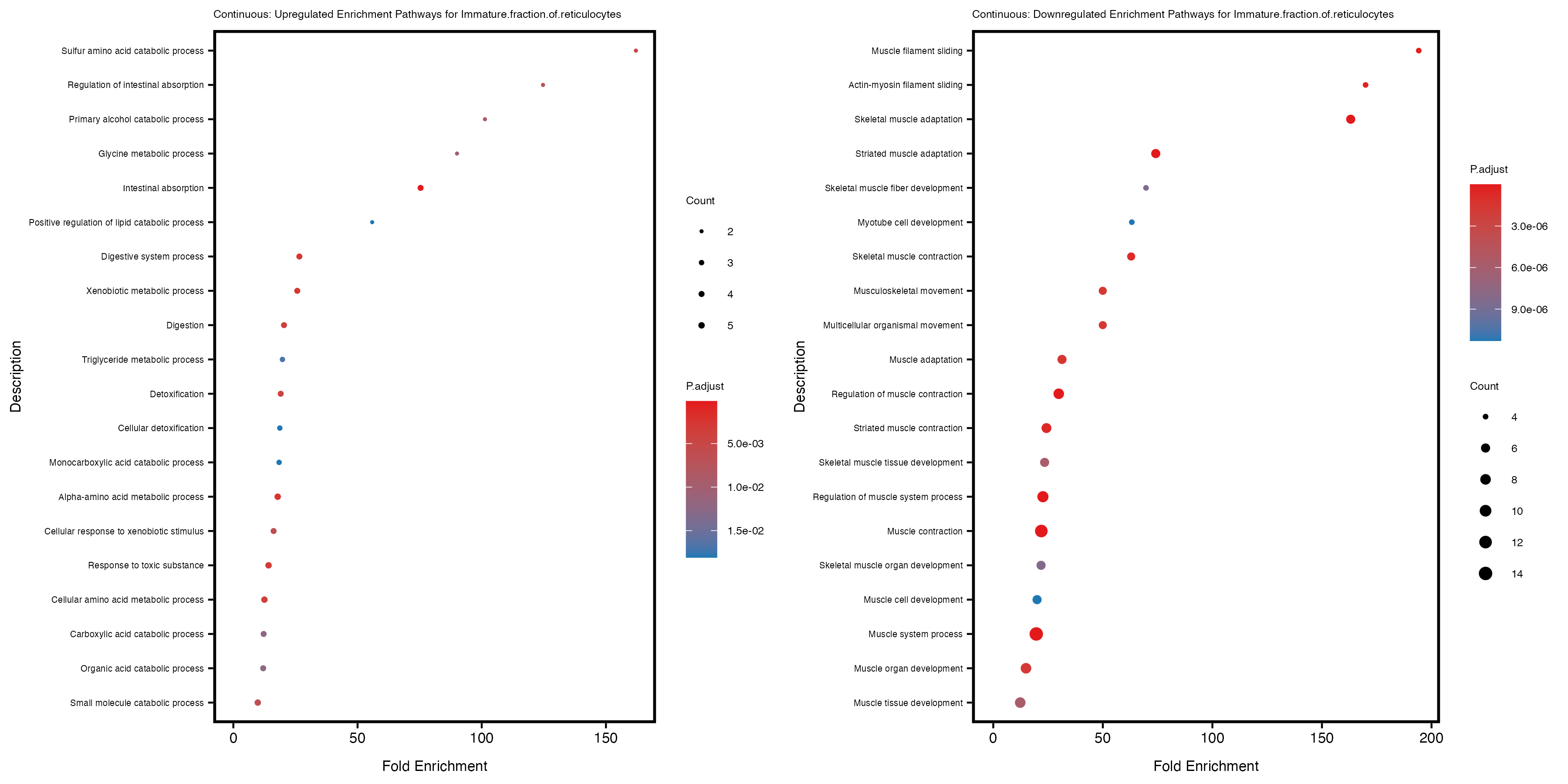
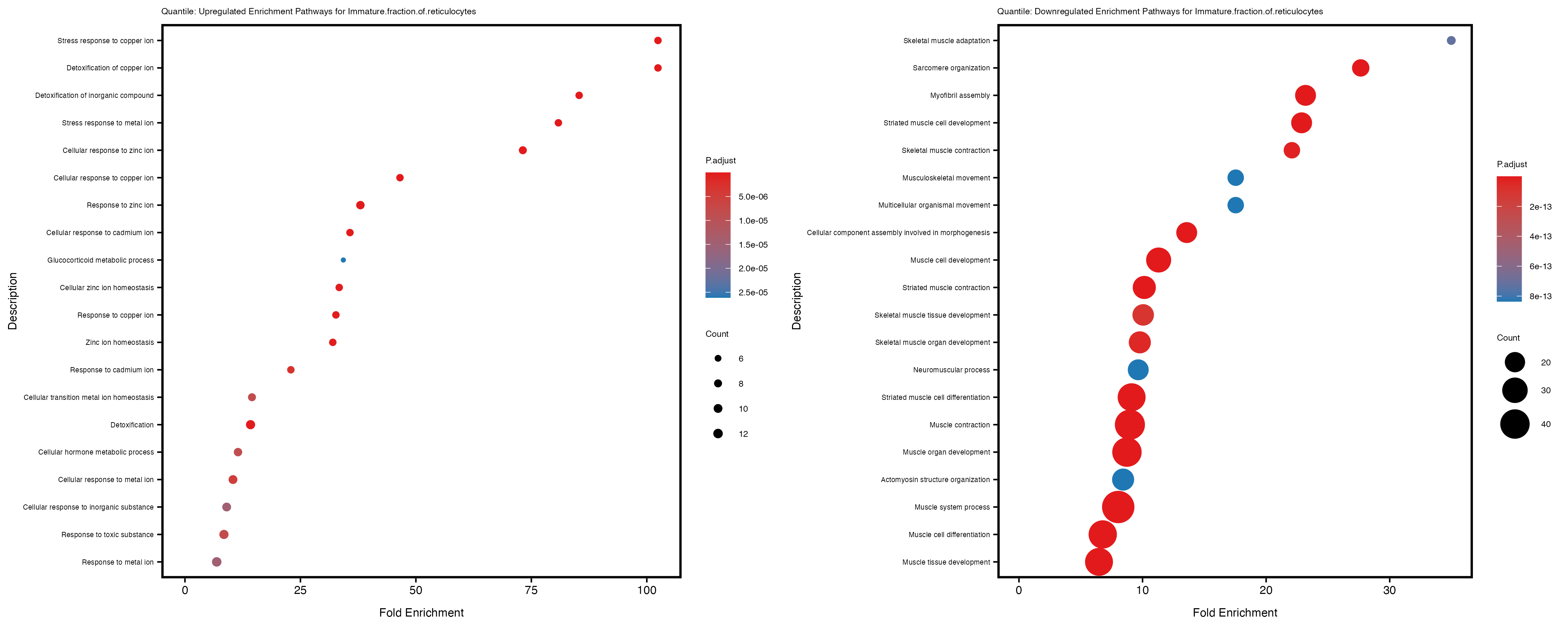
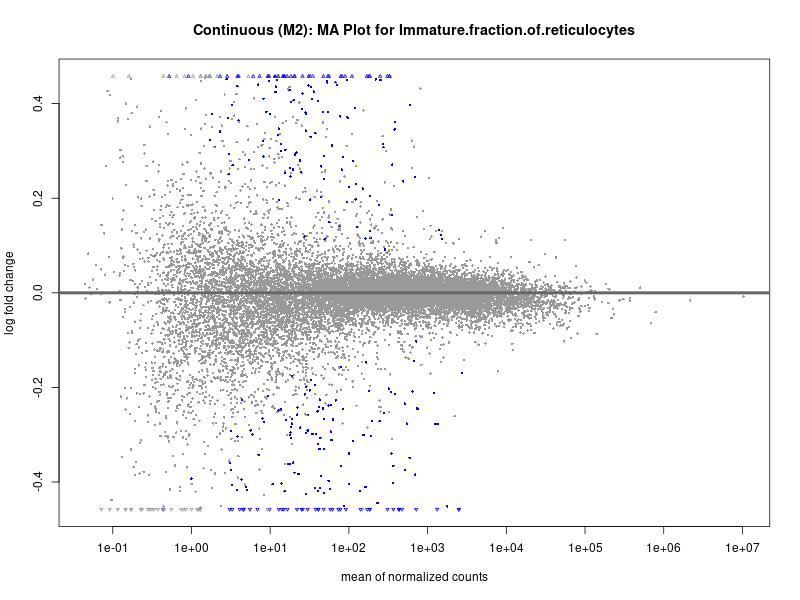
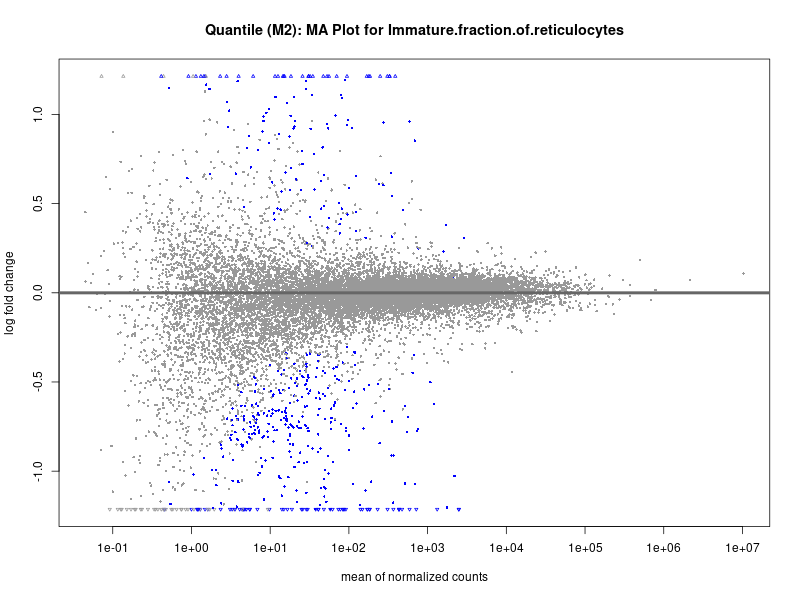
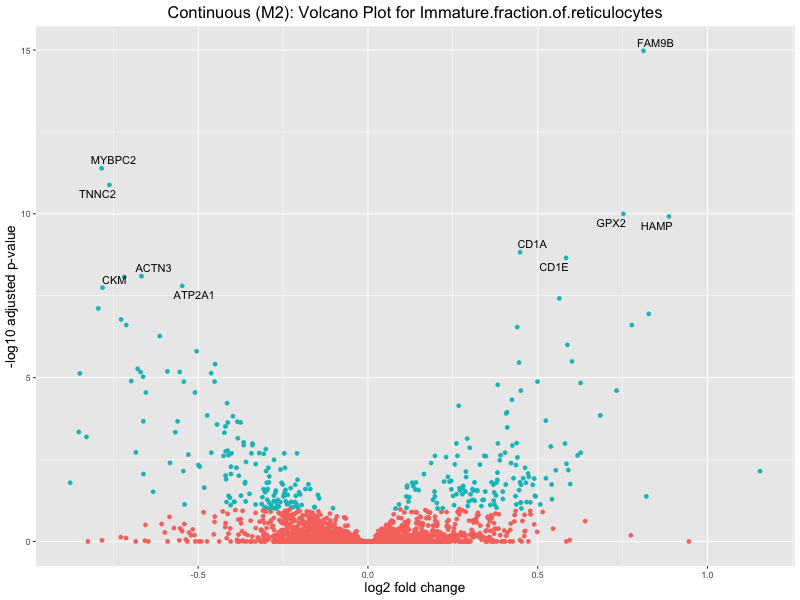
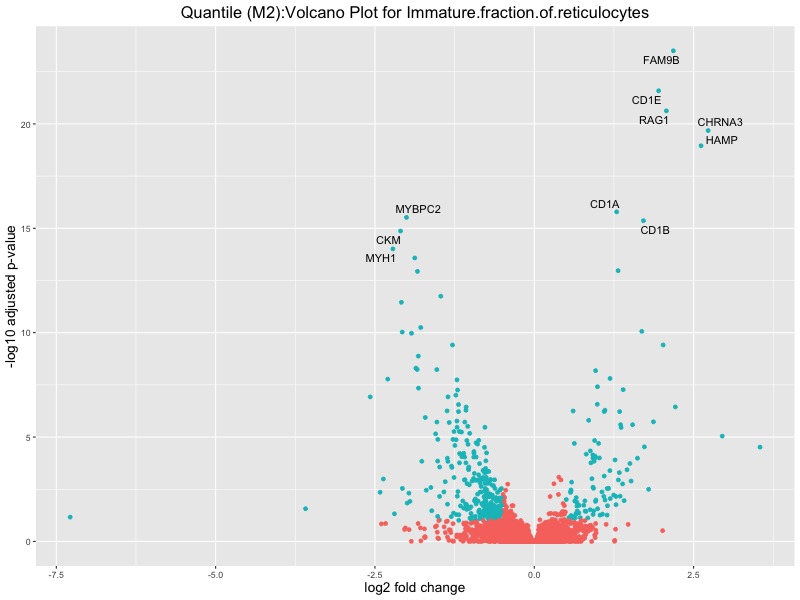
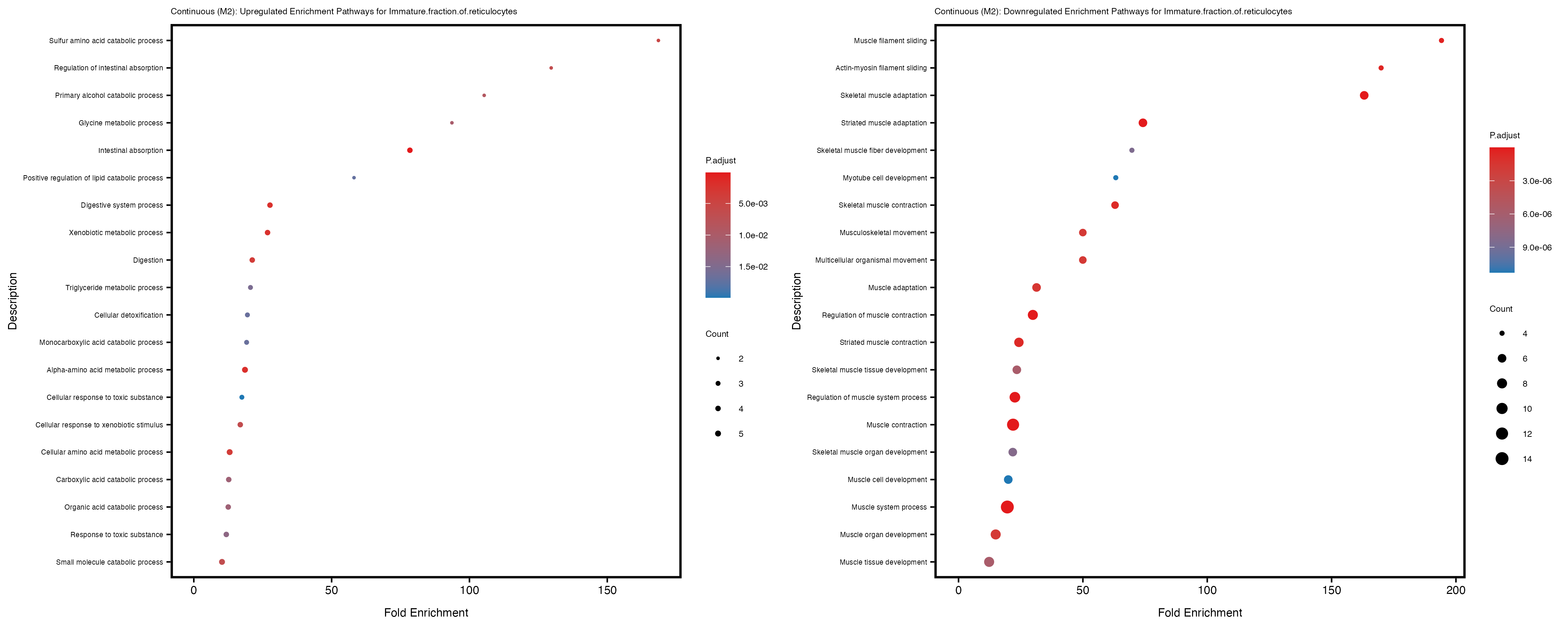
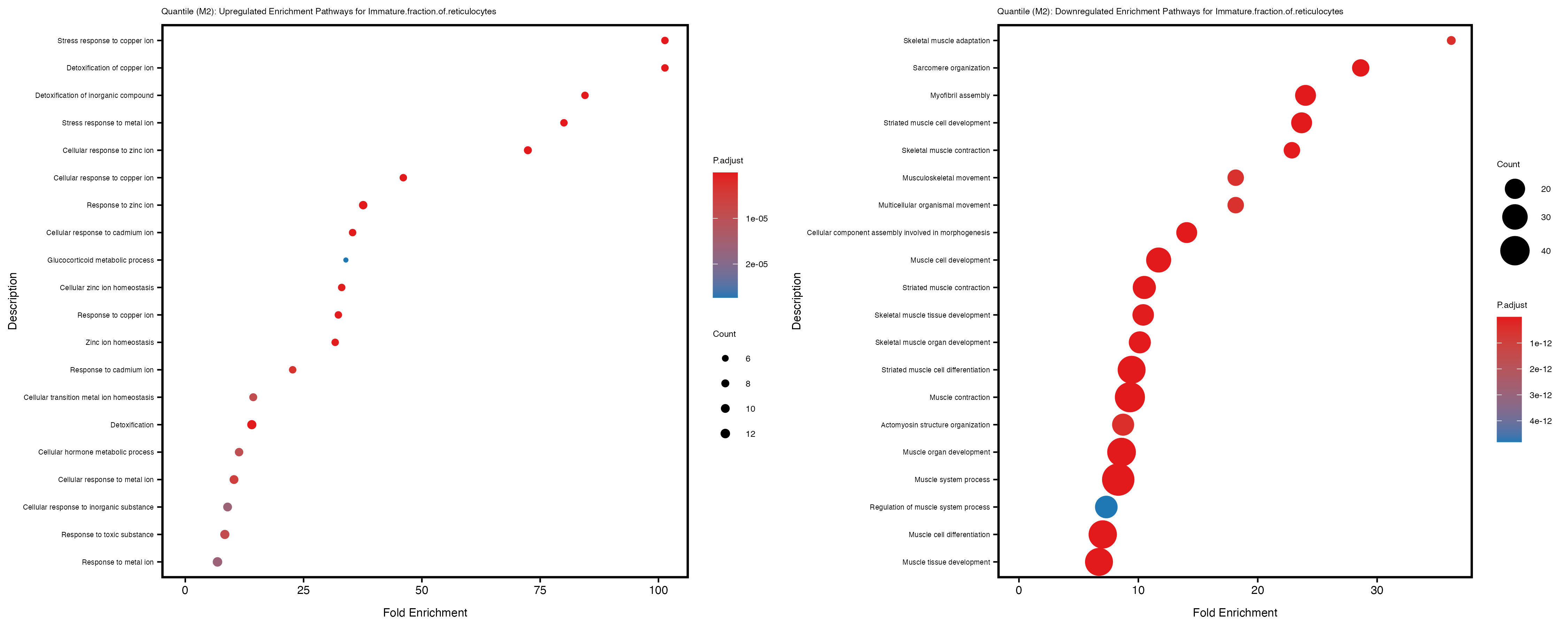
Lymphocyte.count
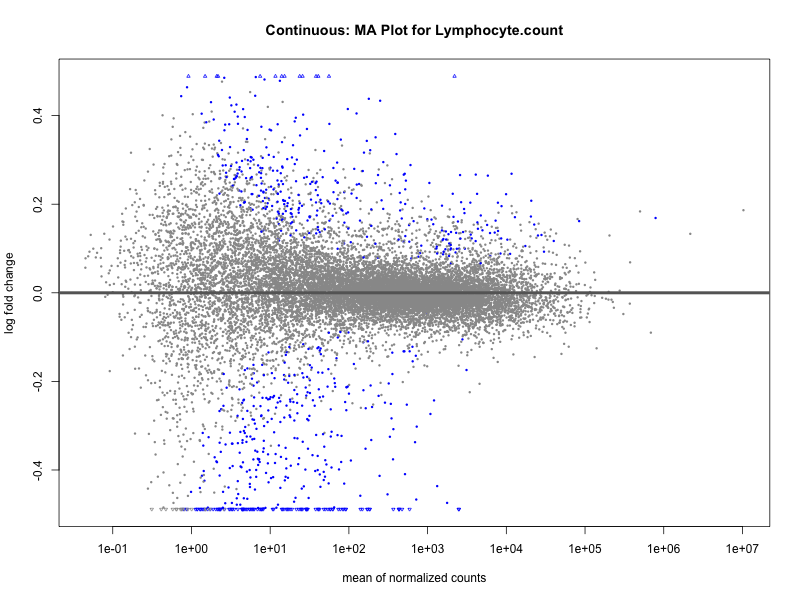
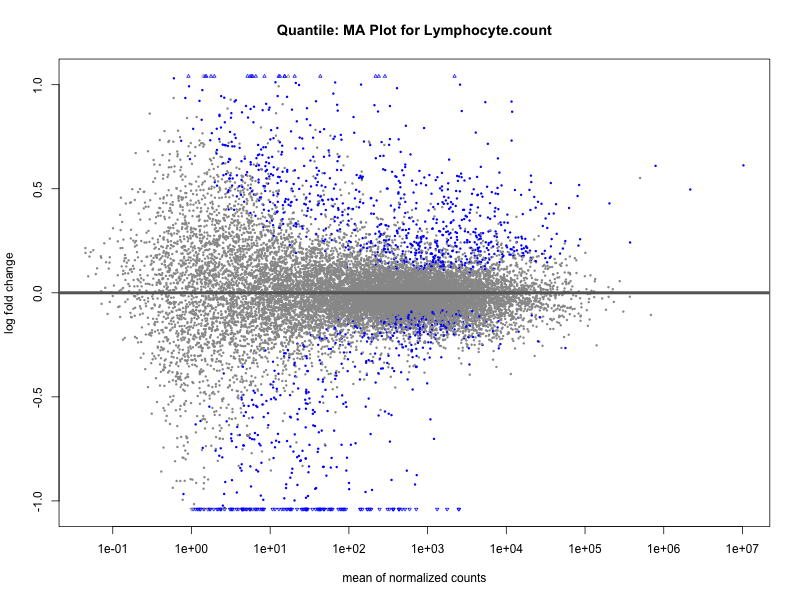
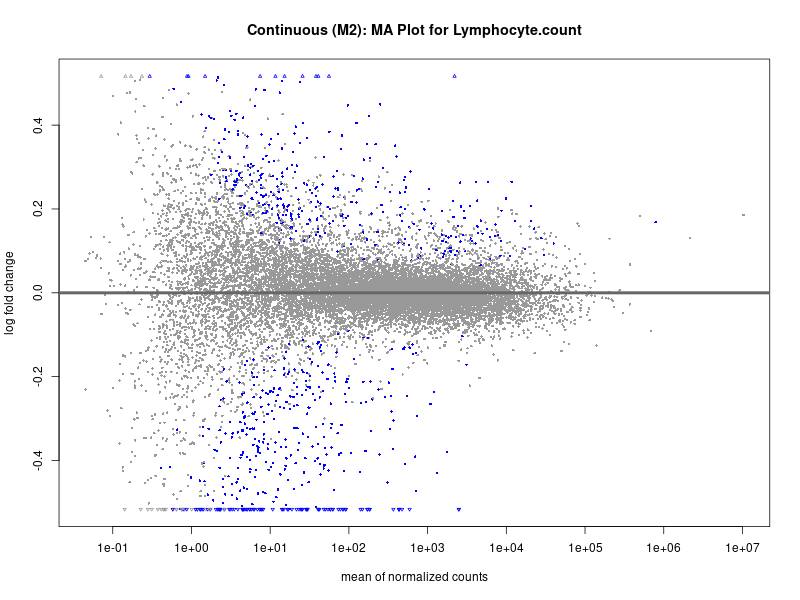
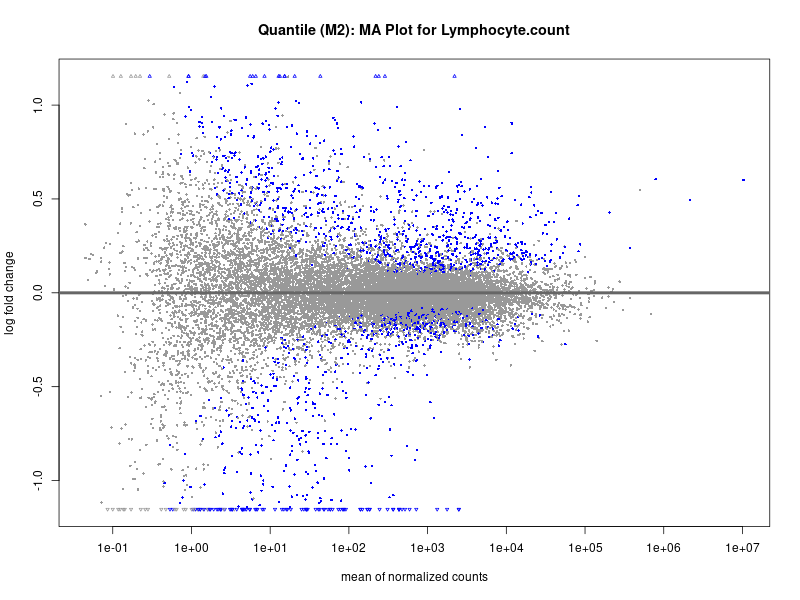
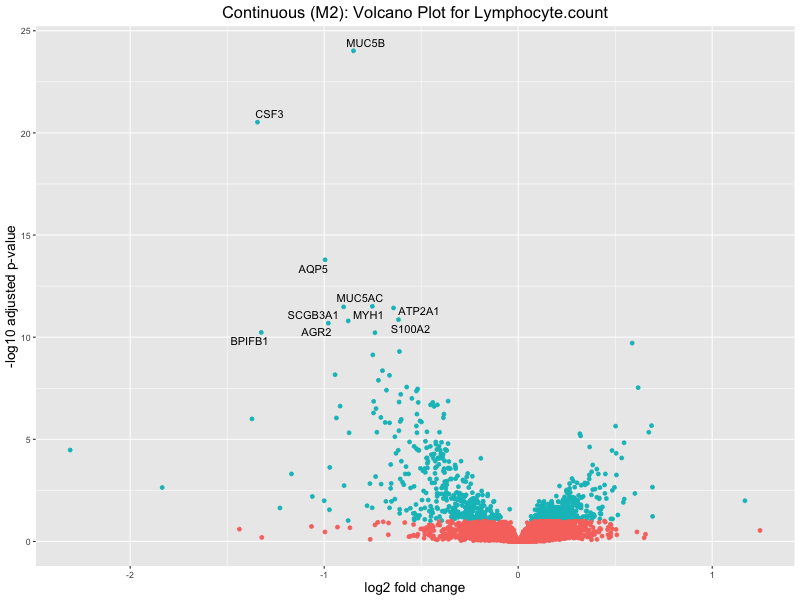
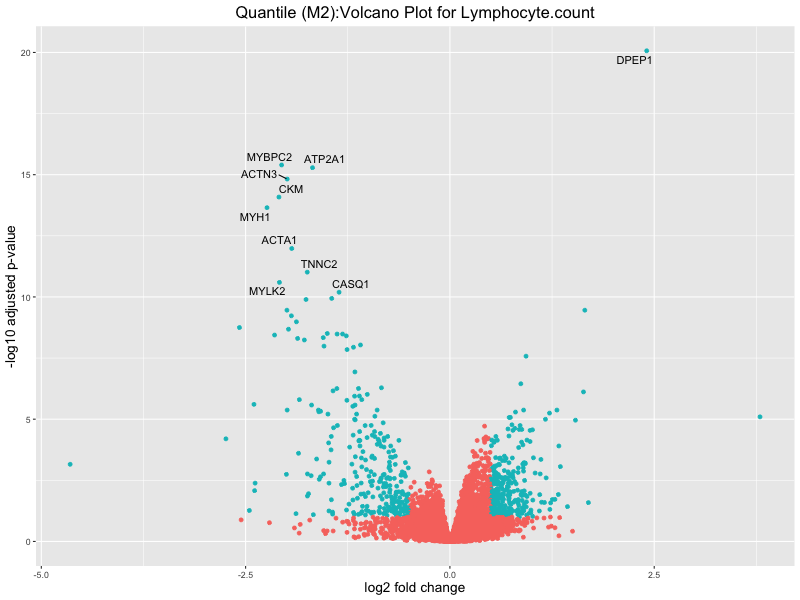
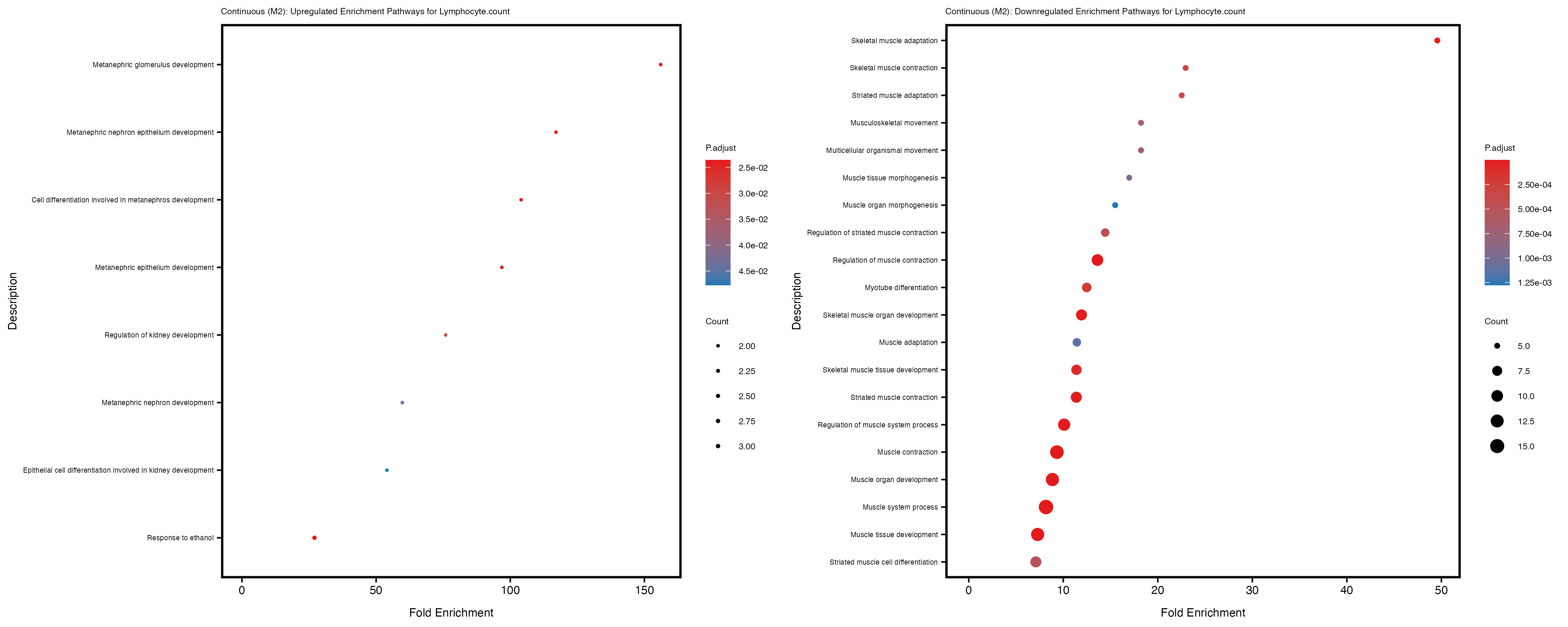
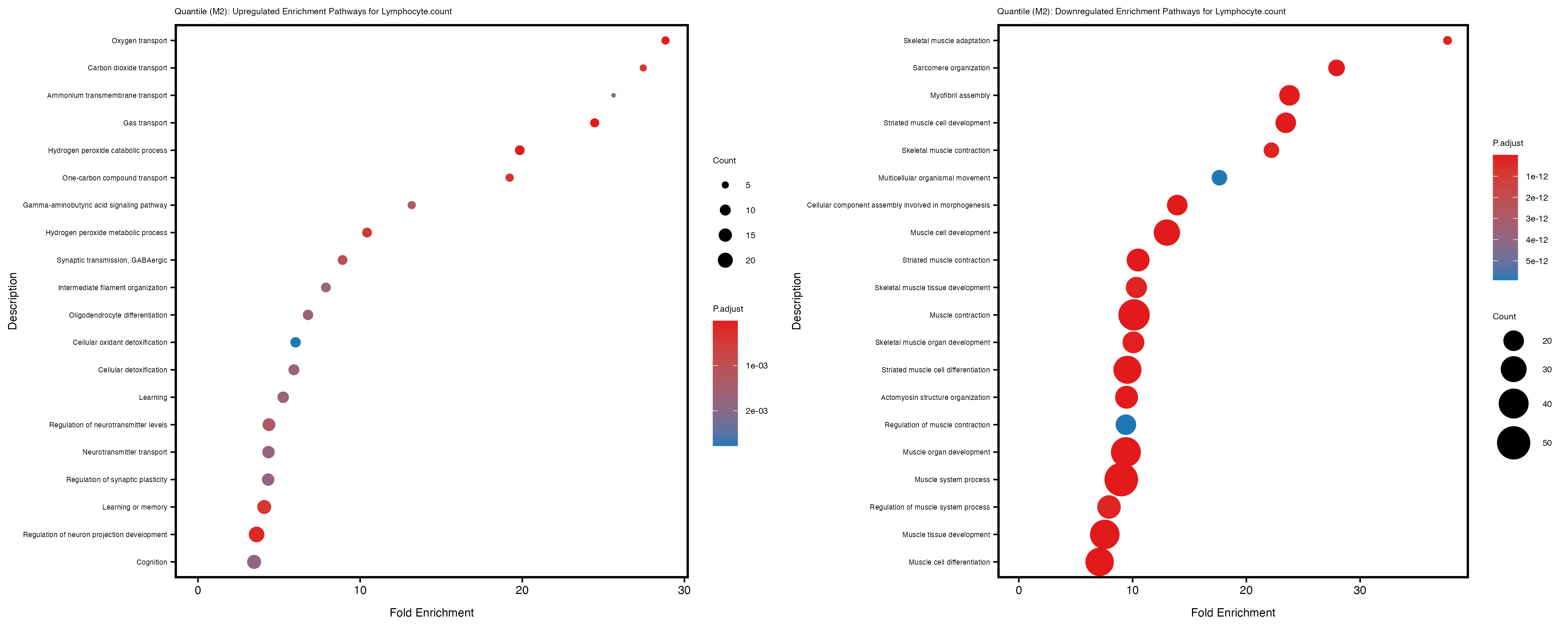
Lymphocyte.percentage.of.white.cells
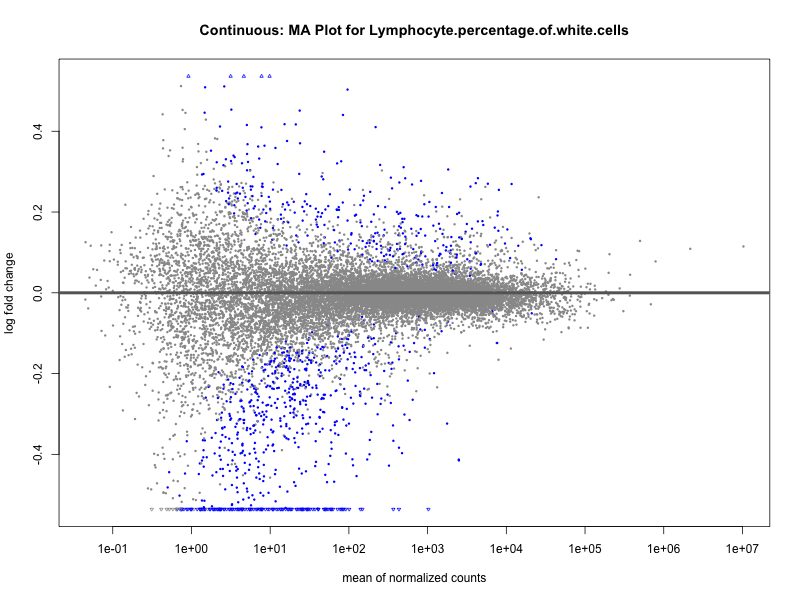
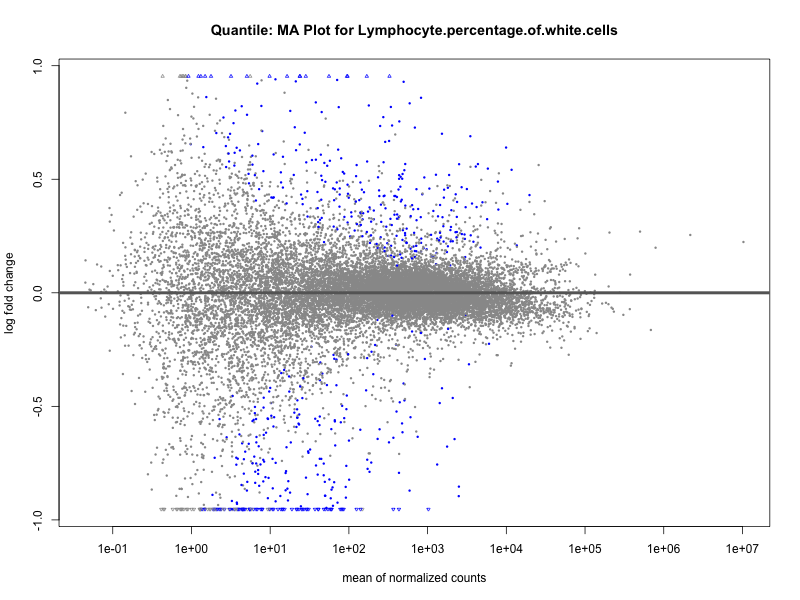
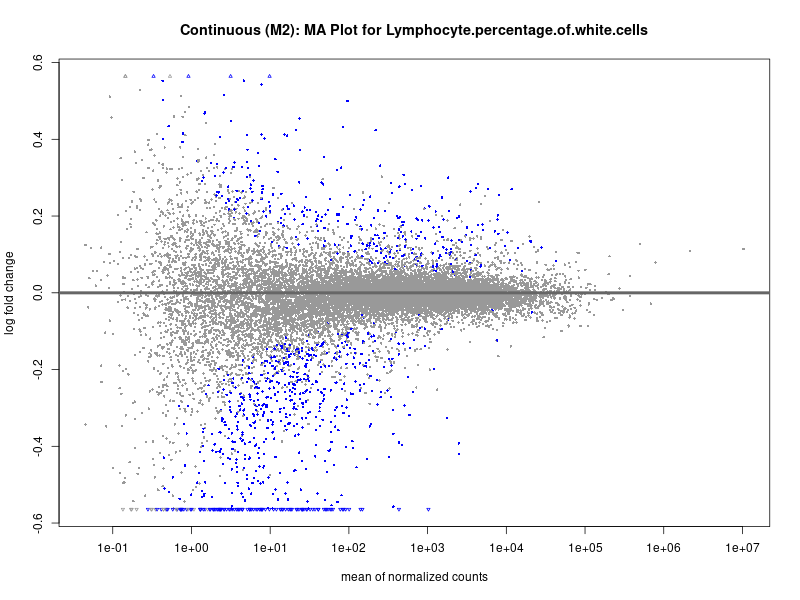
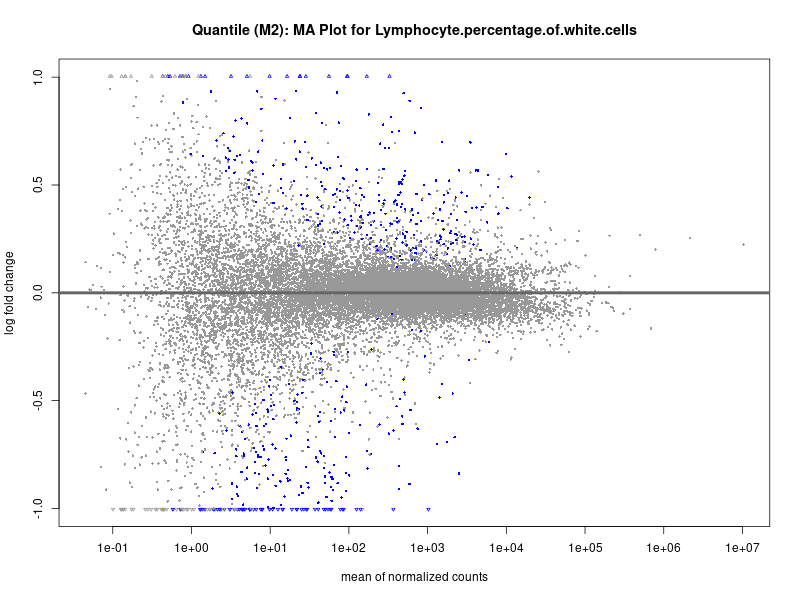
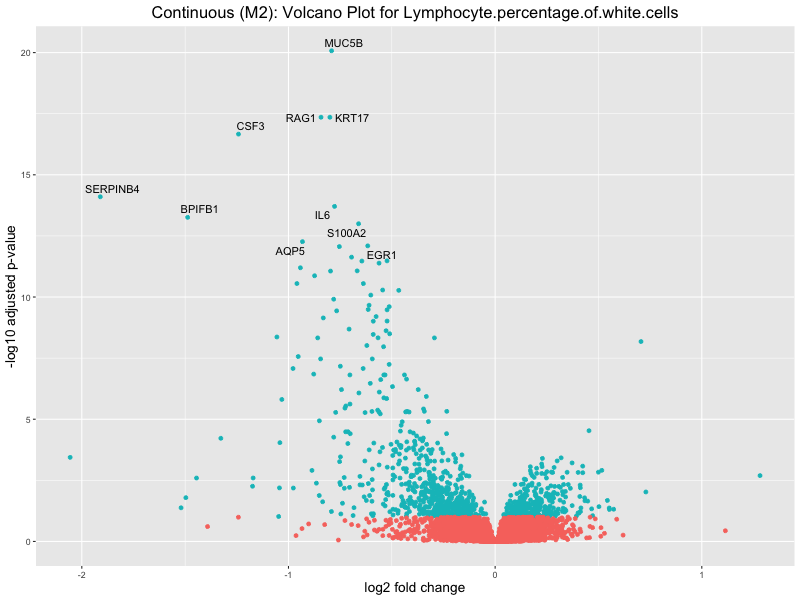
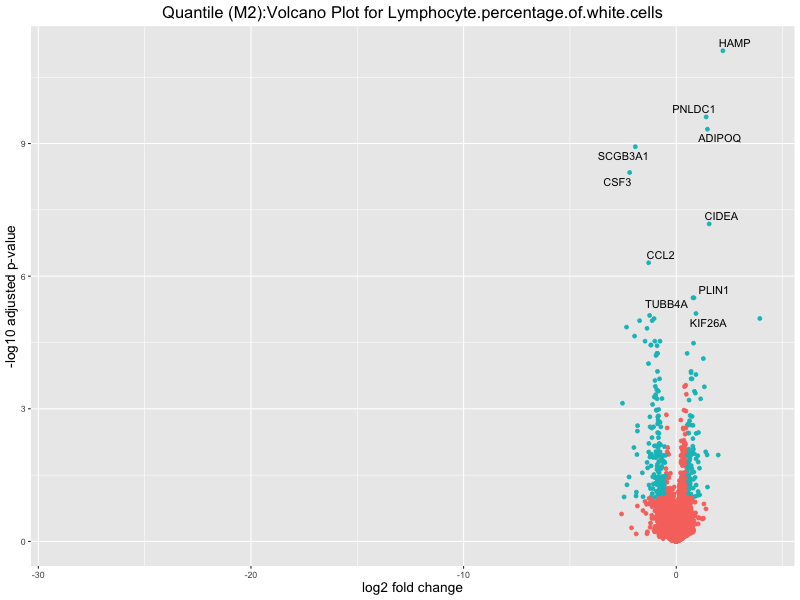
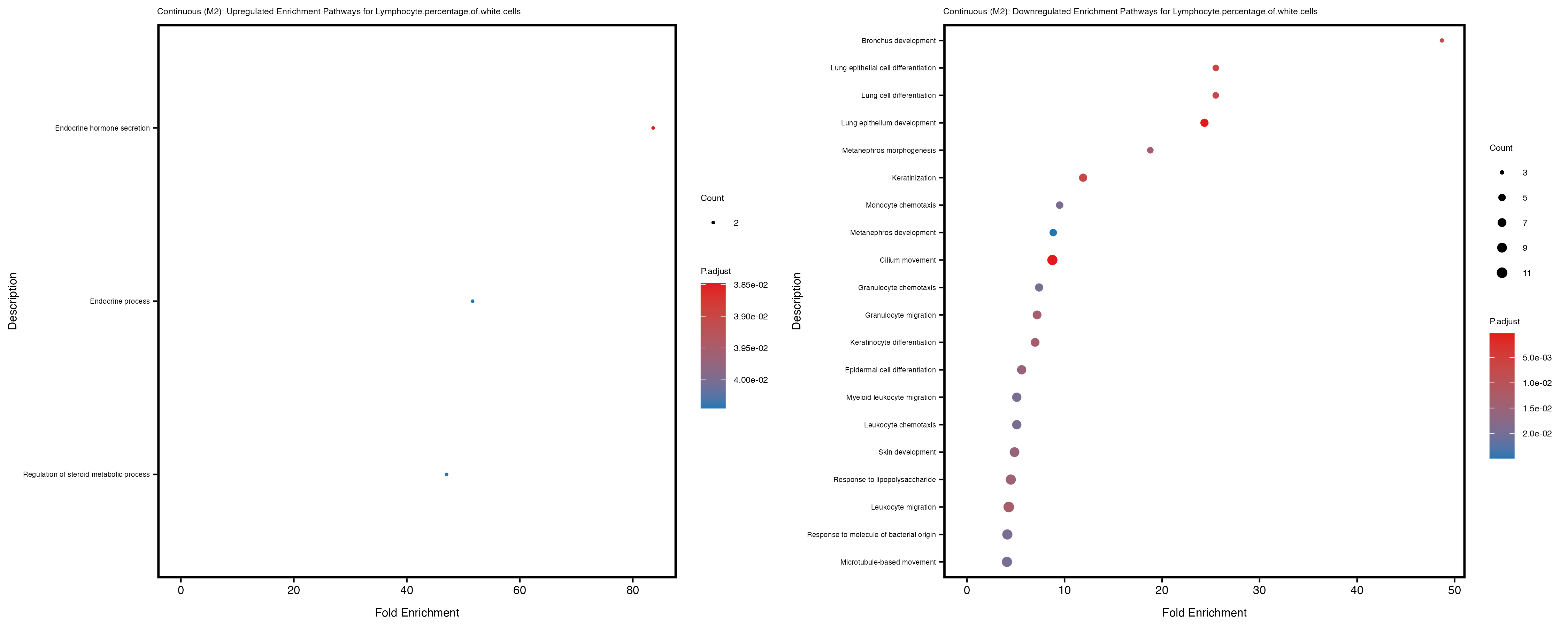
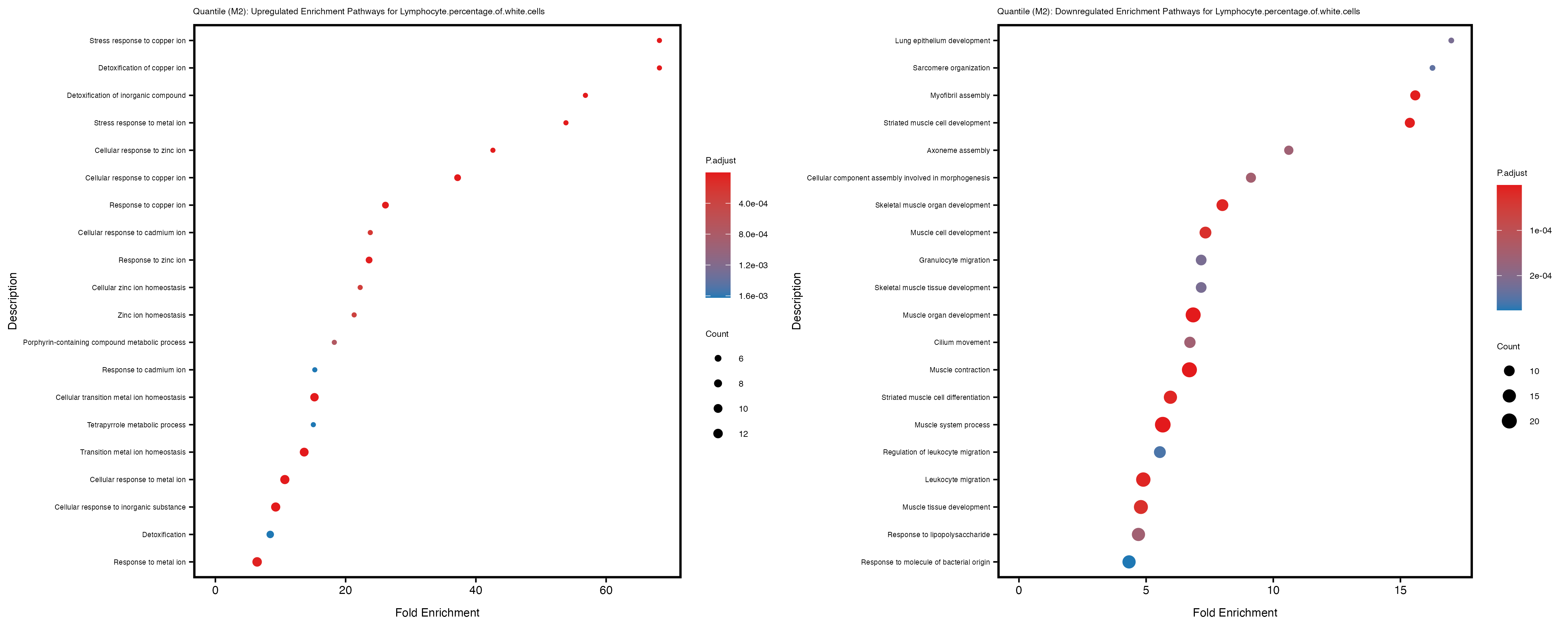
Mean.corpuscular.hemoglobin.concentration
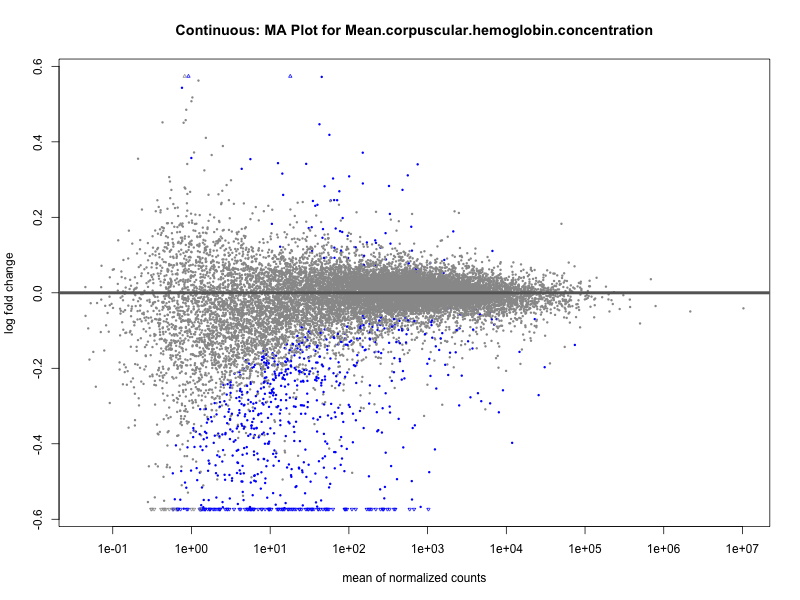
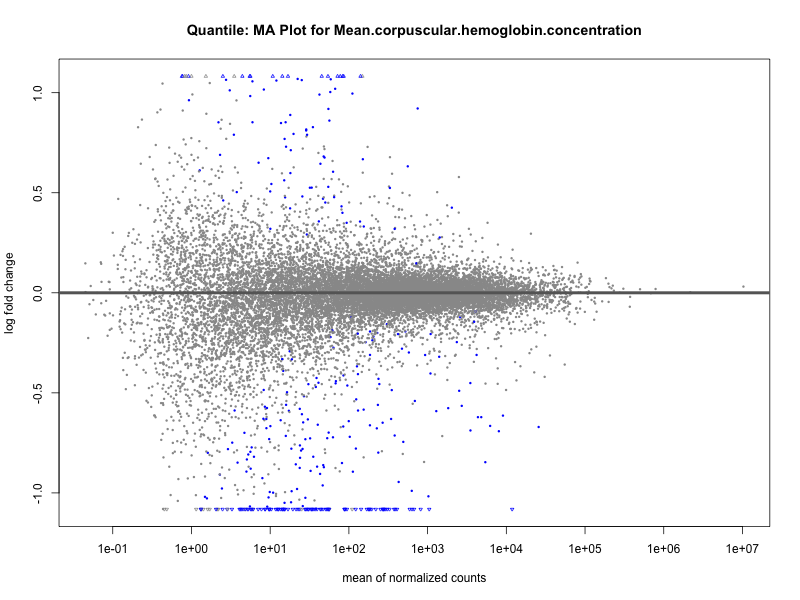
Mean.corpuscular.hemoglobin
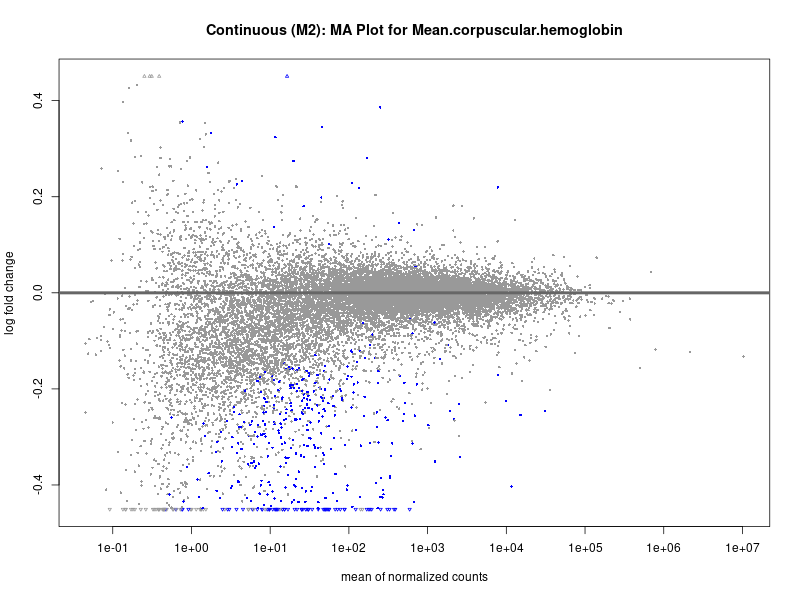
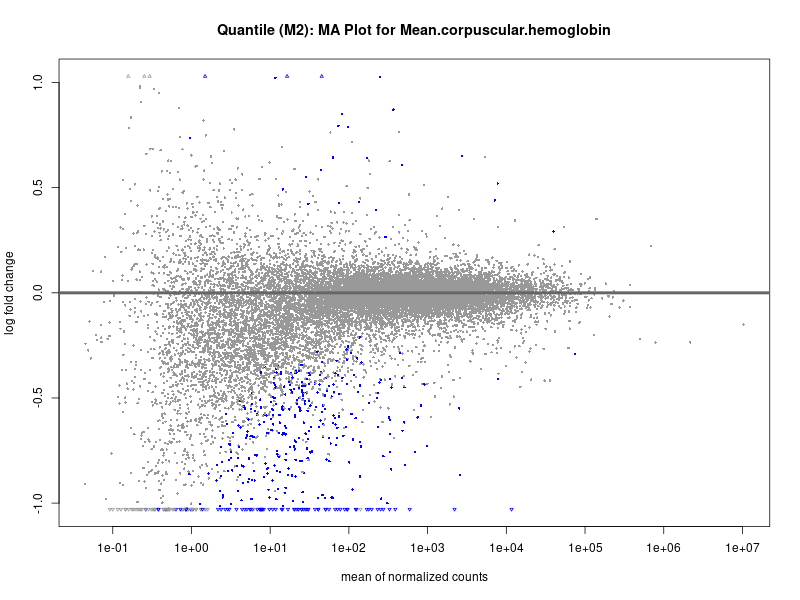
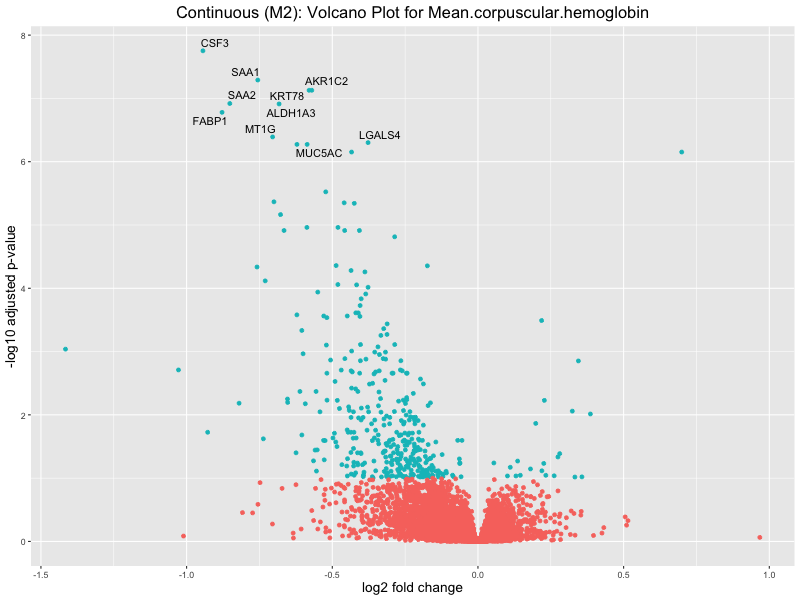
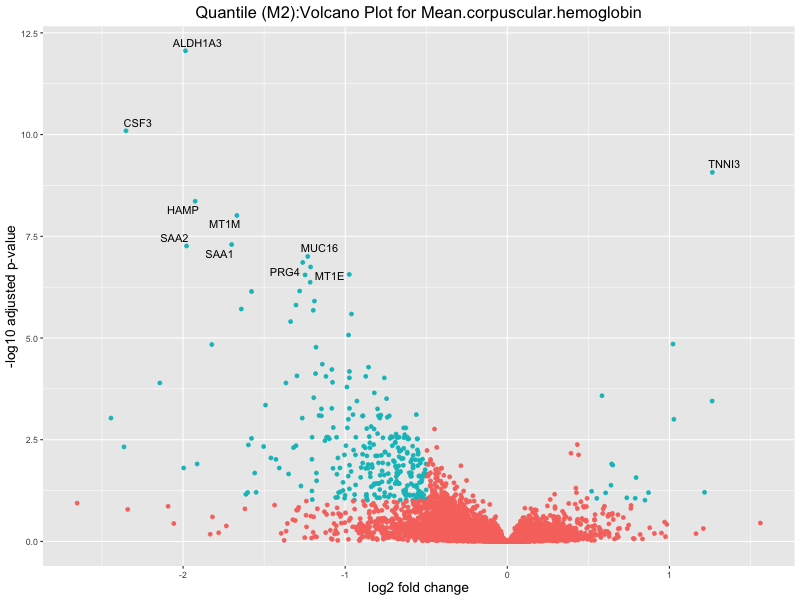
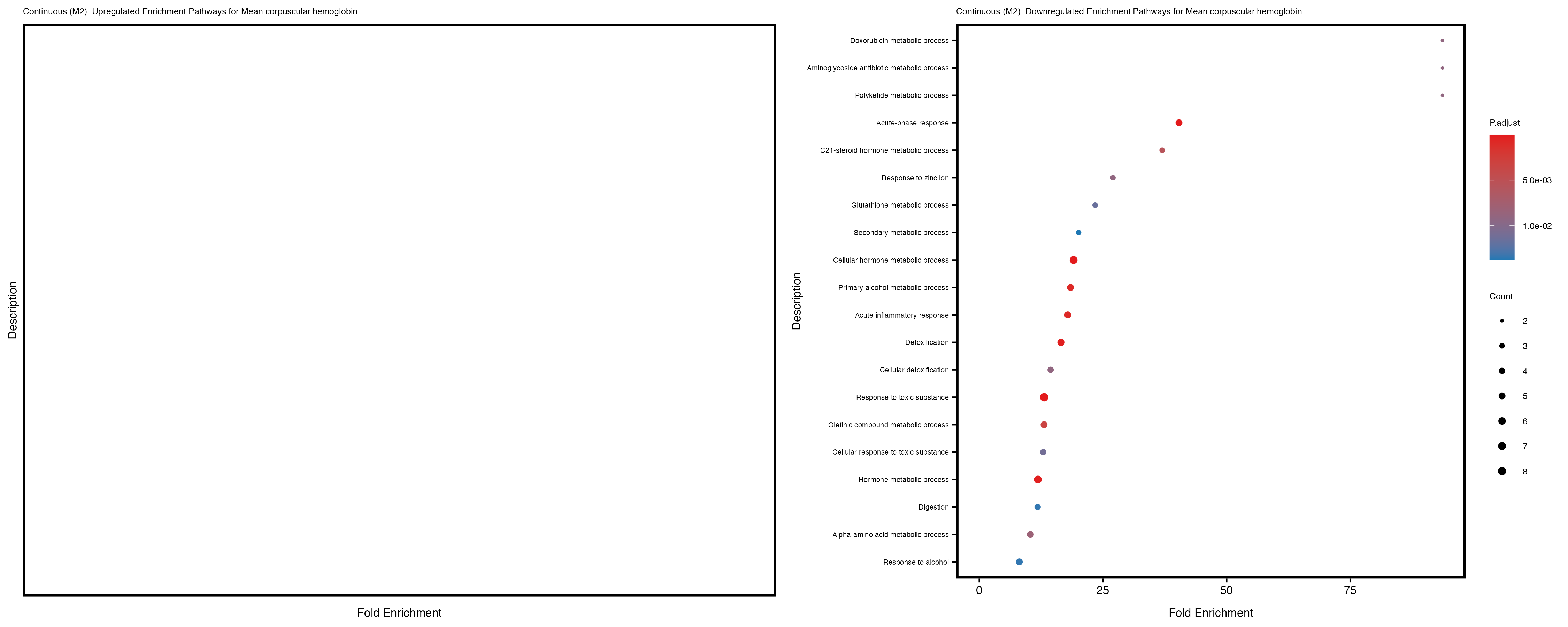
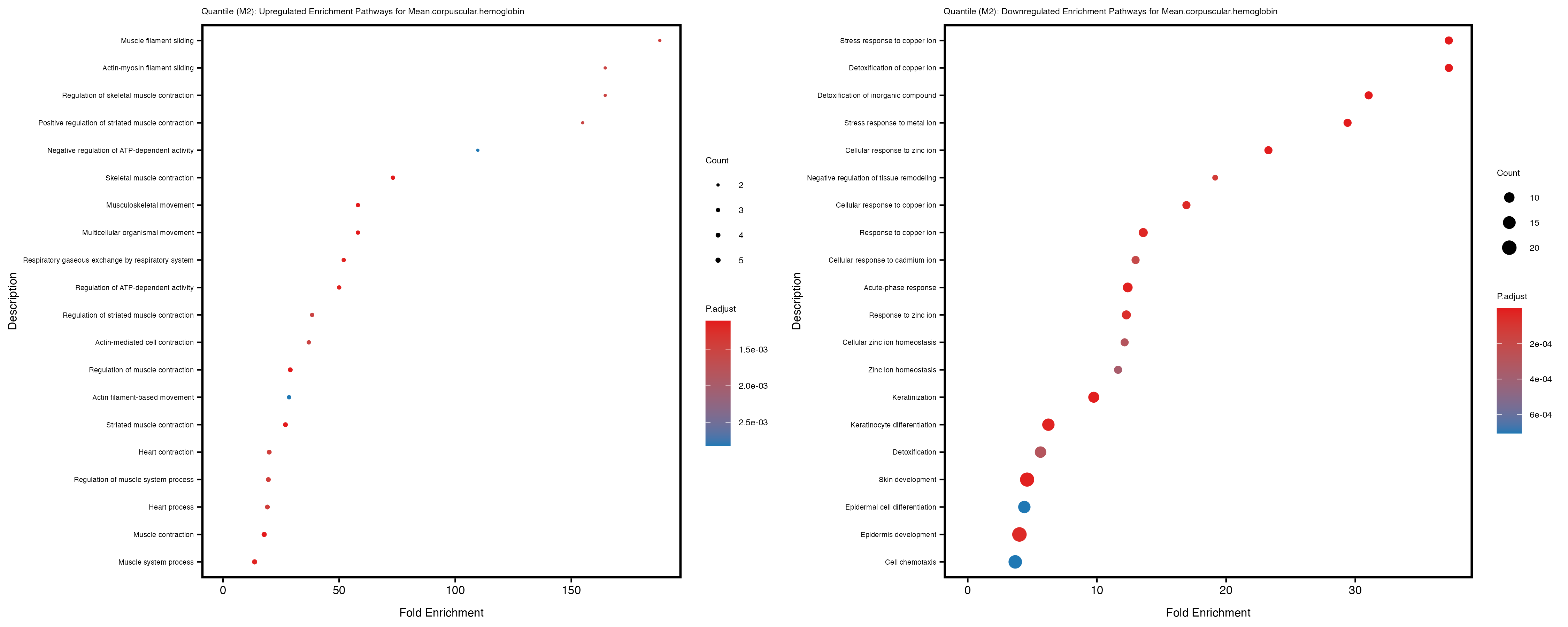
Mean.corpuscular.volume
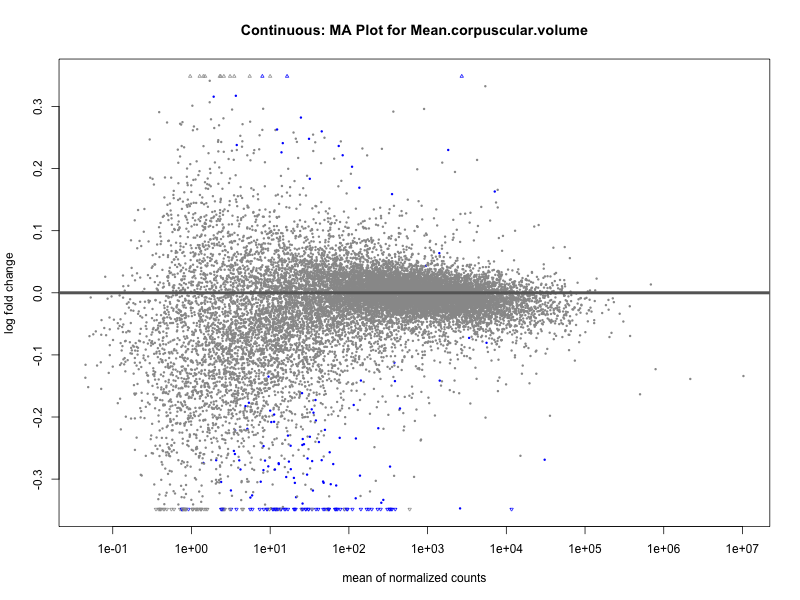
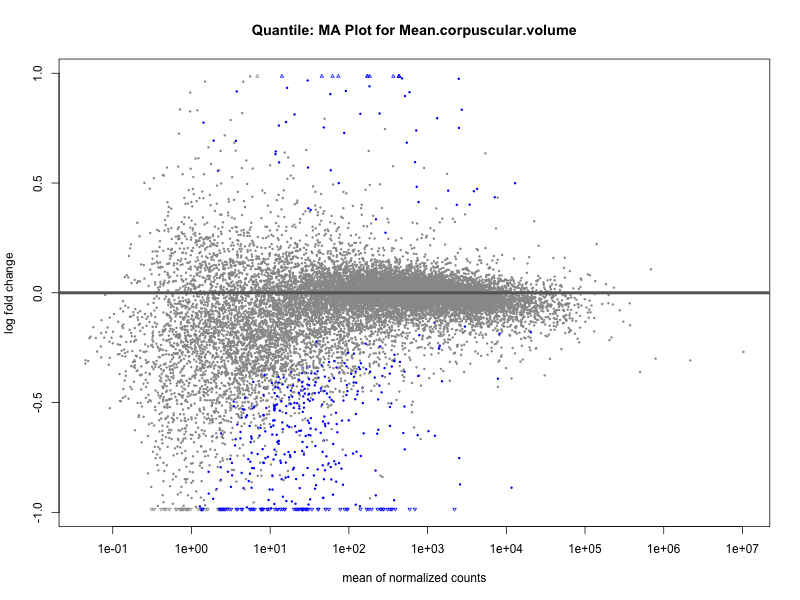
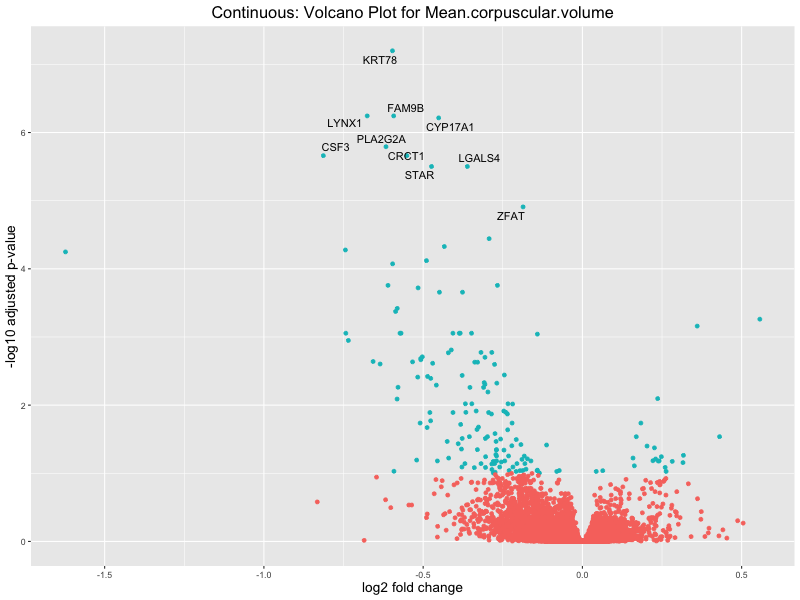
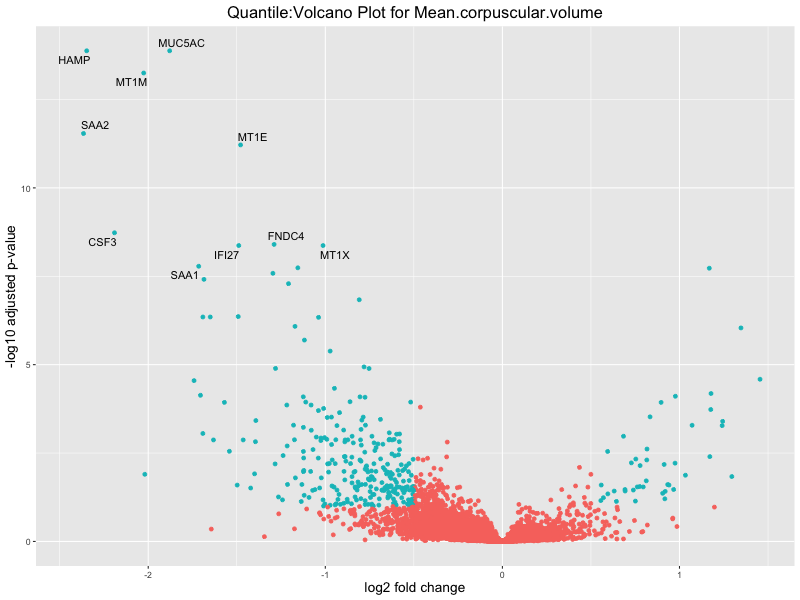
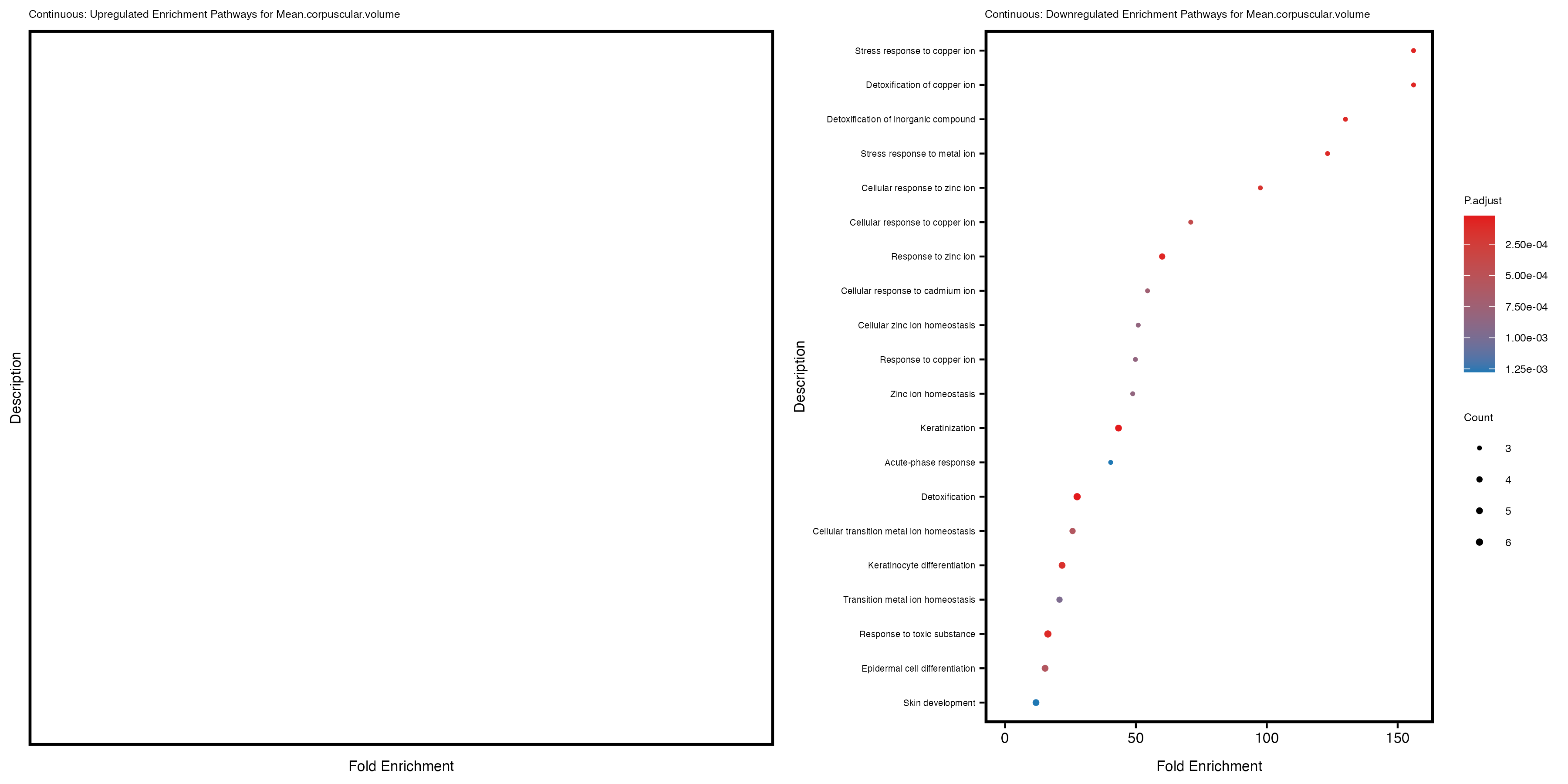
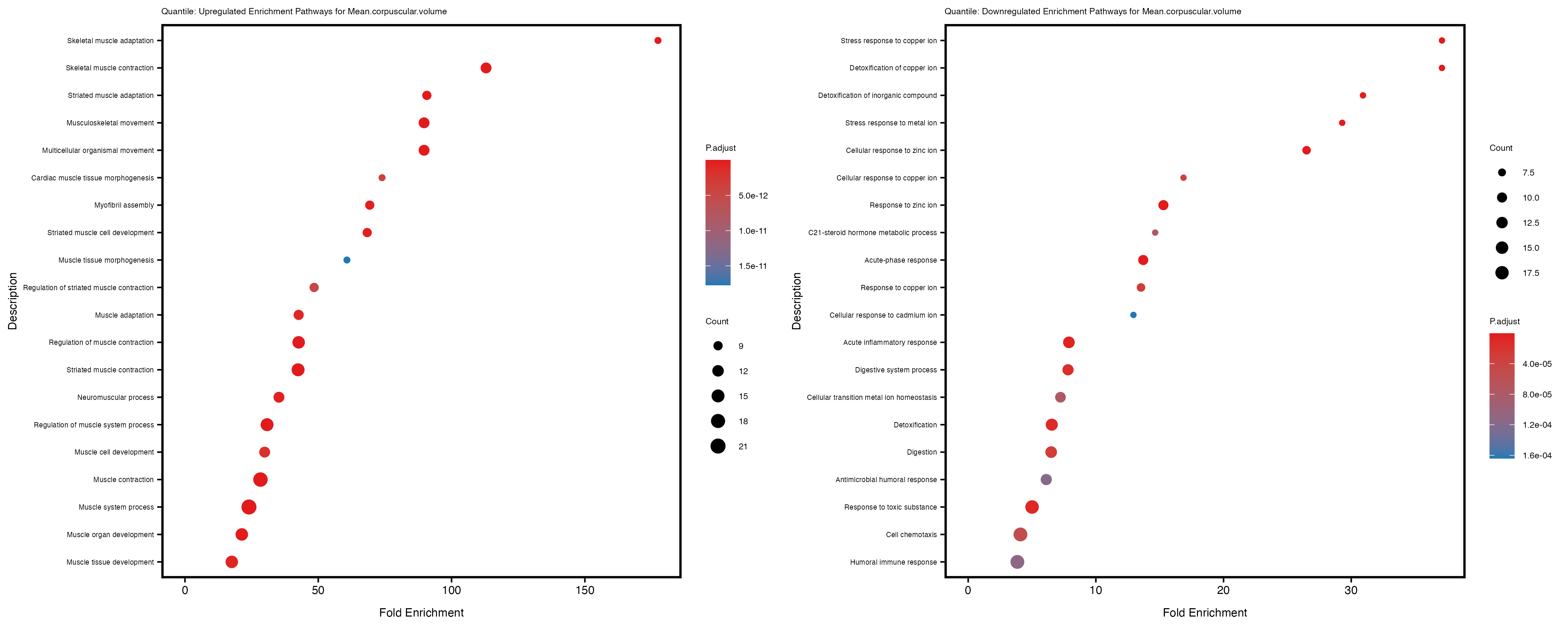
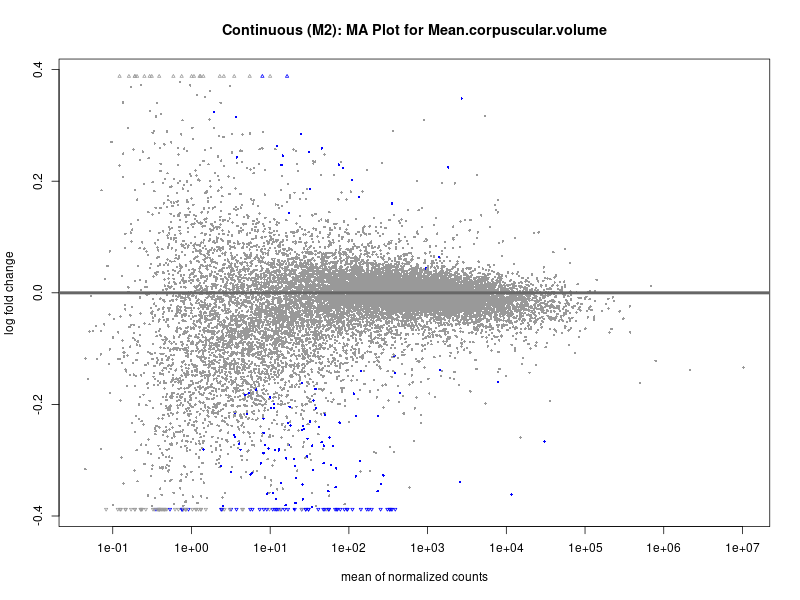
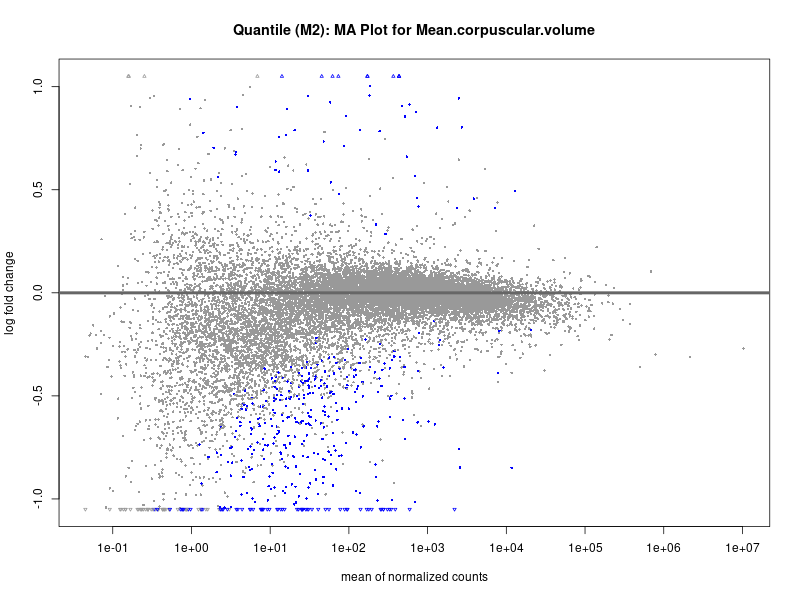
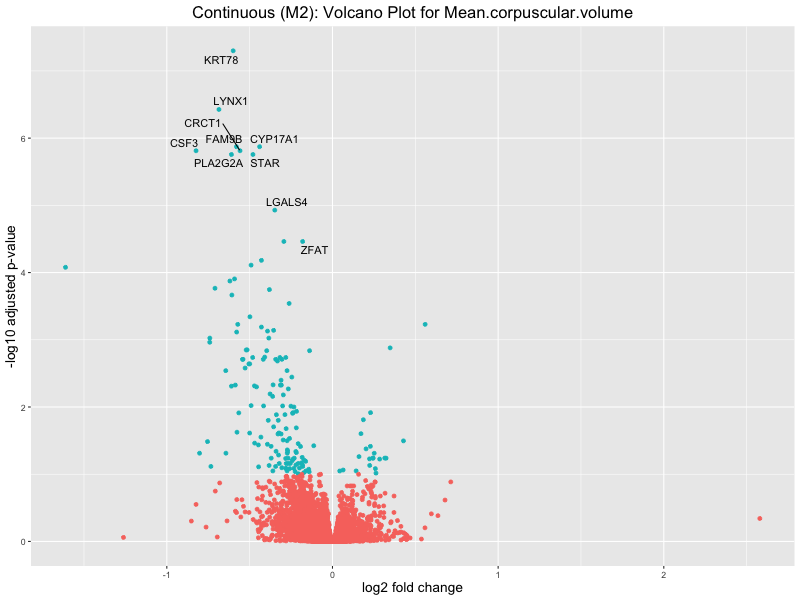
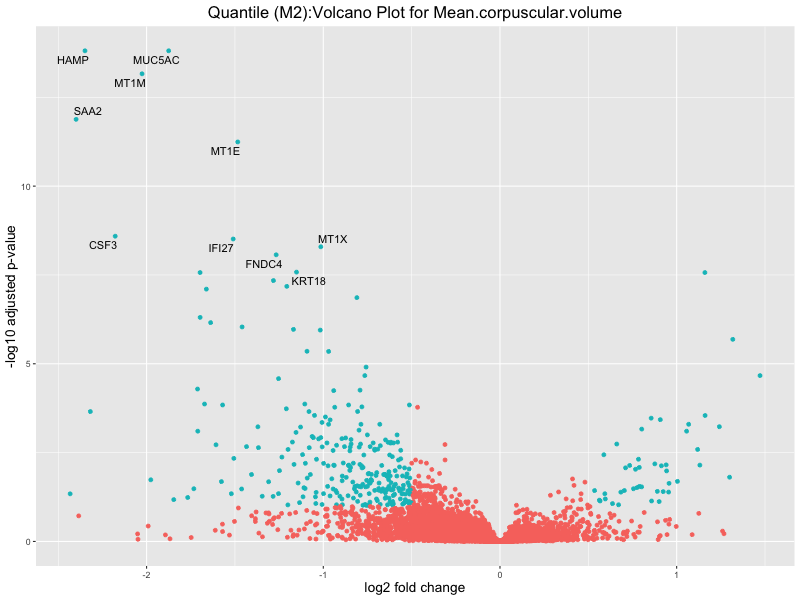
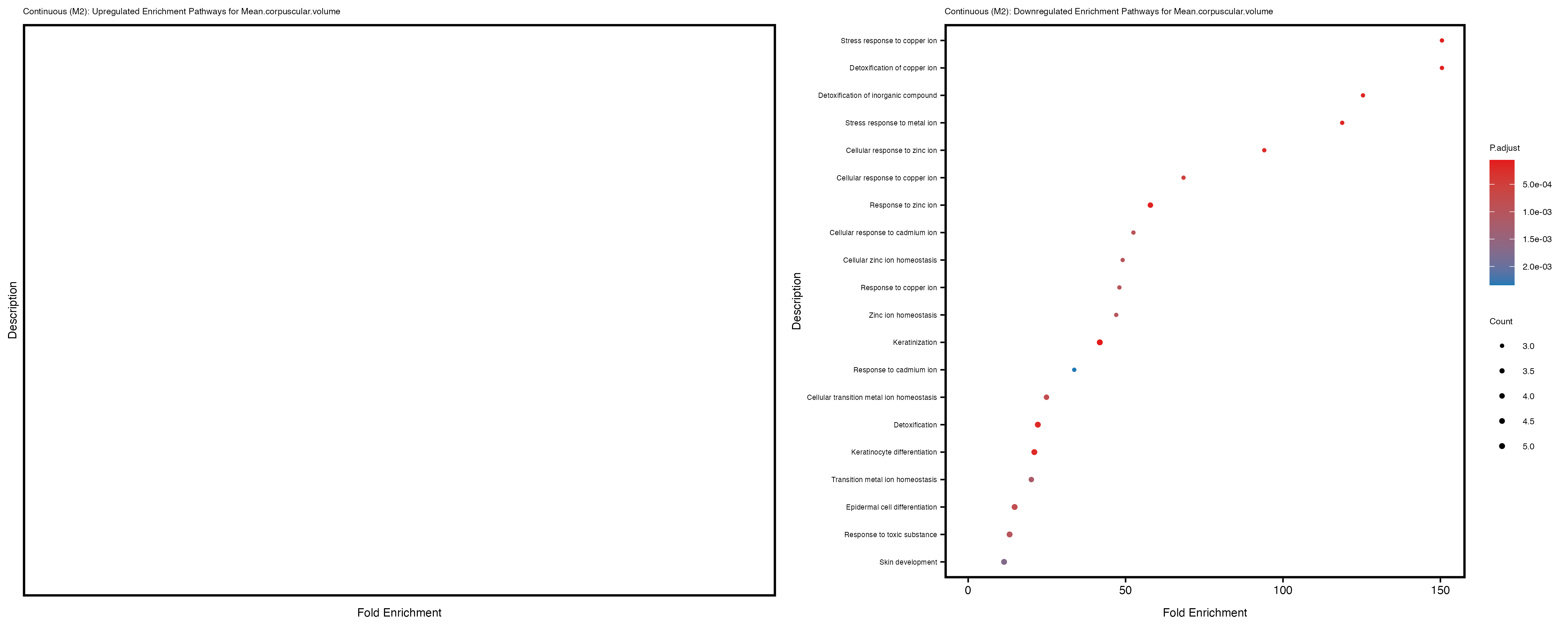
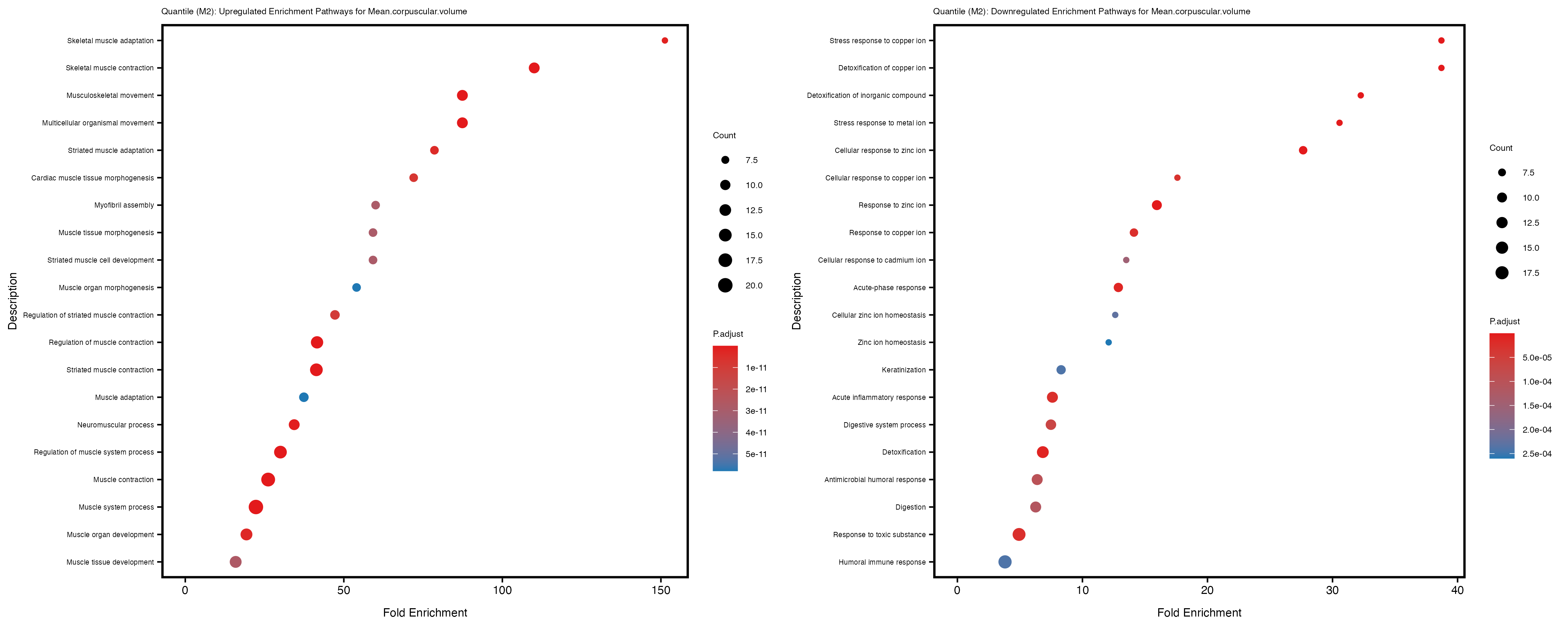
Mean.platelet.volume
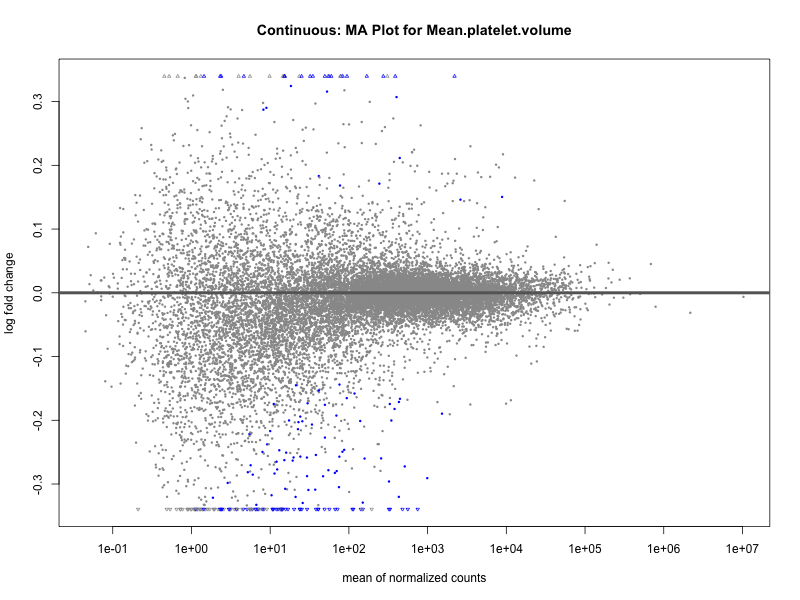
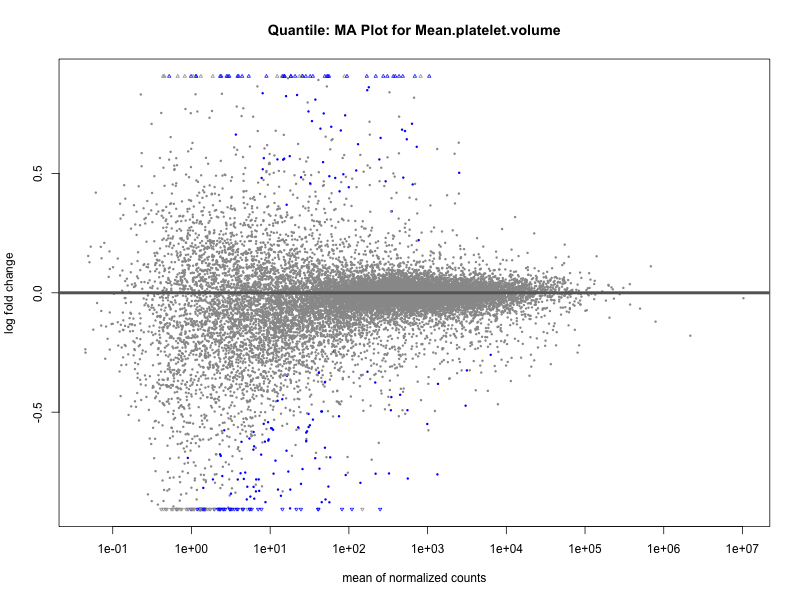
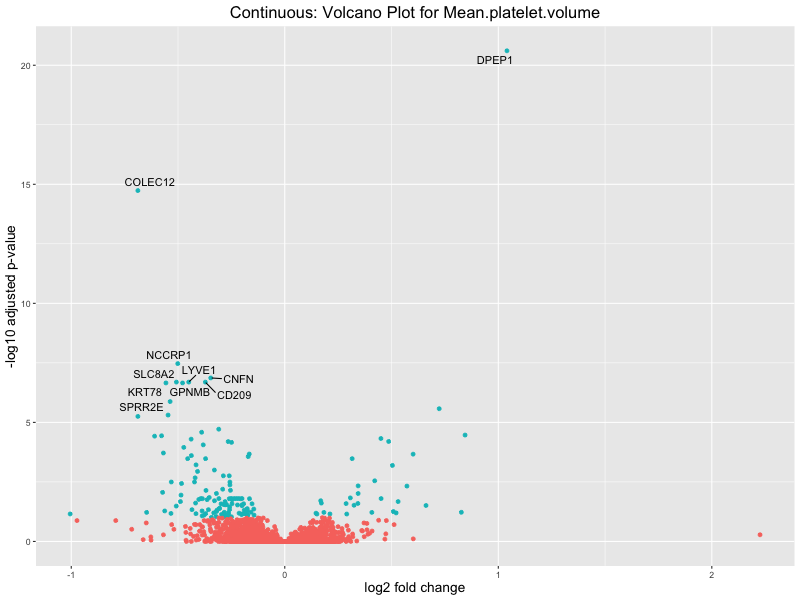
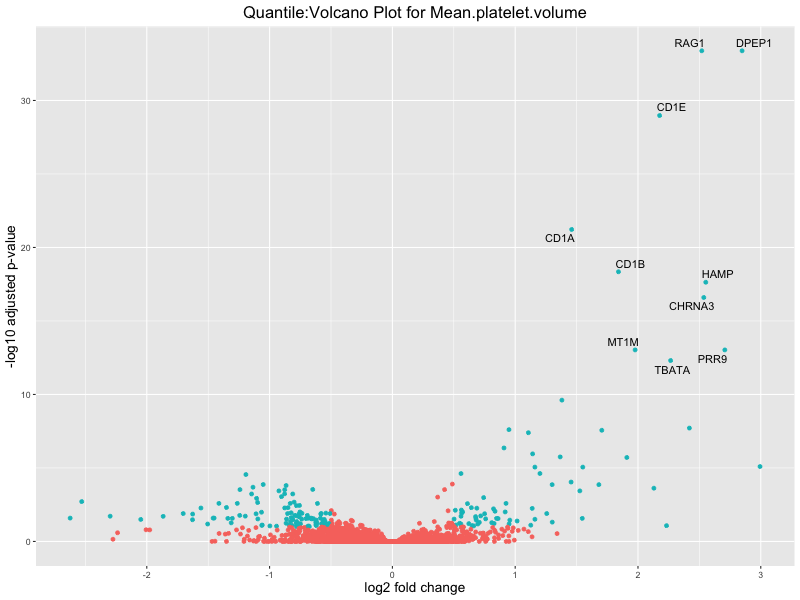
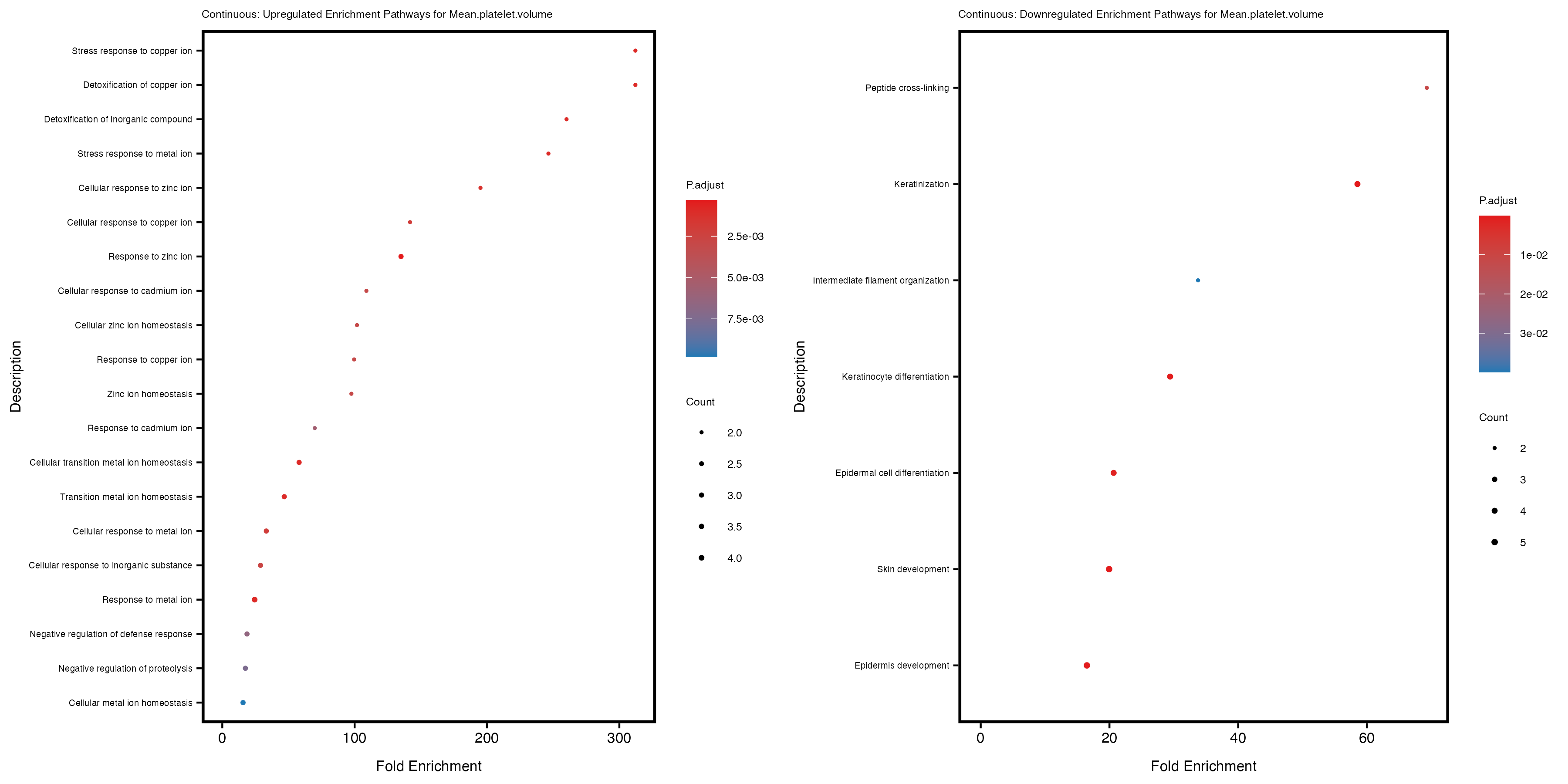
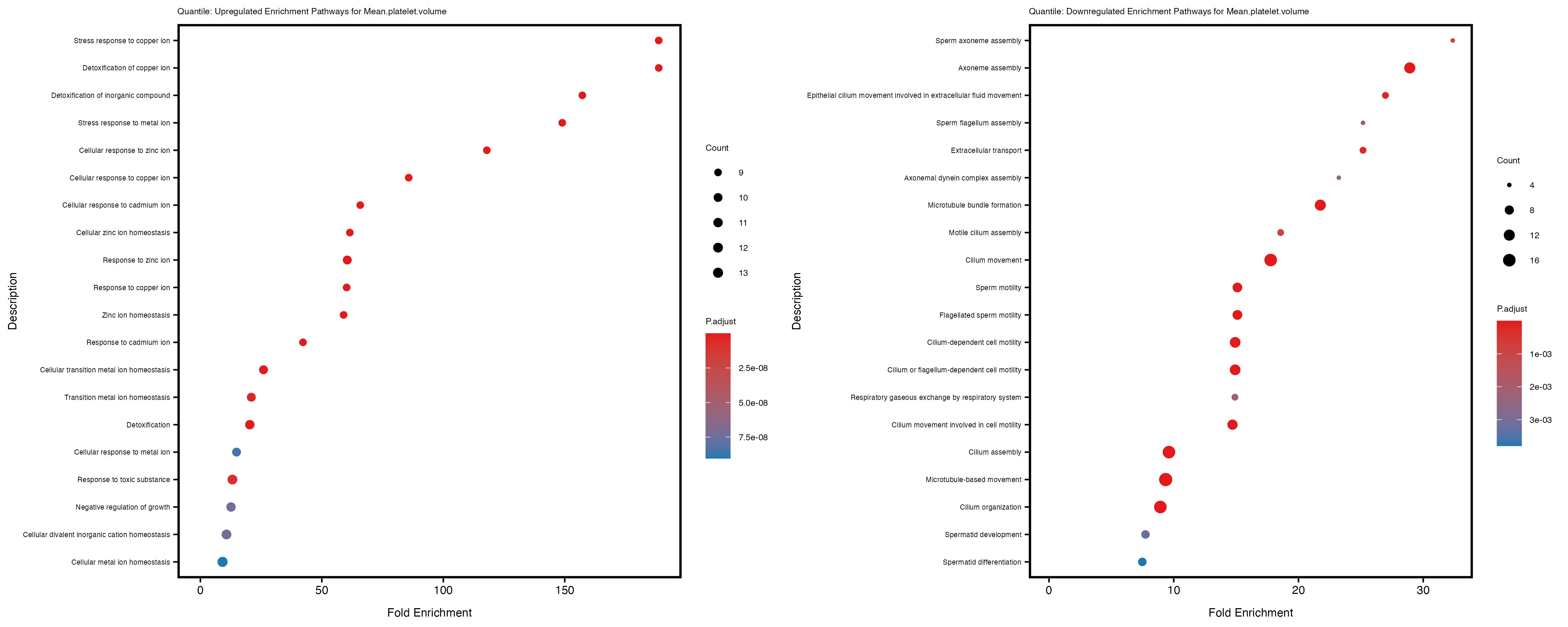
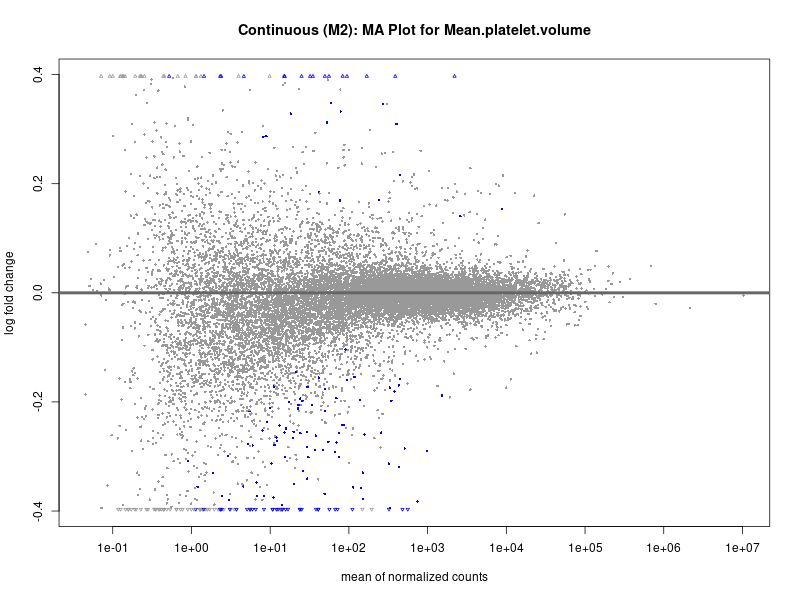
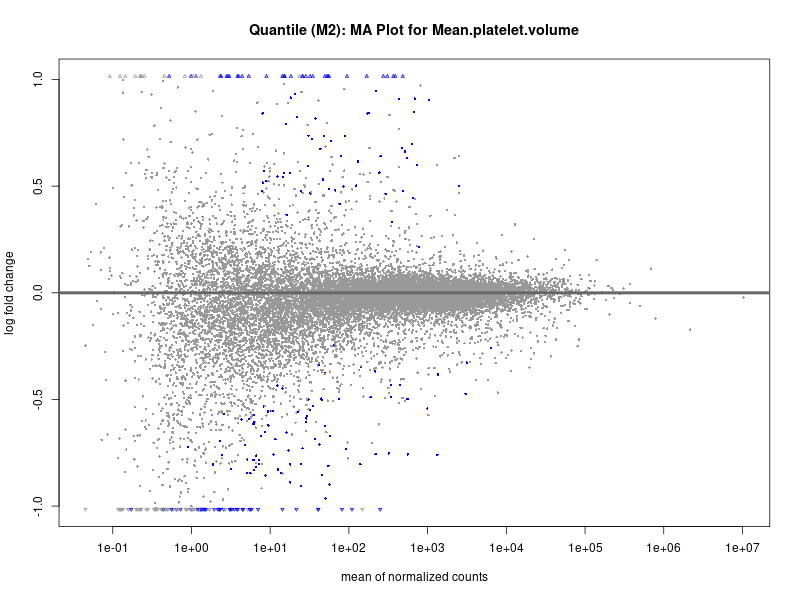
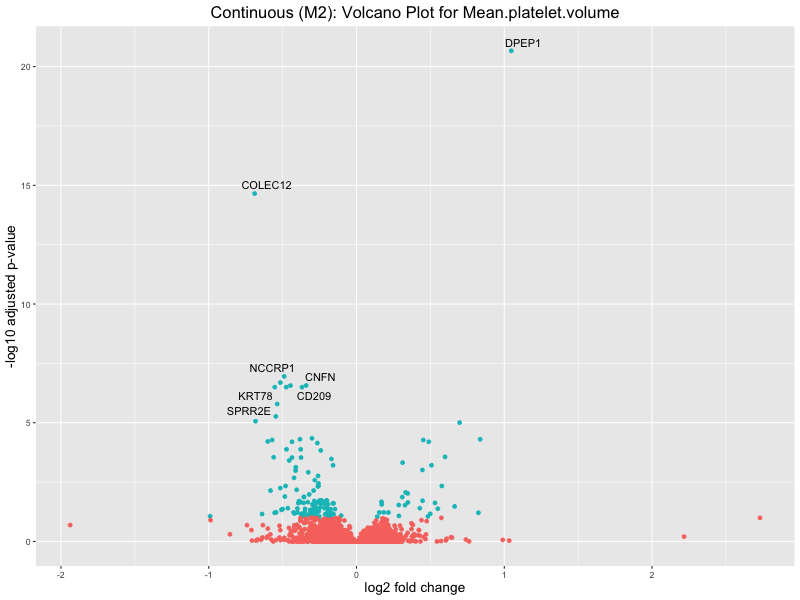
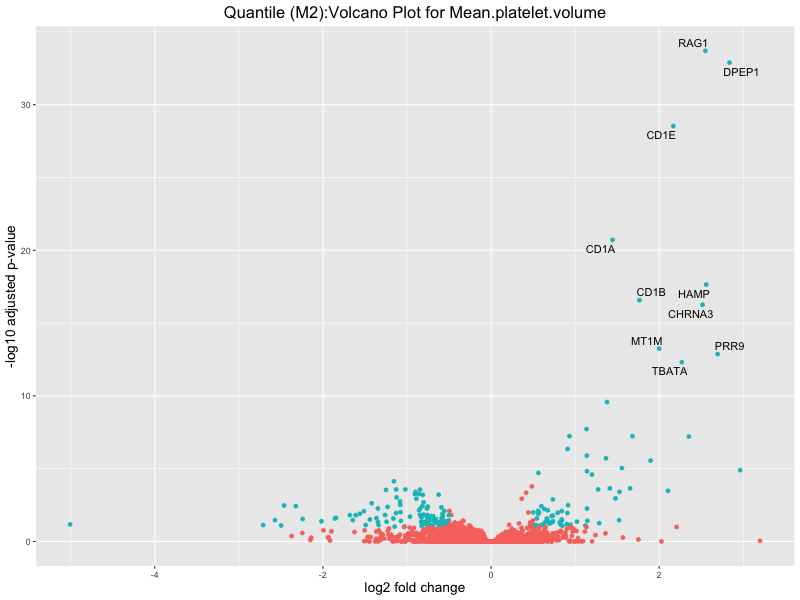
Mean.reticulocyte.volume
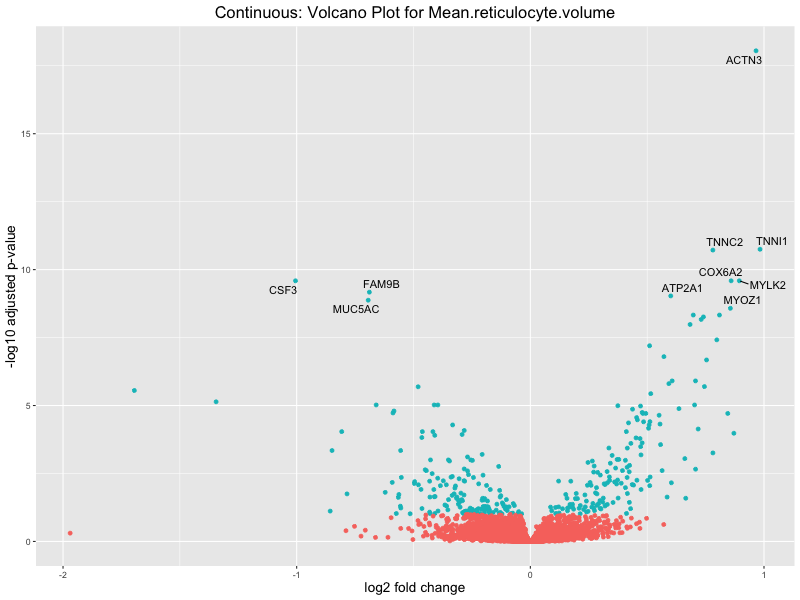
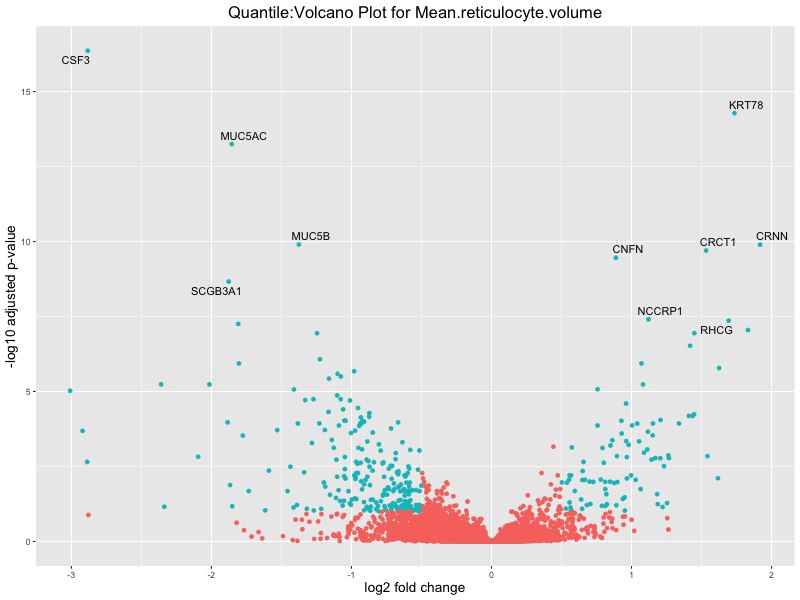
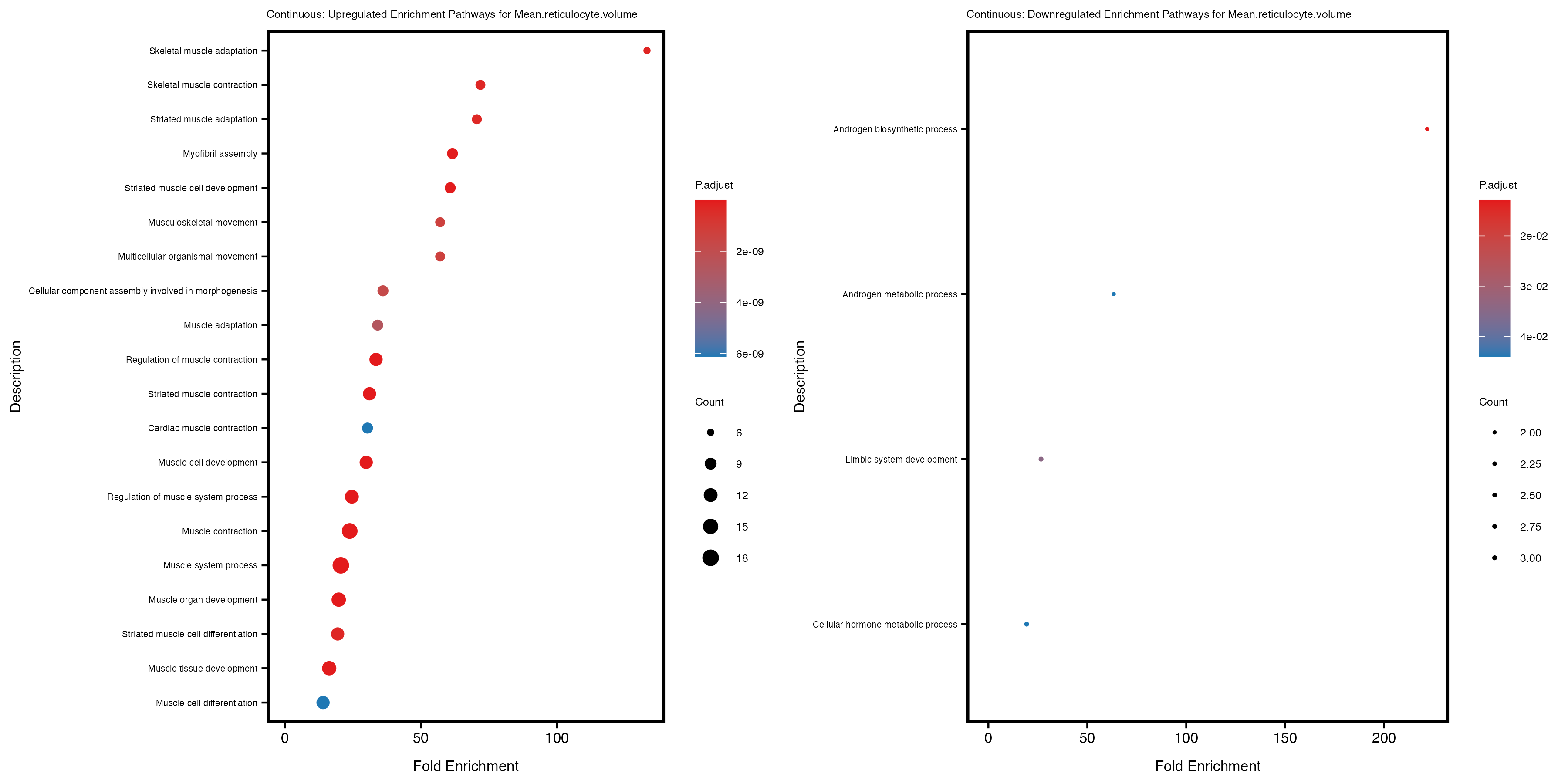
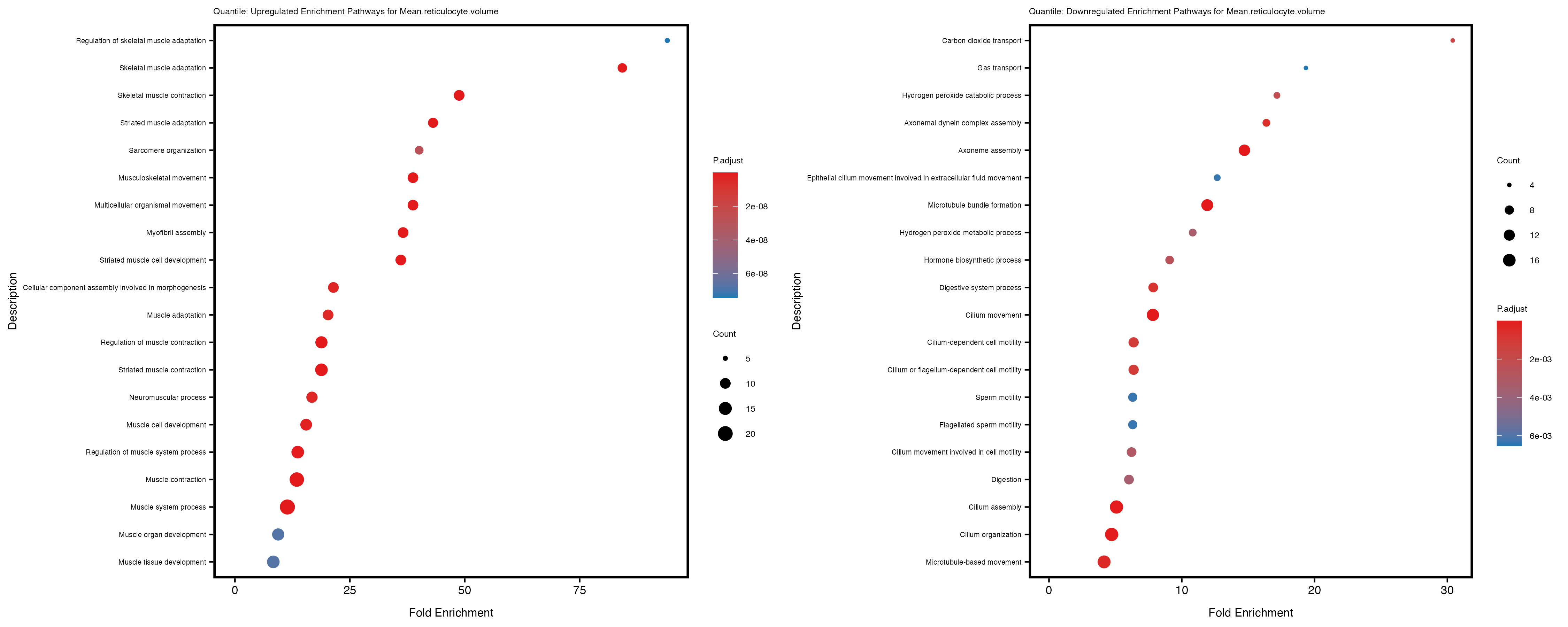
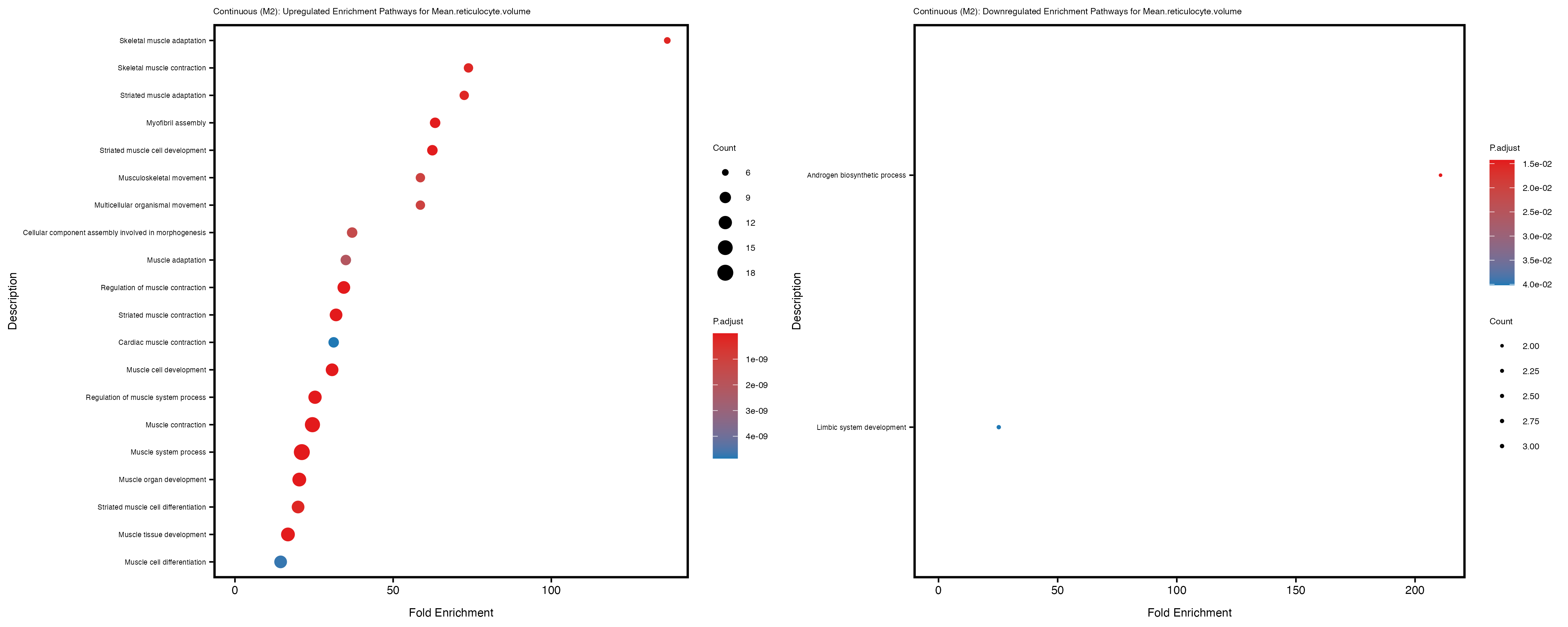
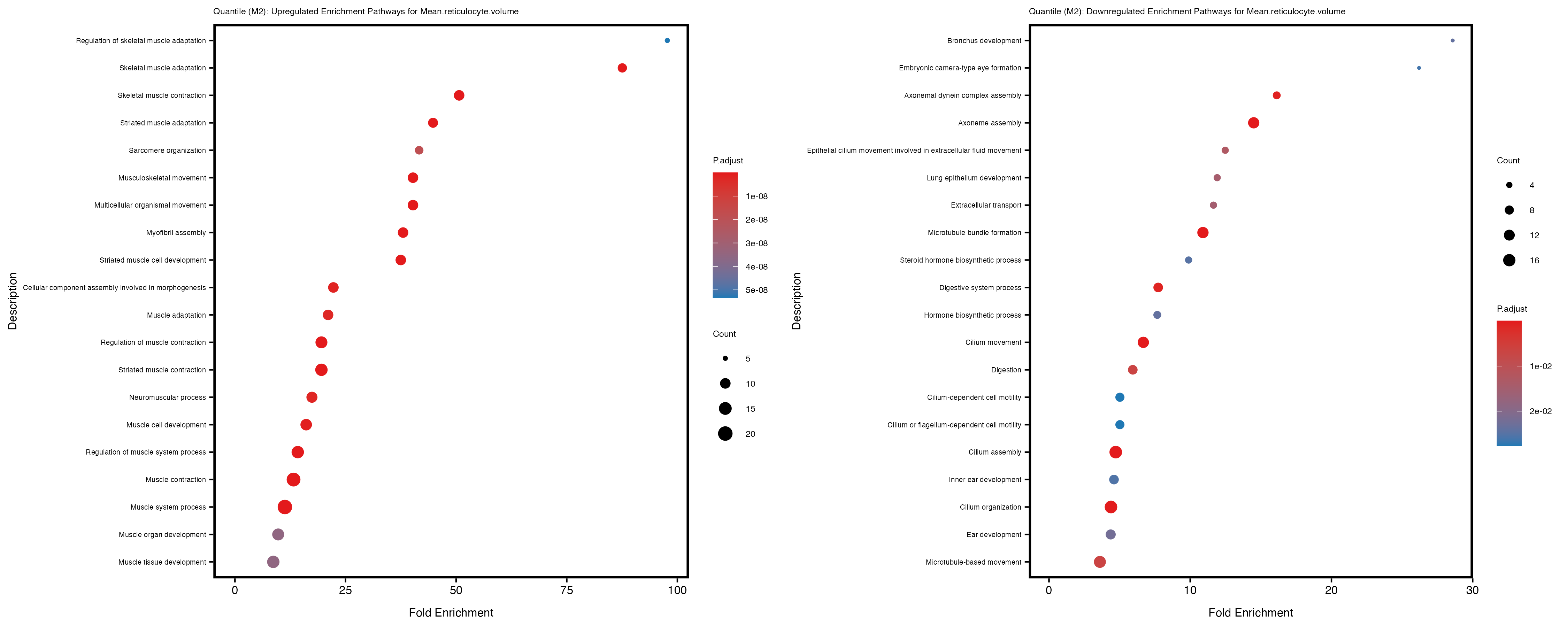
Mean.sphered.corpuscular.volume
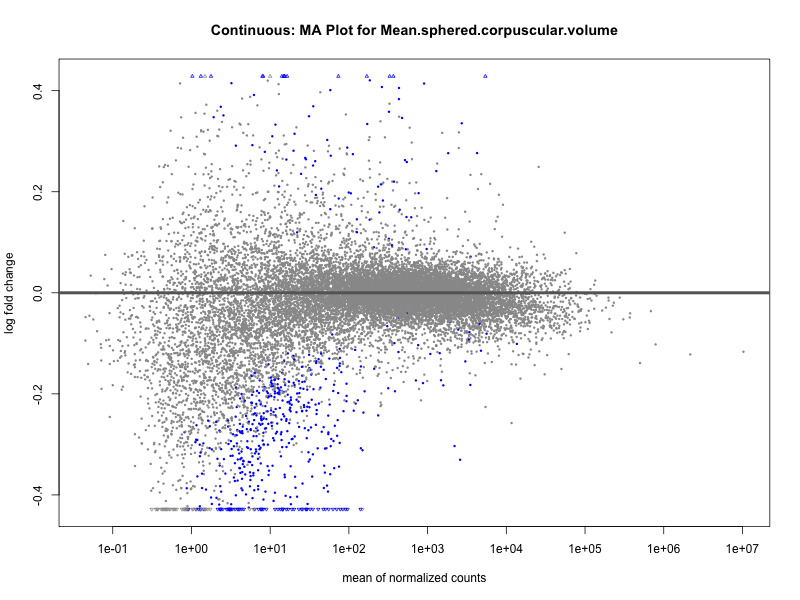
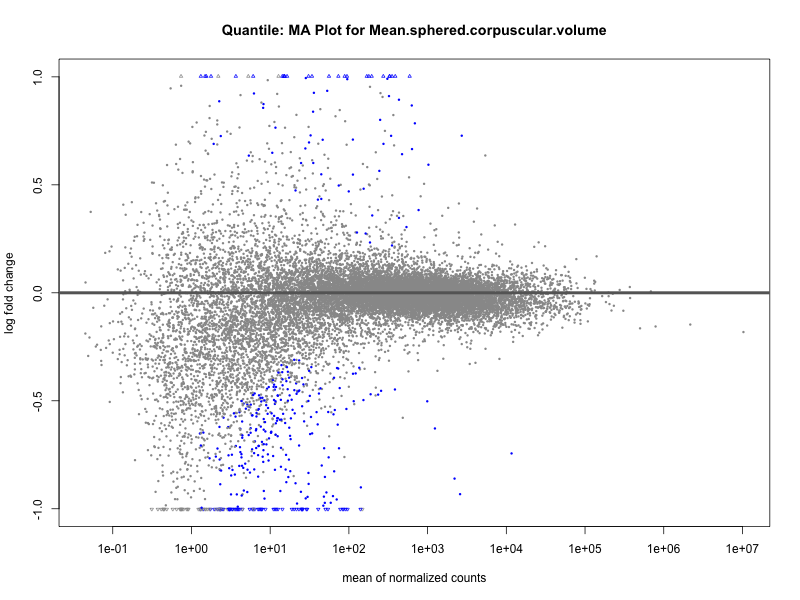
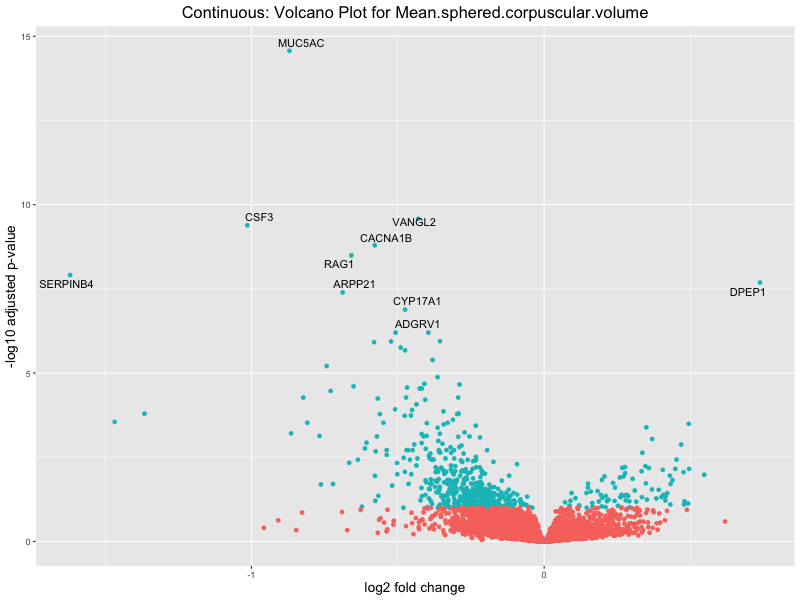
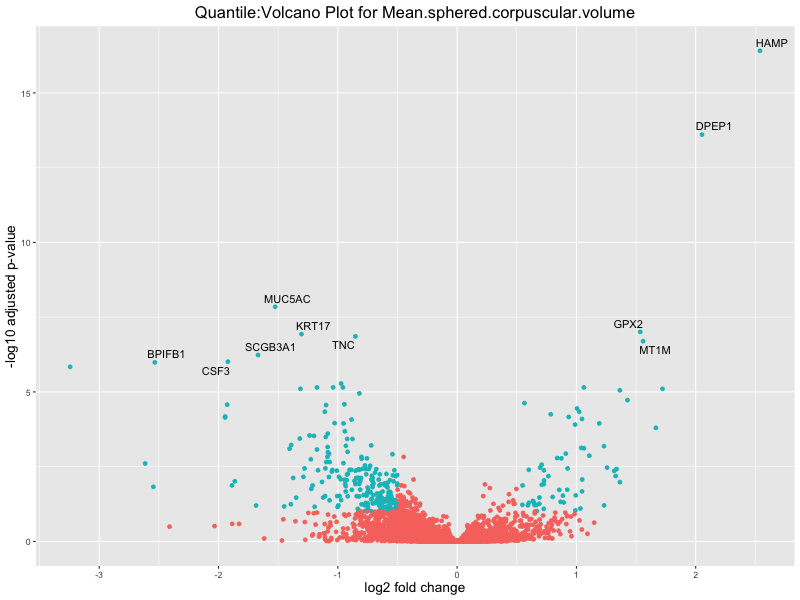
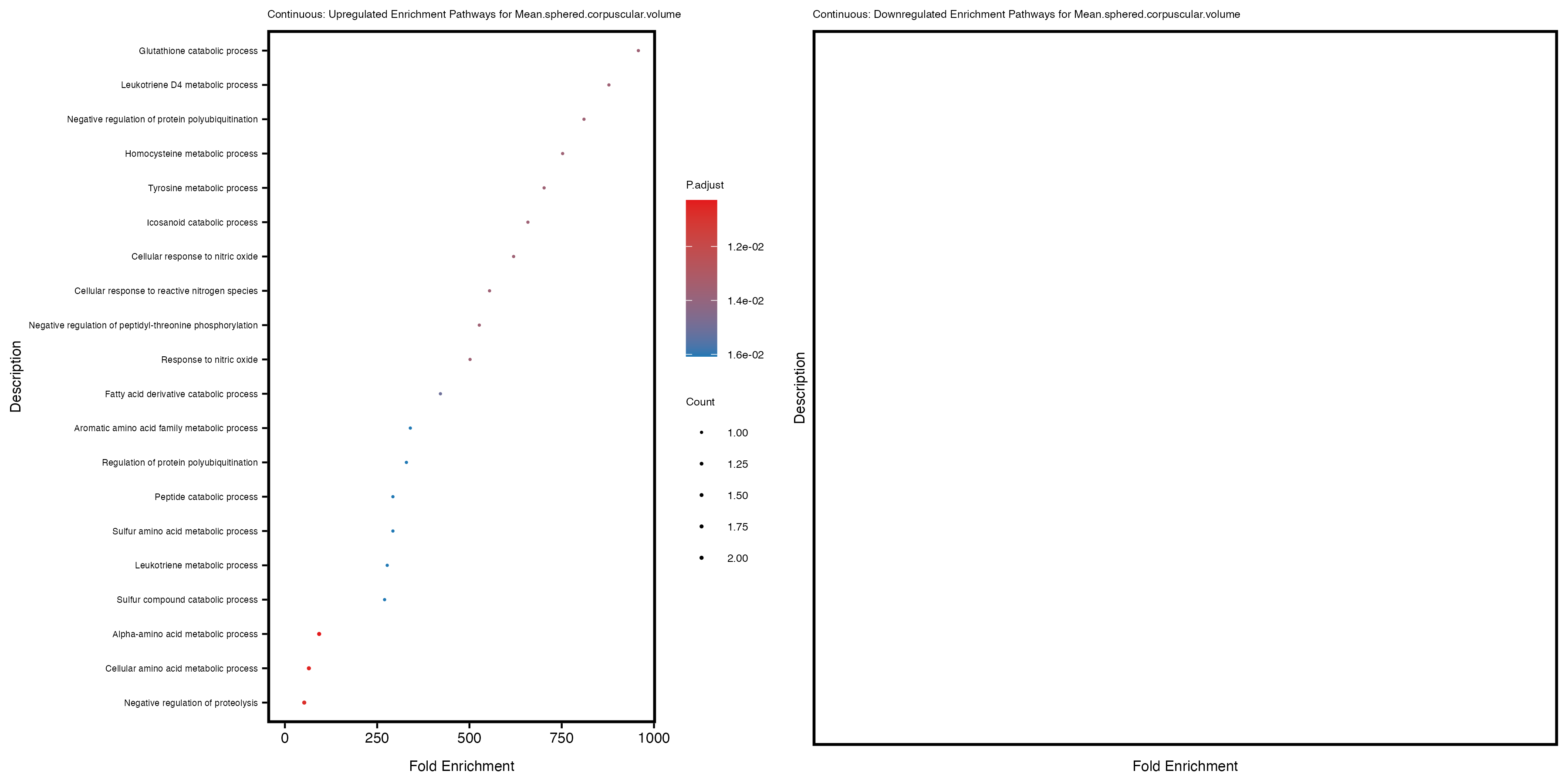
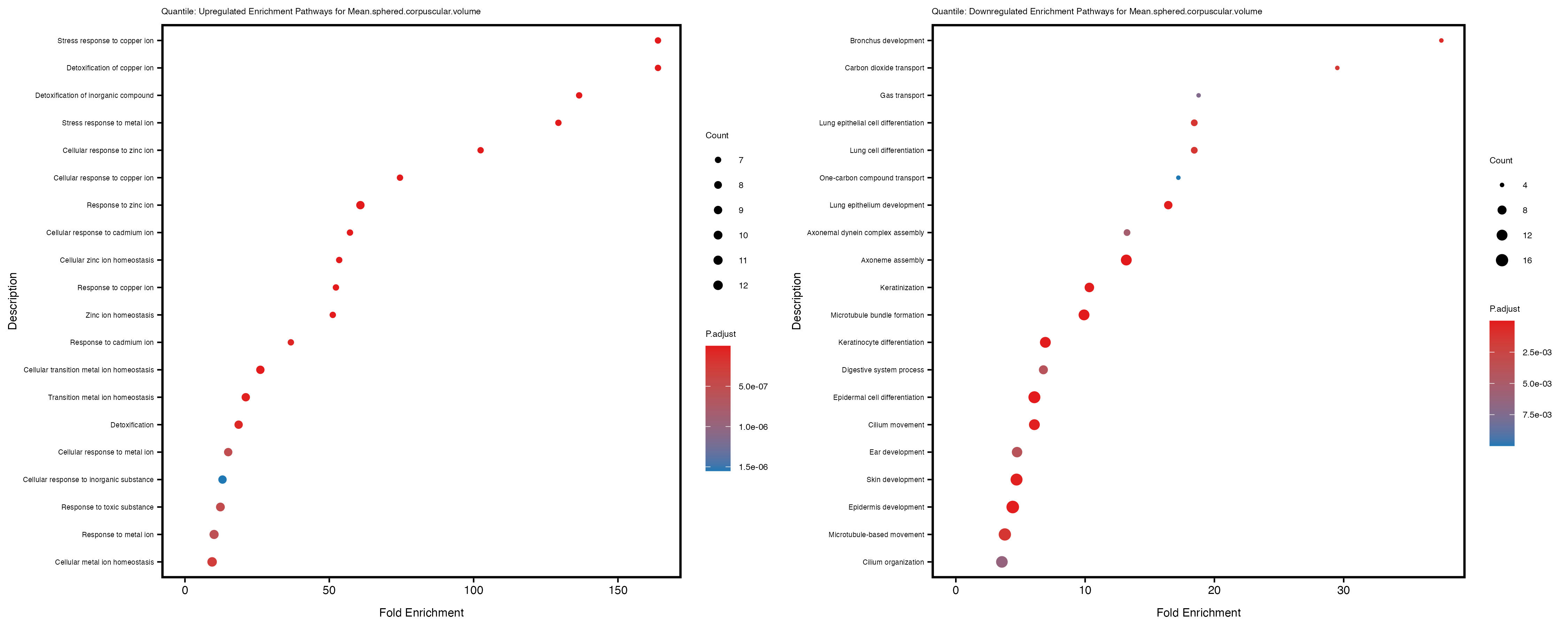
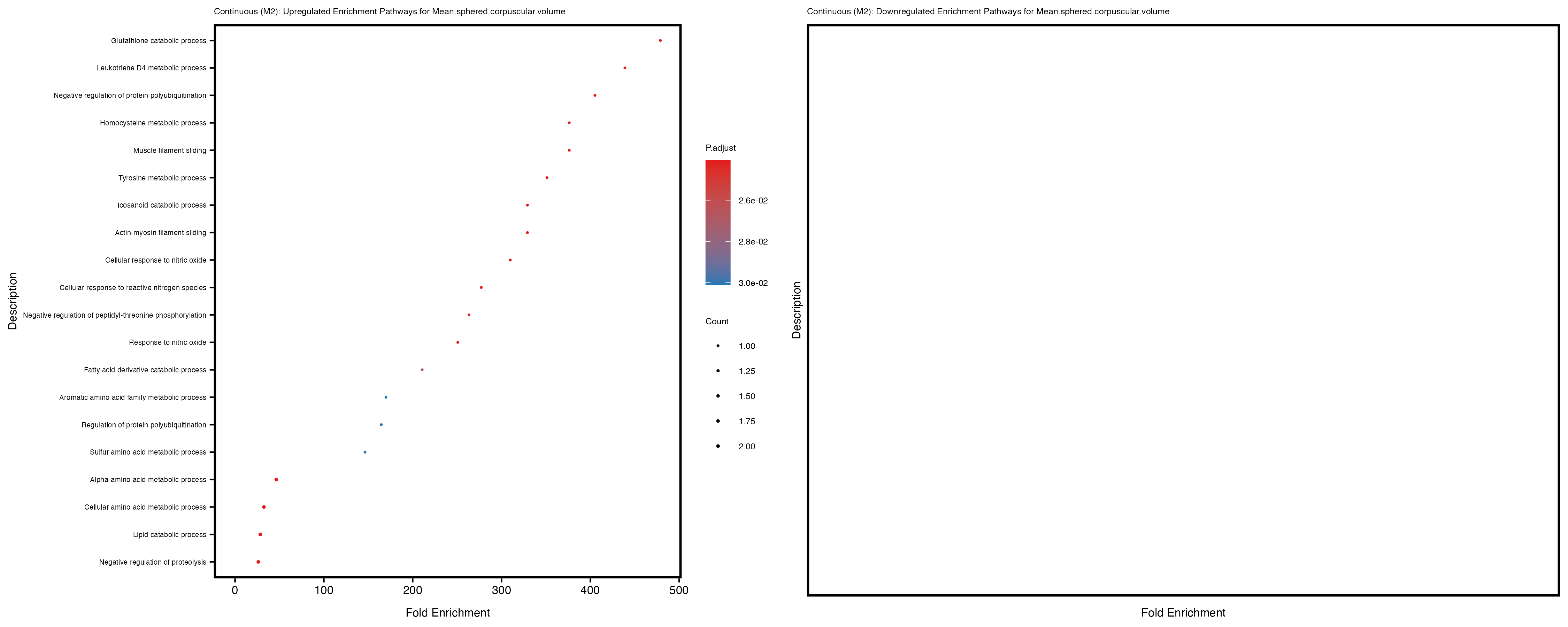
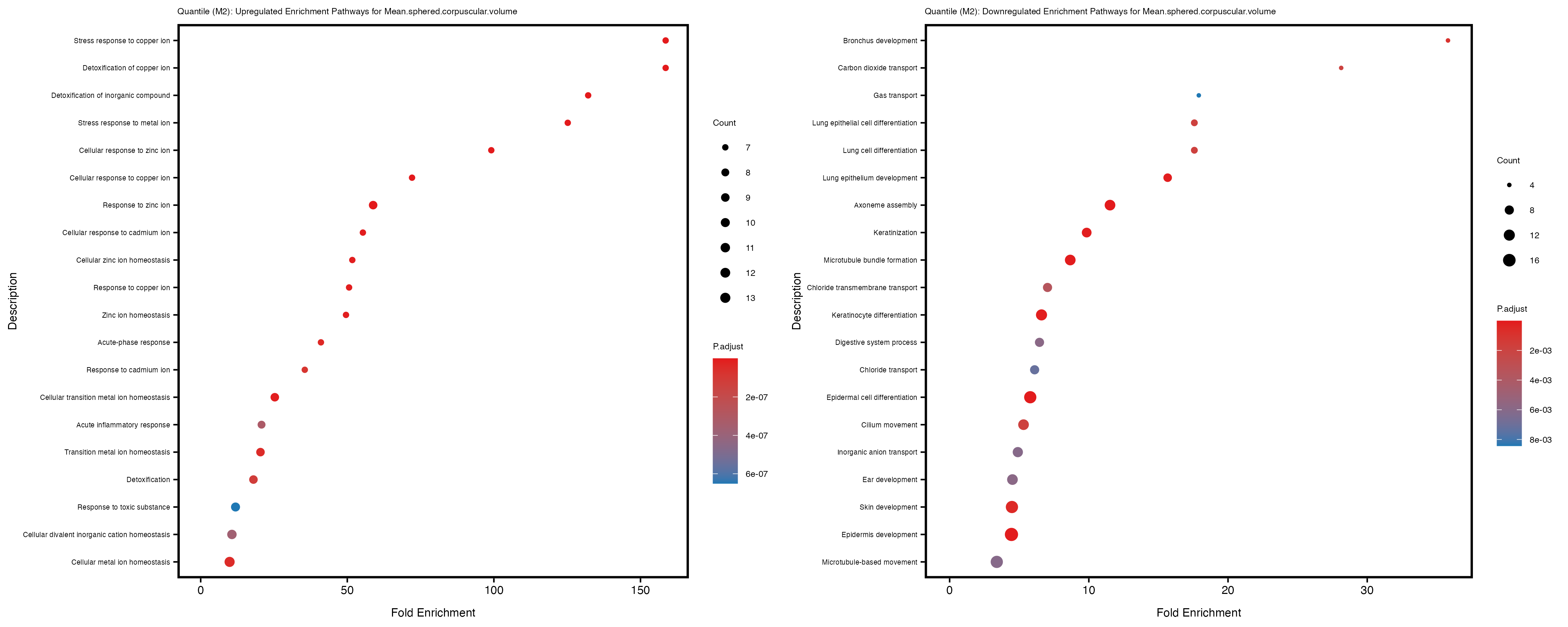
Monocyte.count
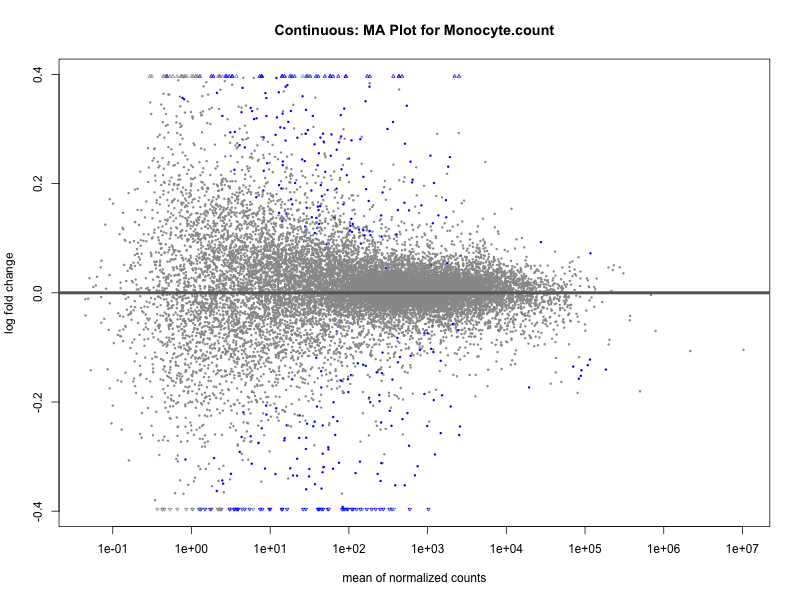
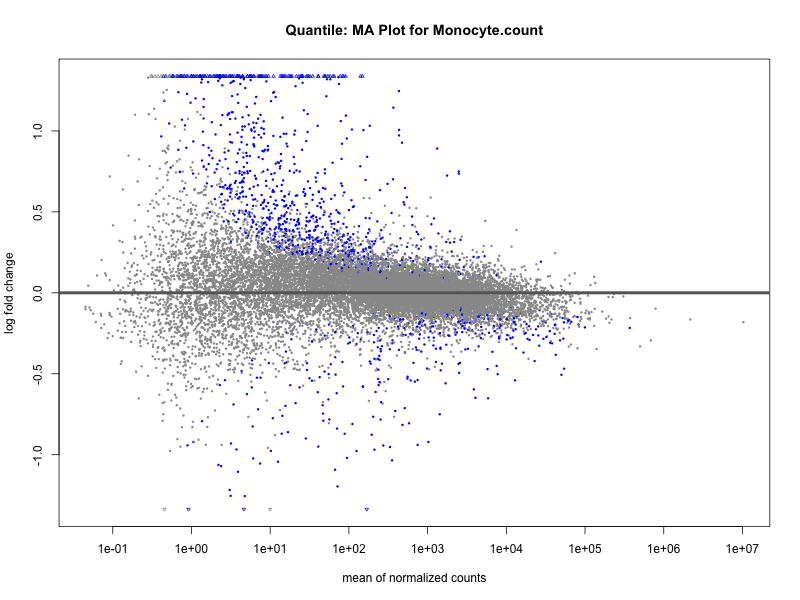
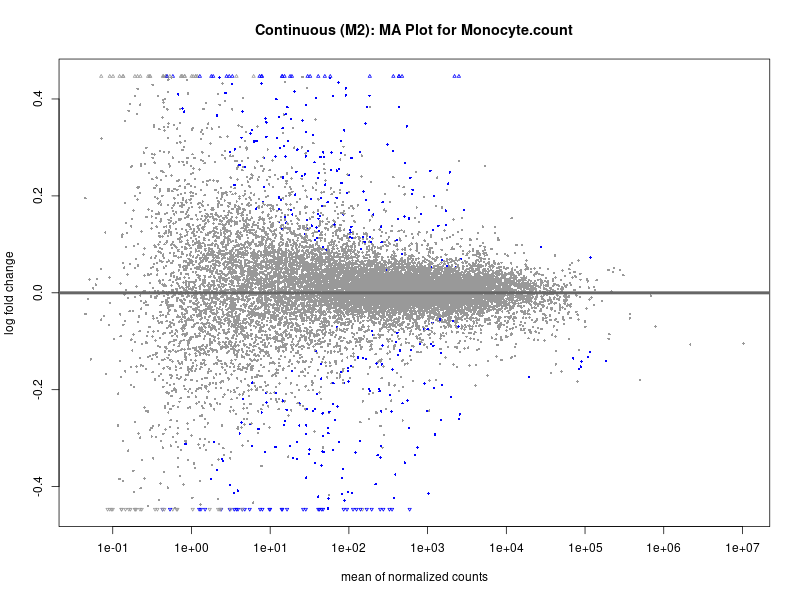
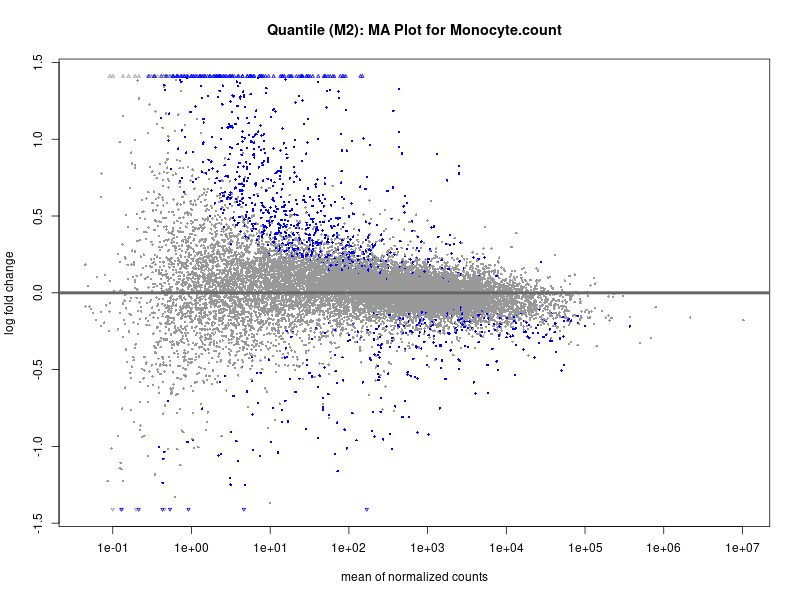
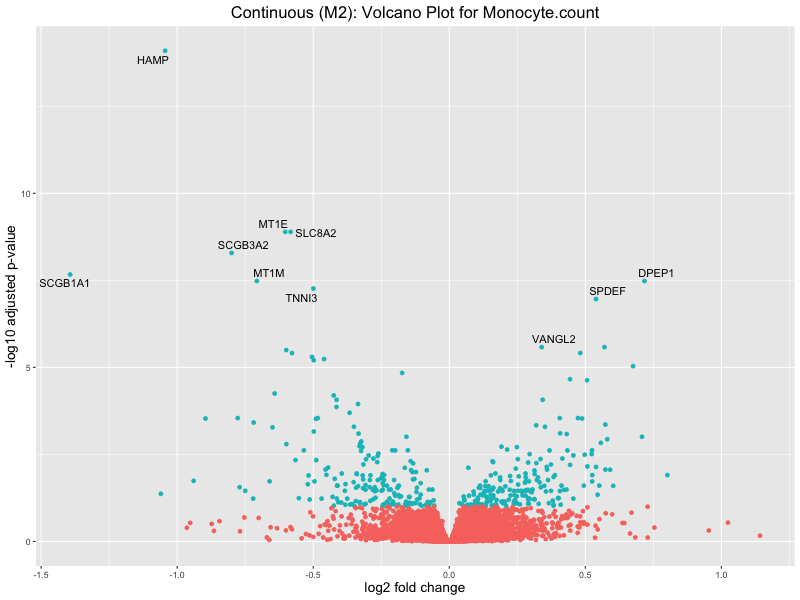
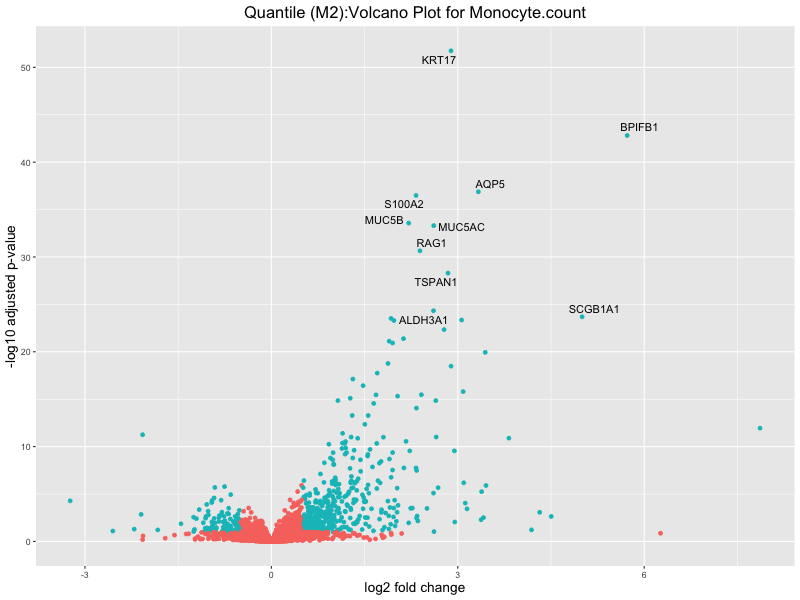
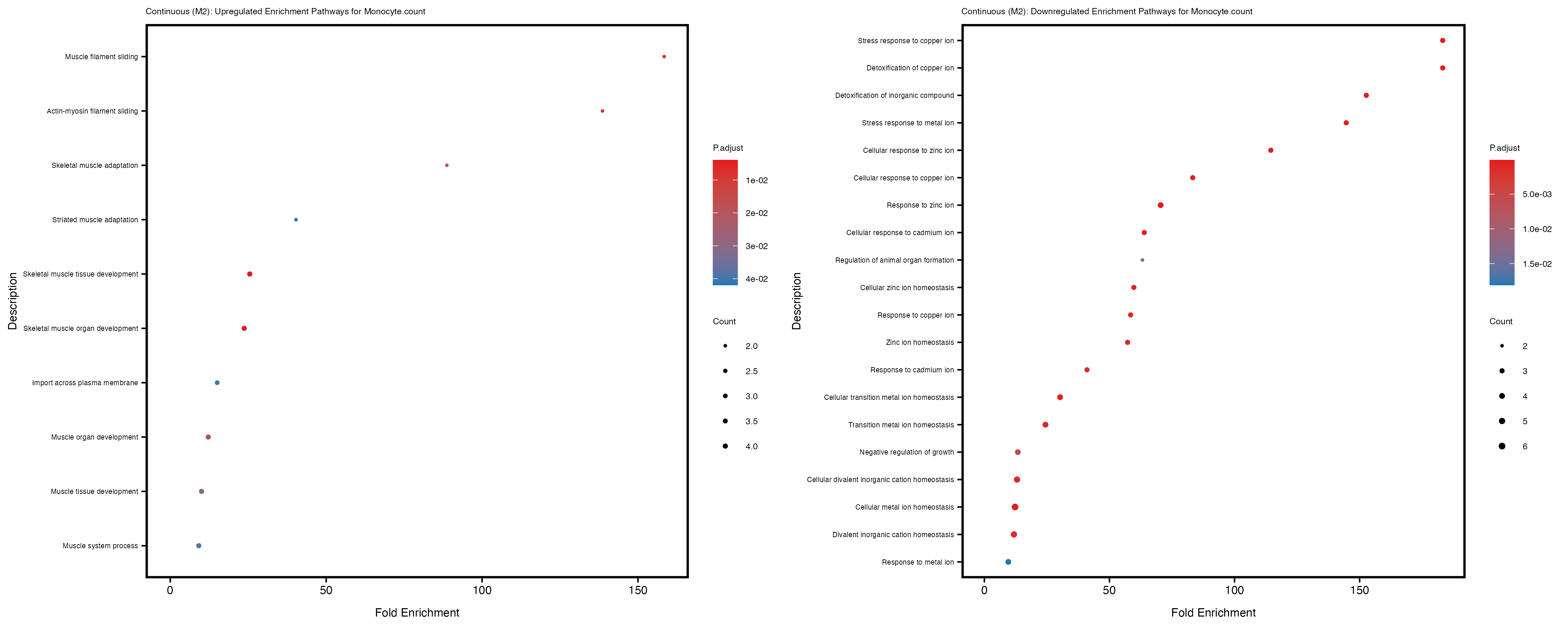
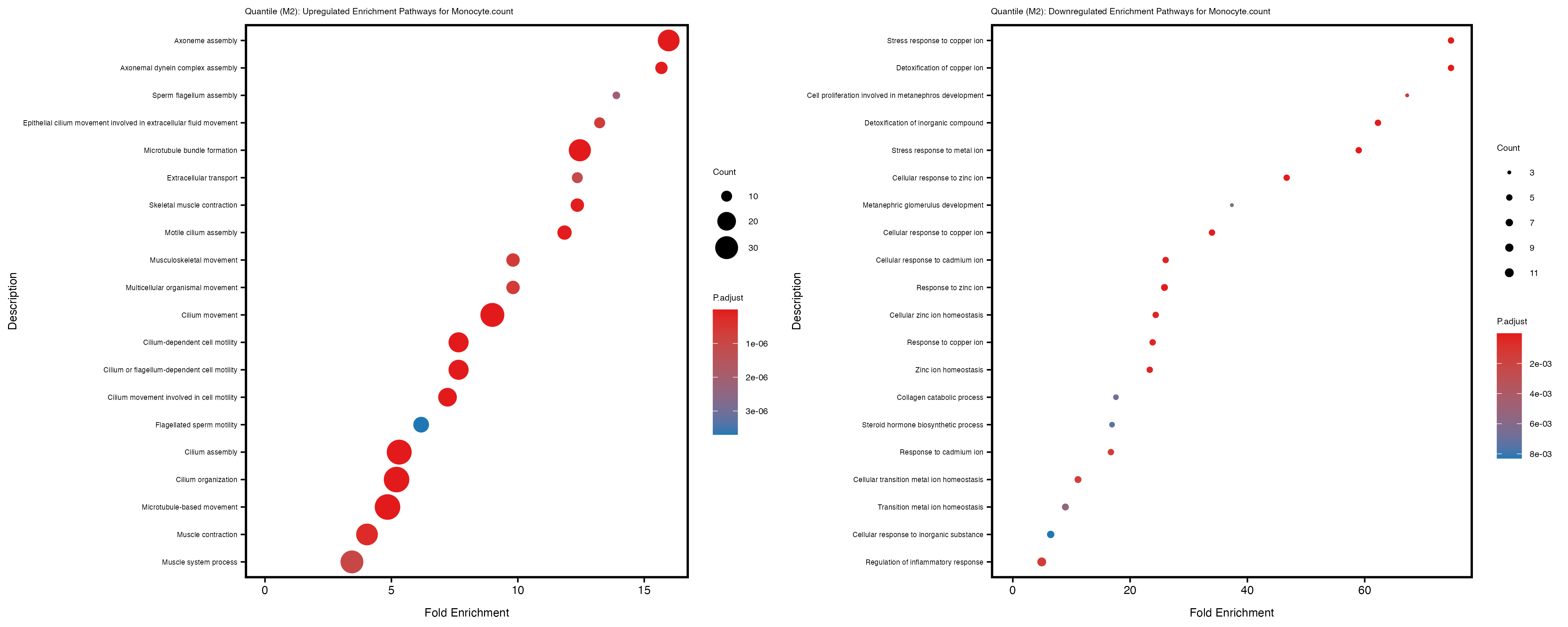
Monocyte.percentage.of.white.cells
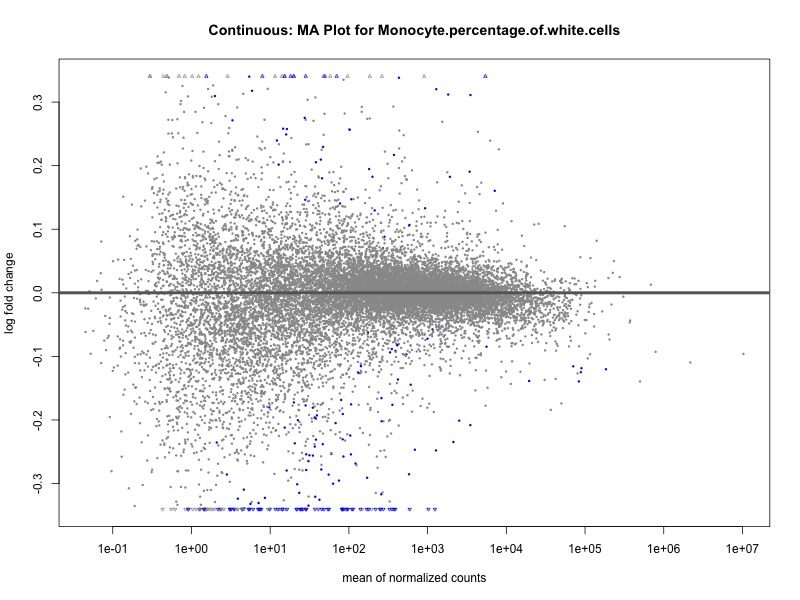
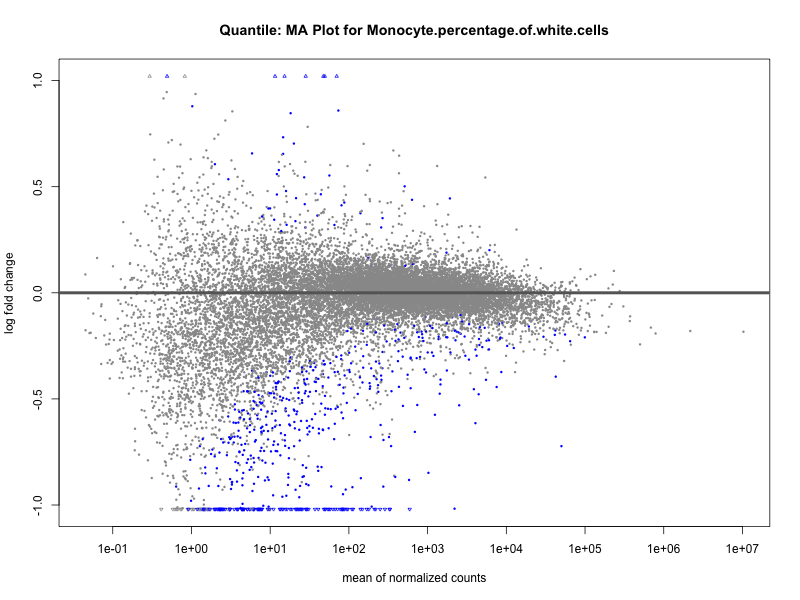
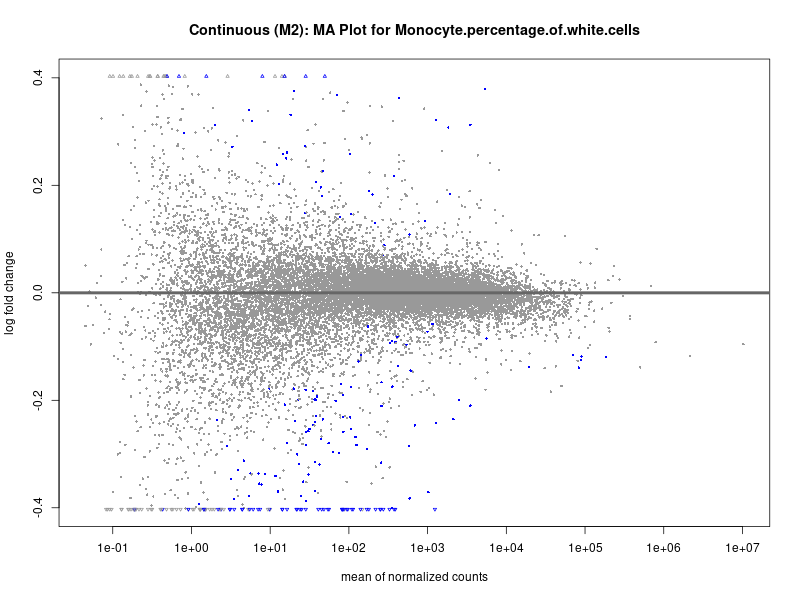
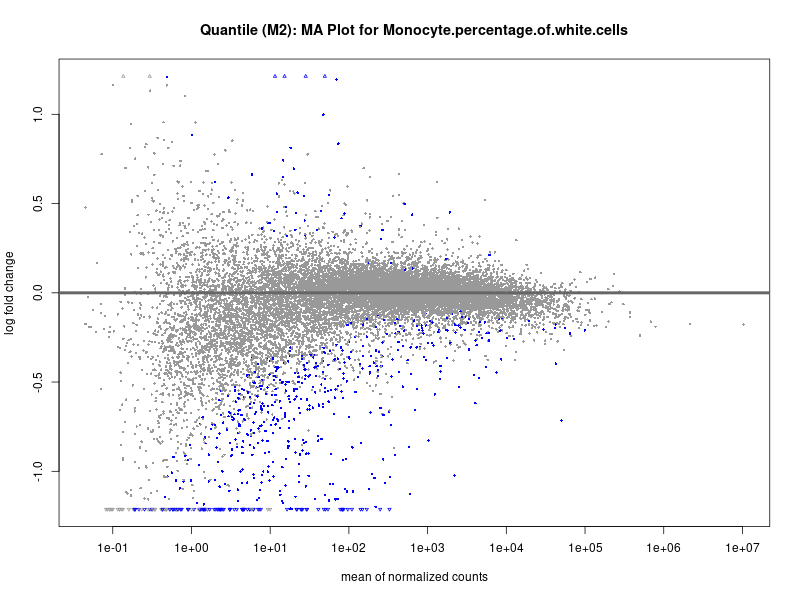
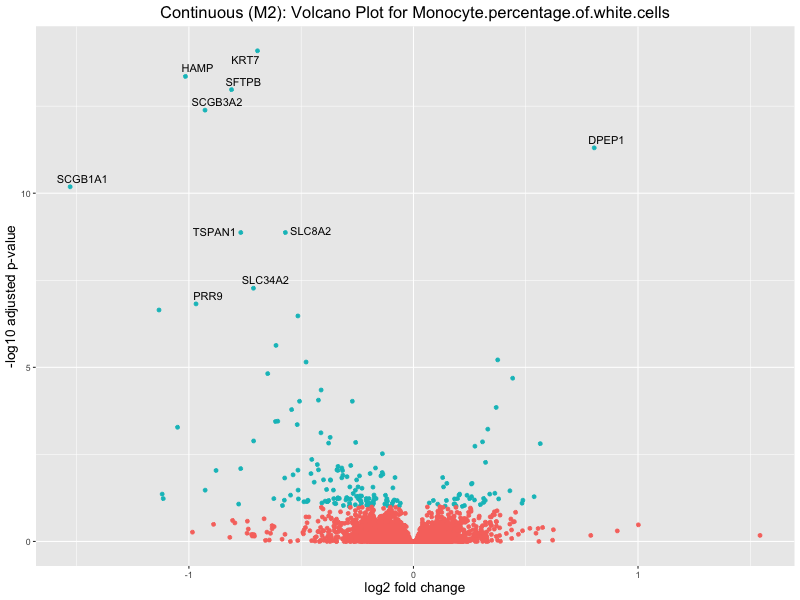
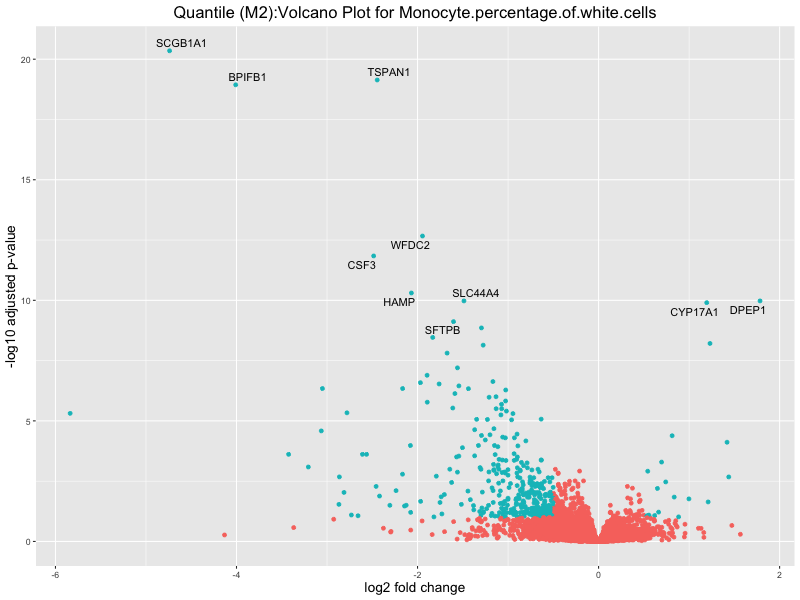
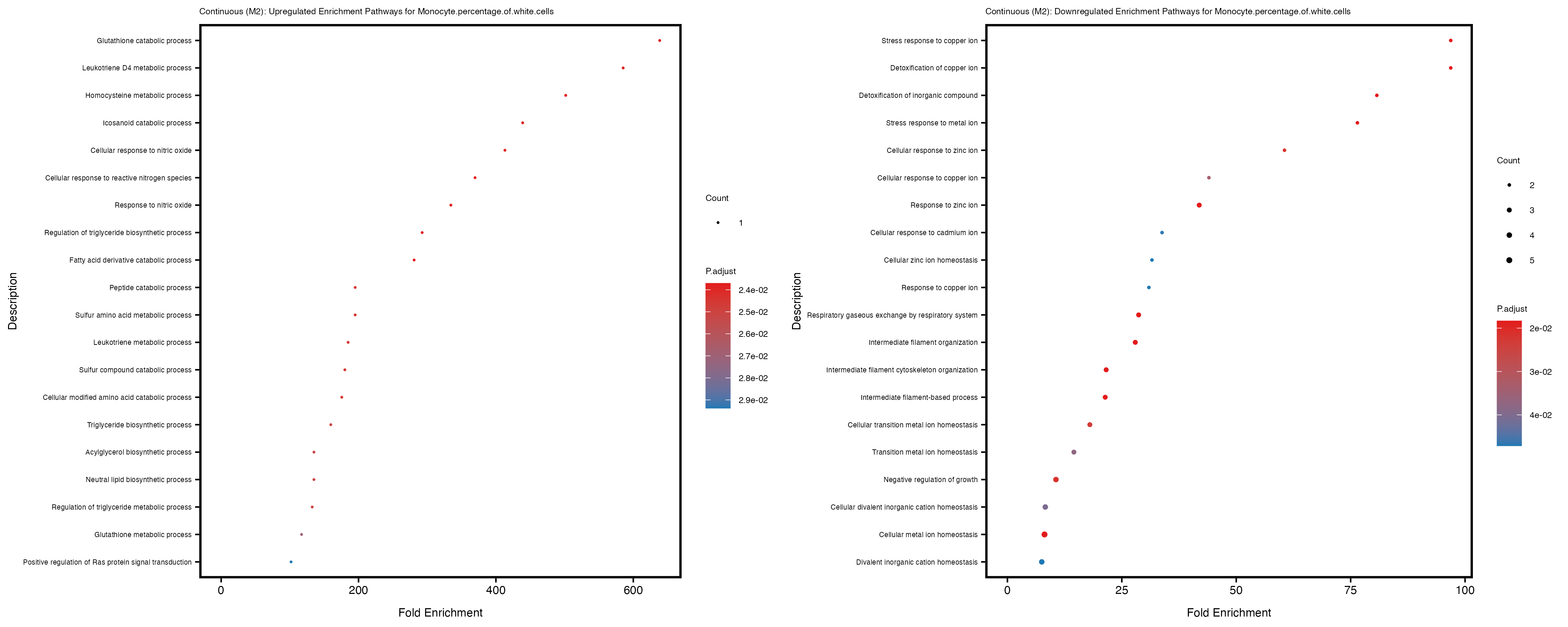
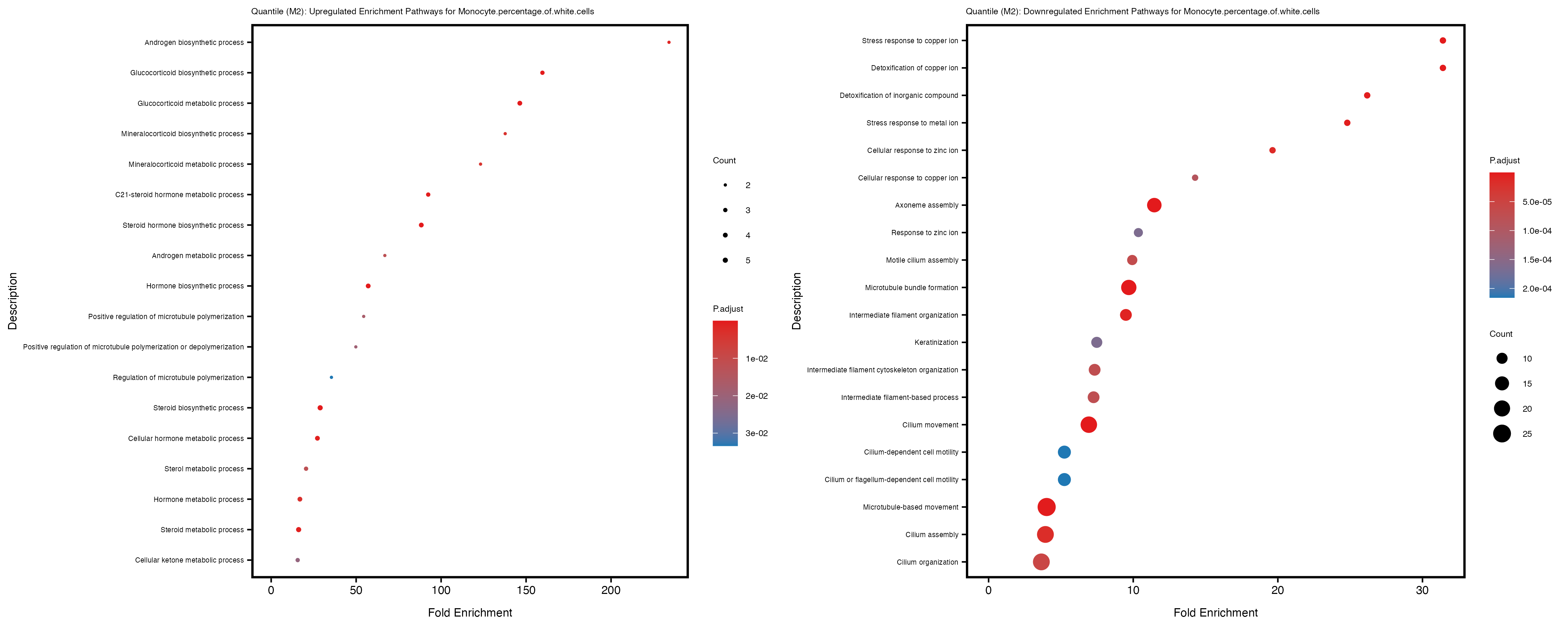
Neutrophil.count
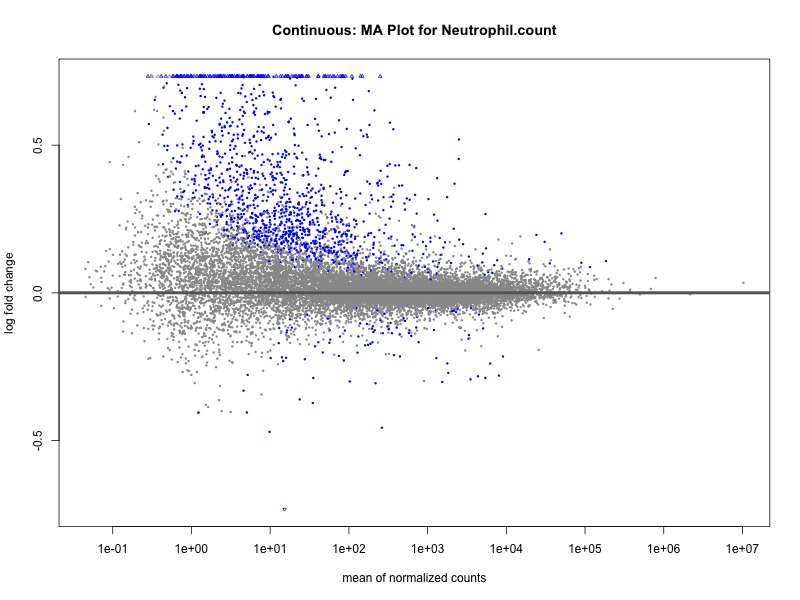
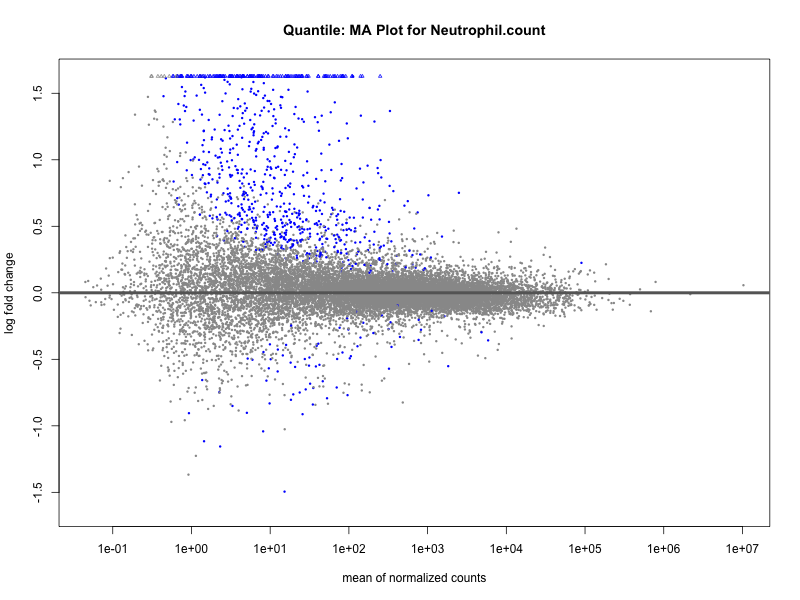
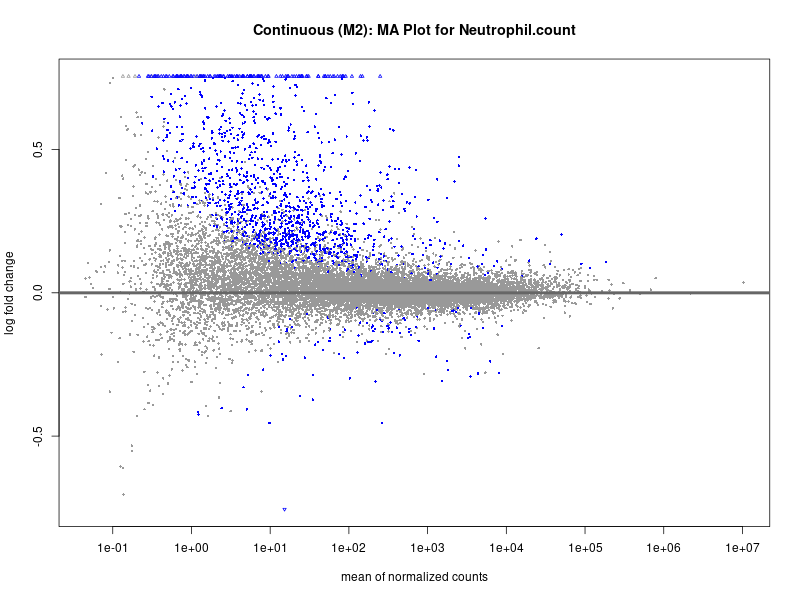
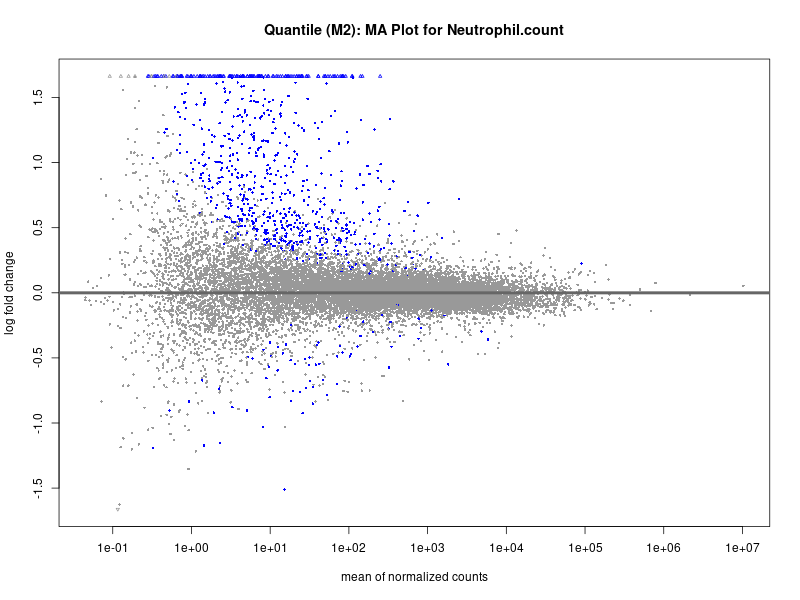
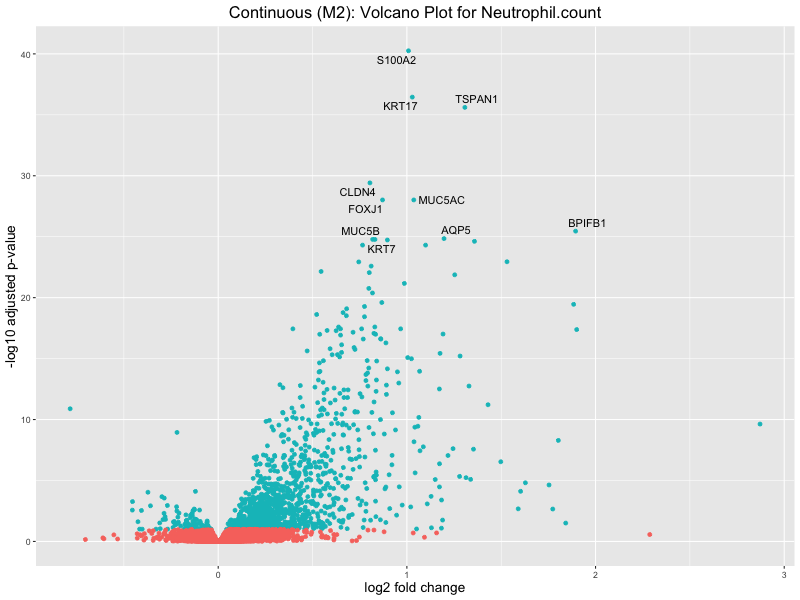
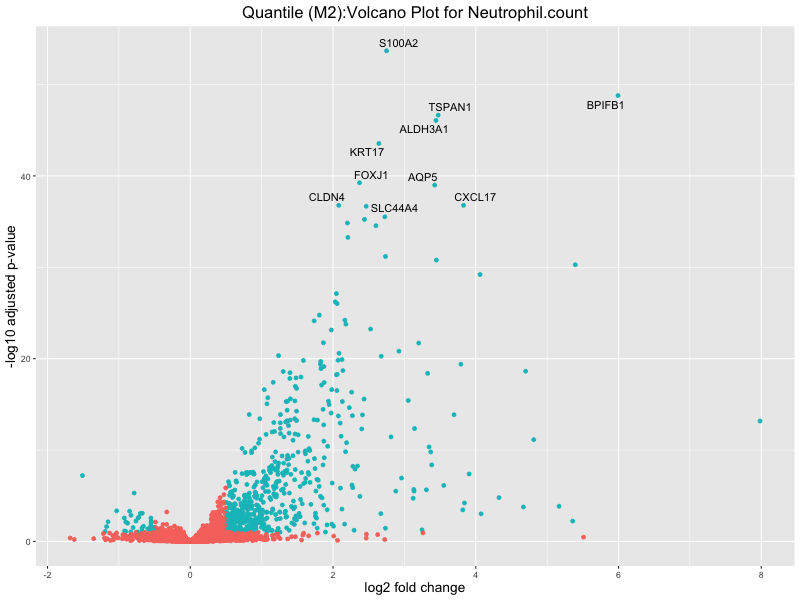
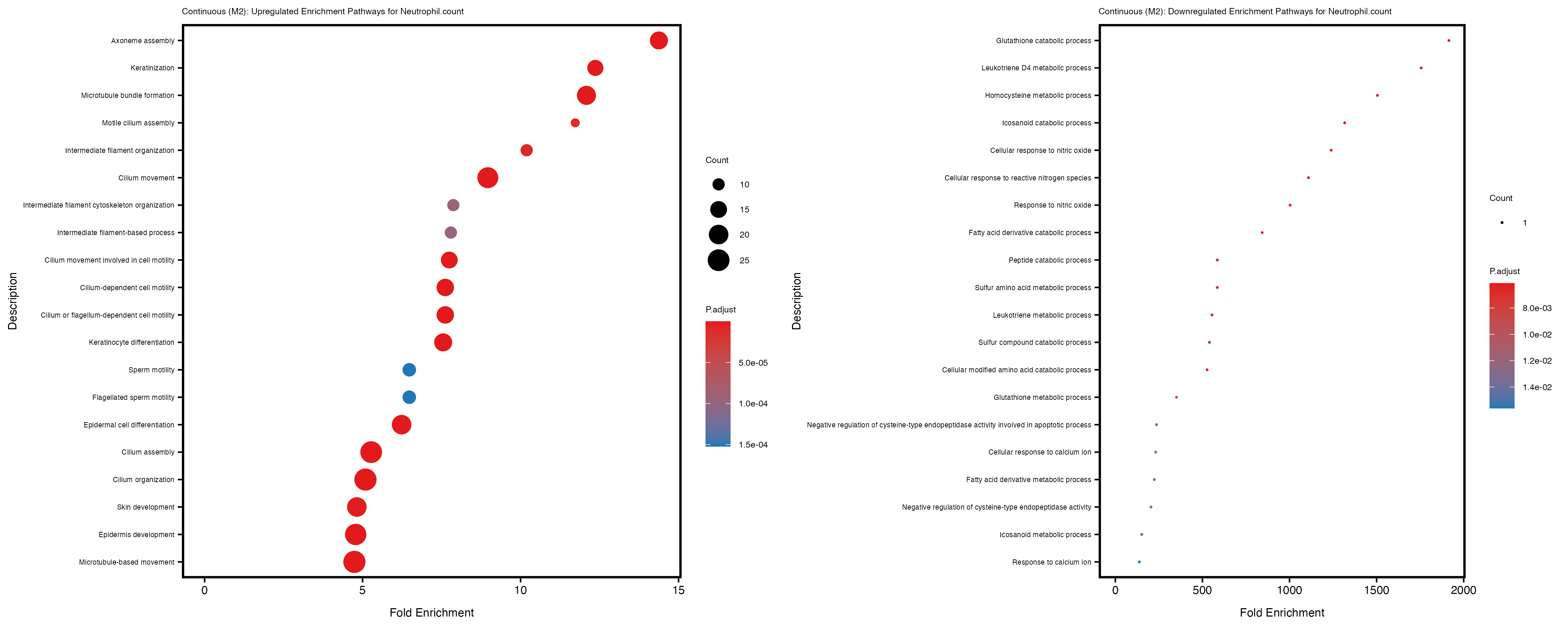
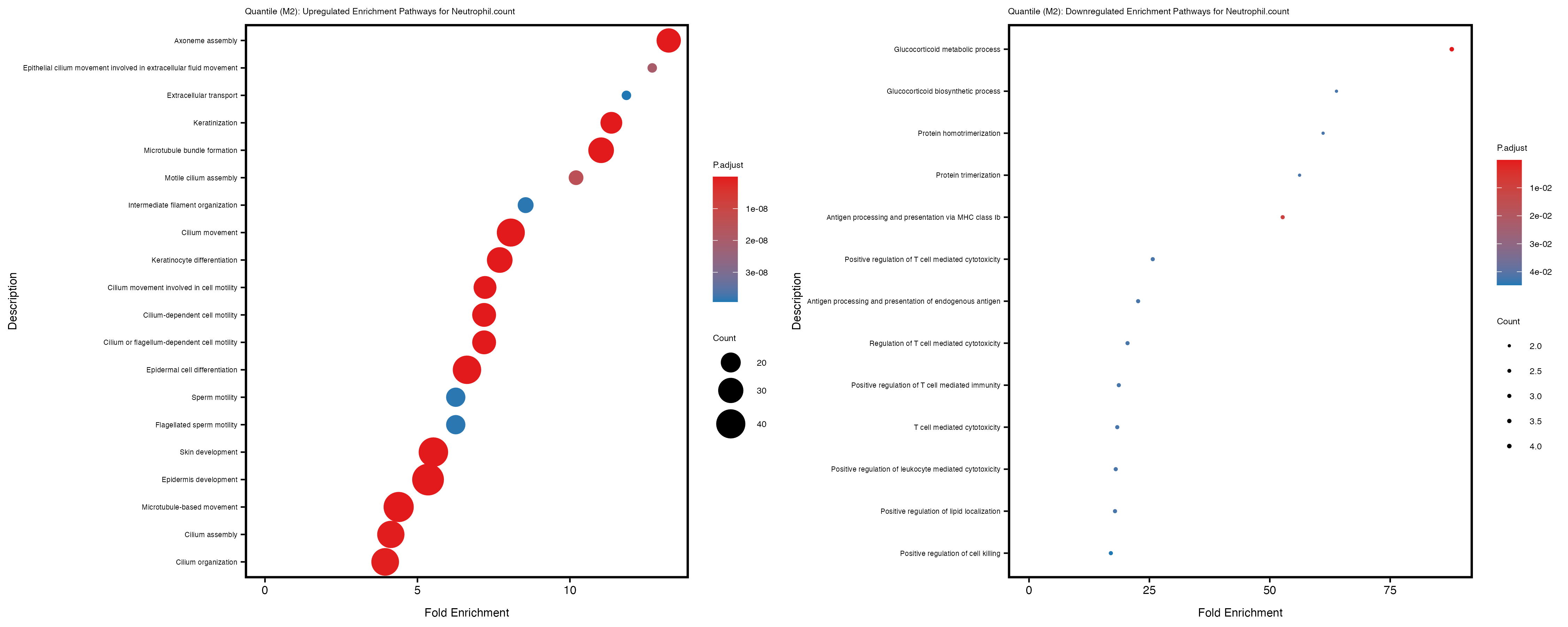
Neutrophil.percentage.of.white.cells
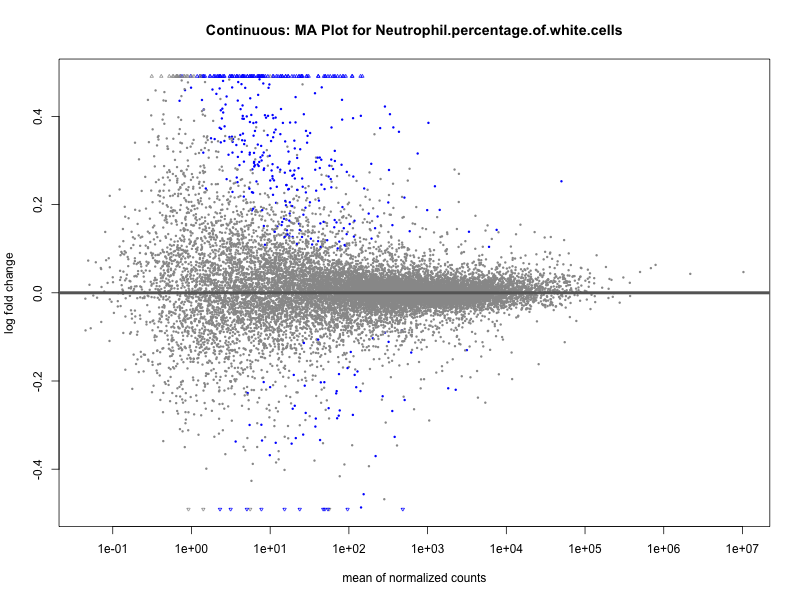
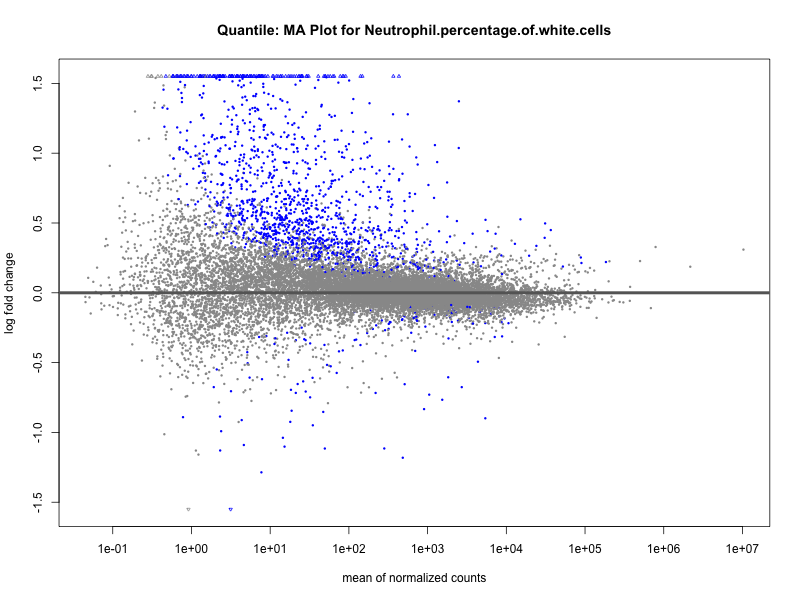
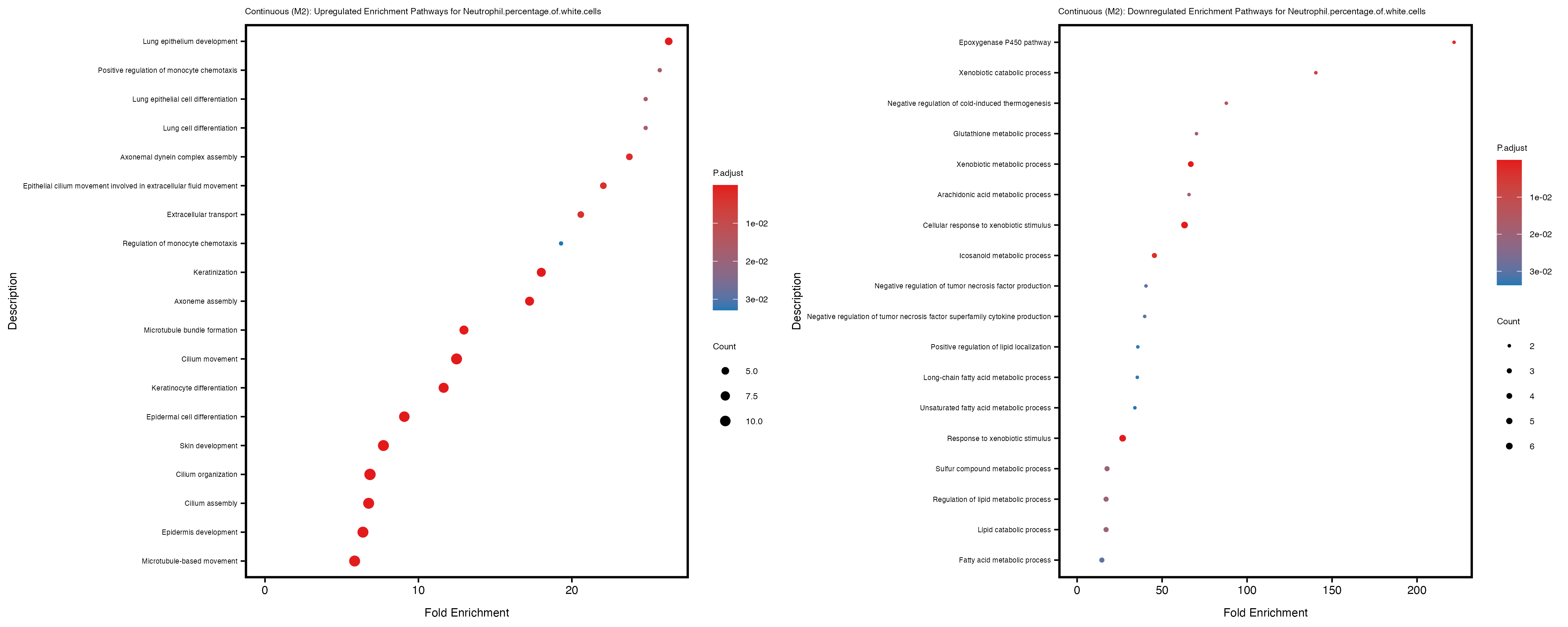
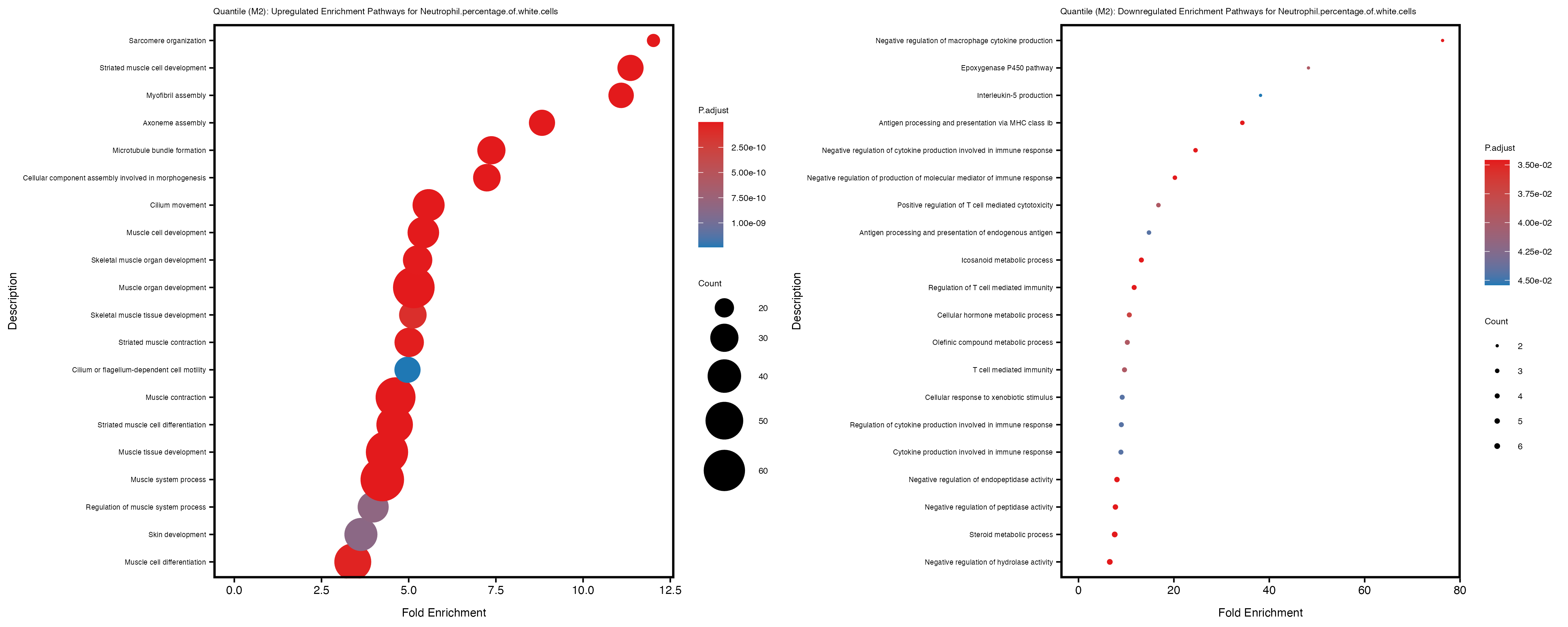
Platelet.count
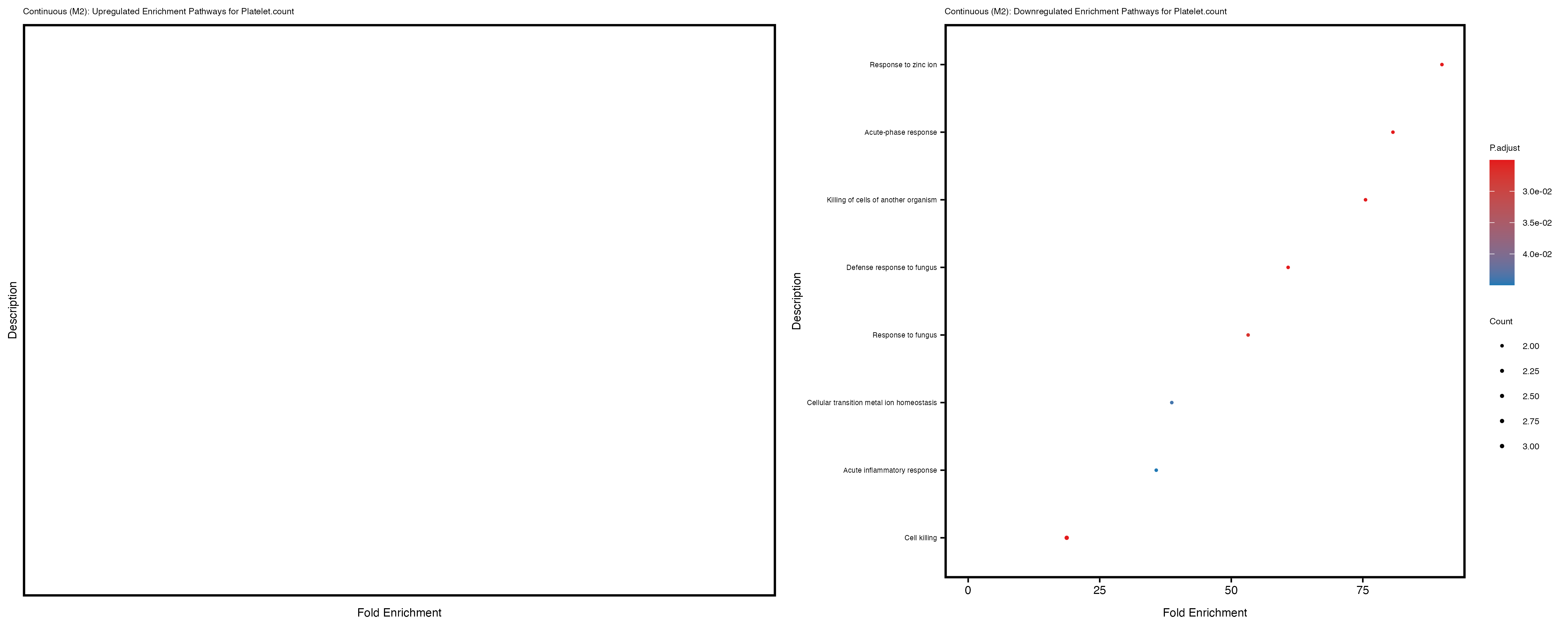
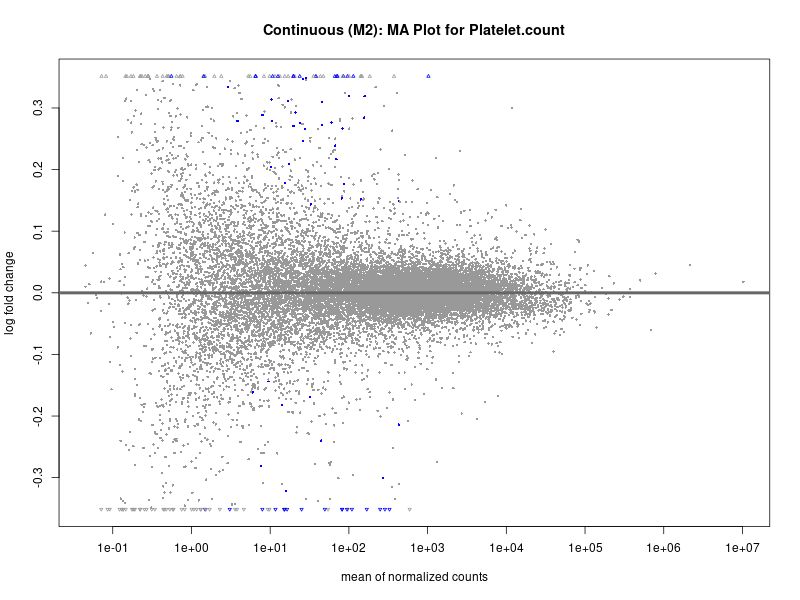
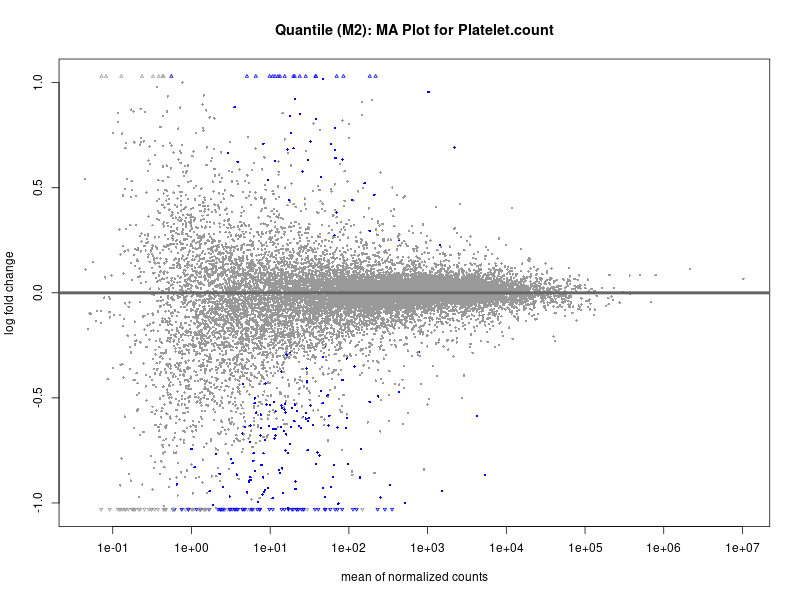
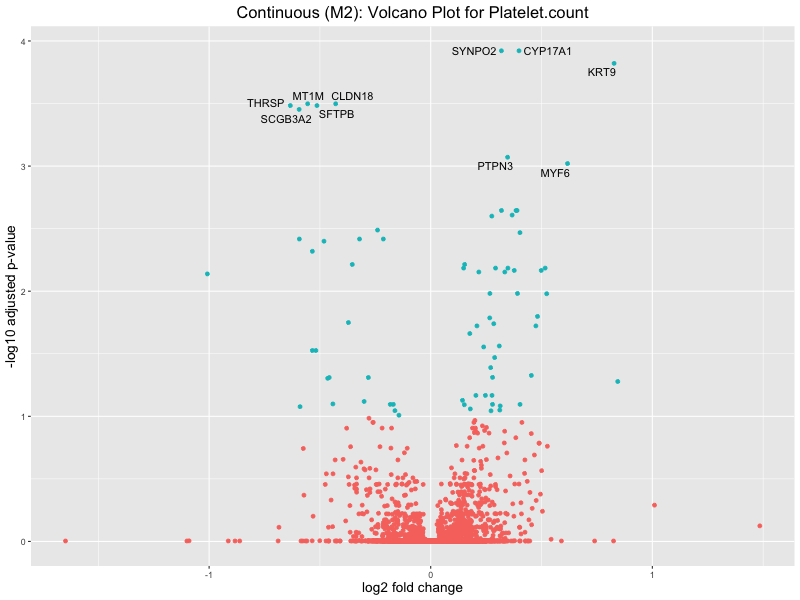
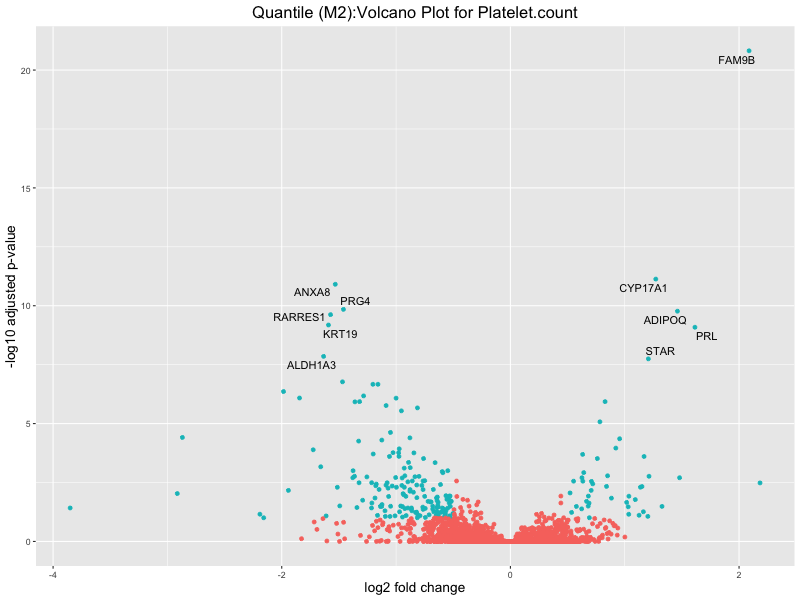

Platelet.crit
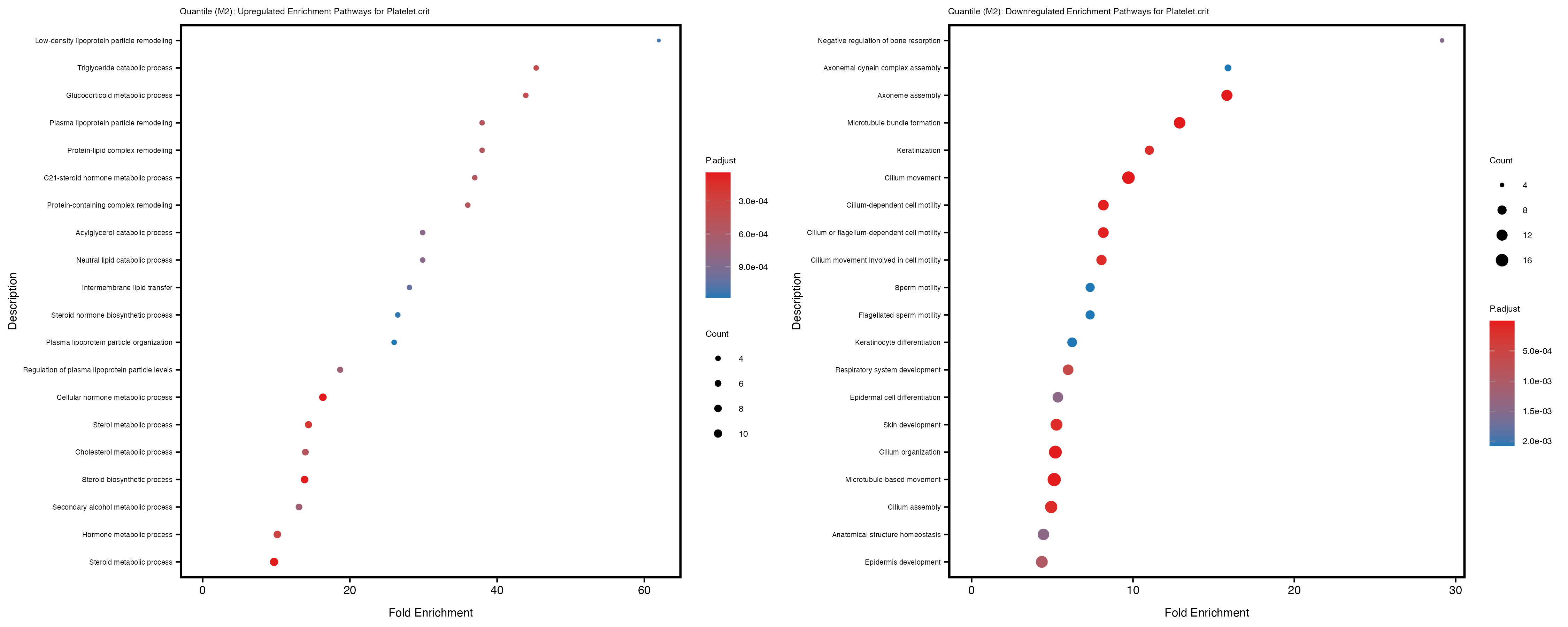
Platelet.distribution.width
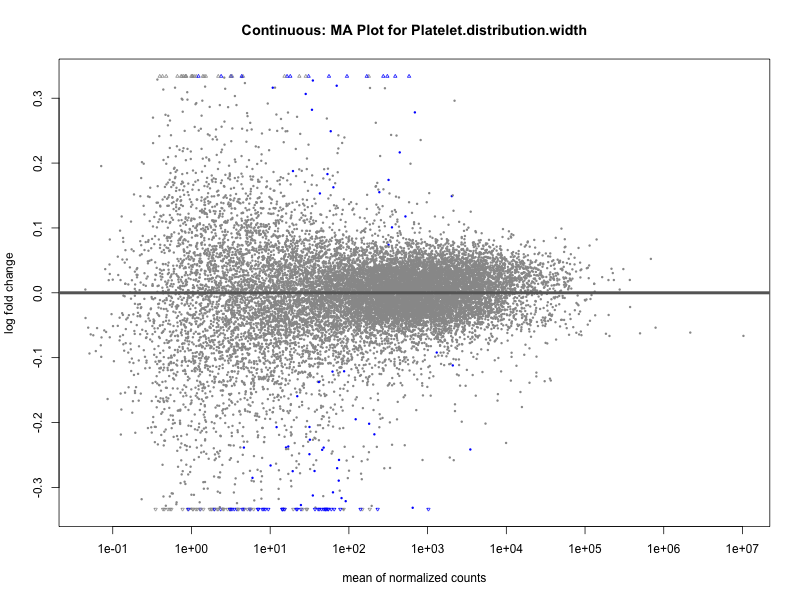
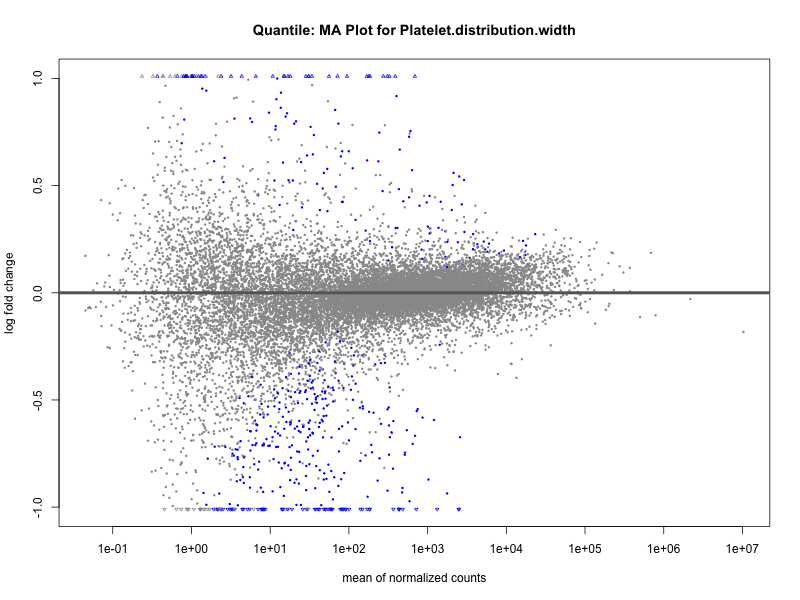
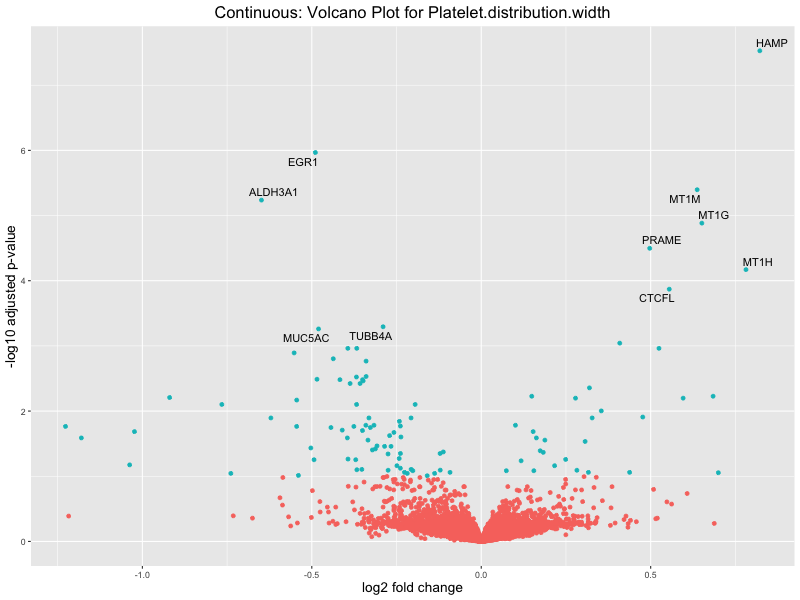
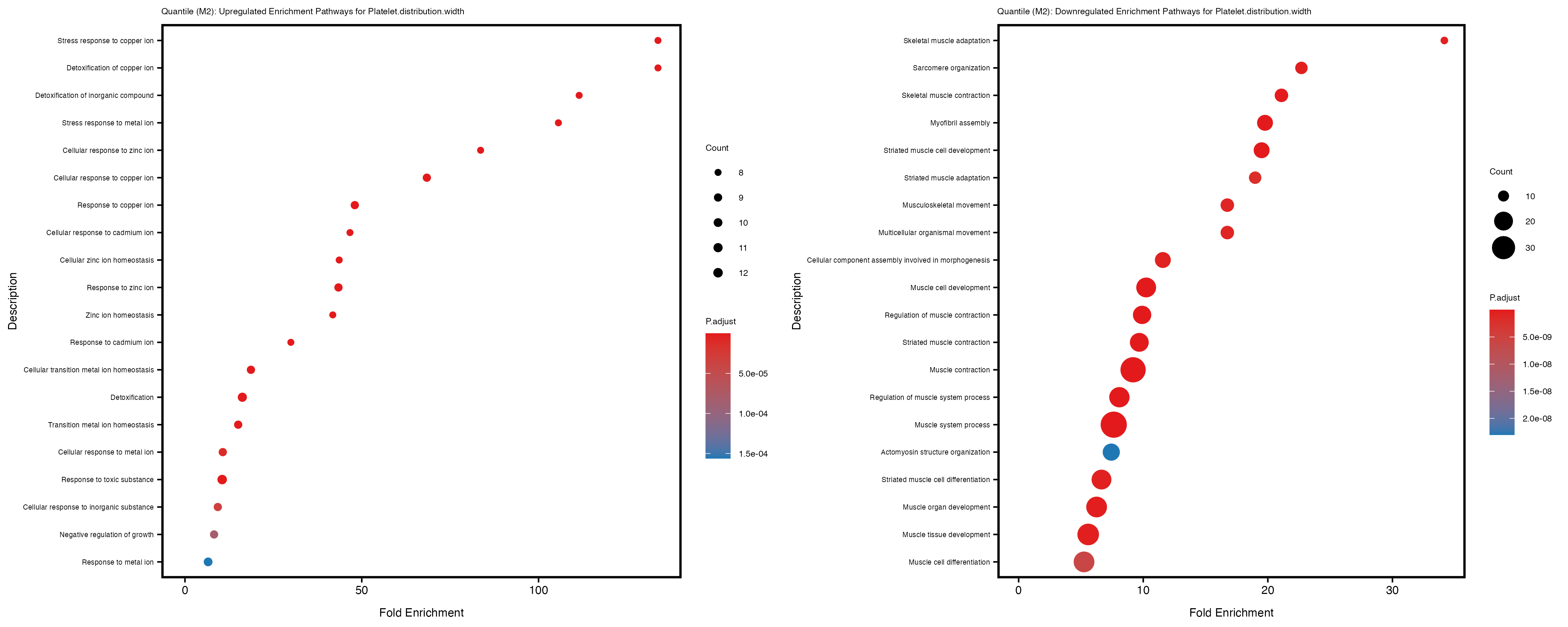
Red.blood.cell.count
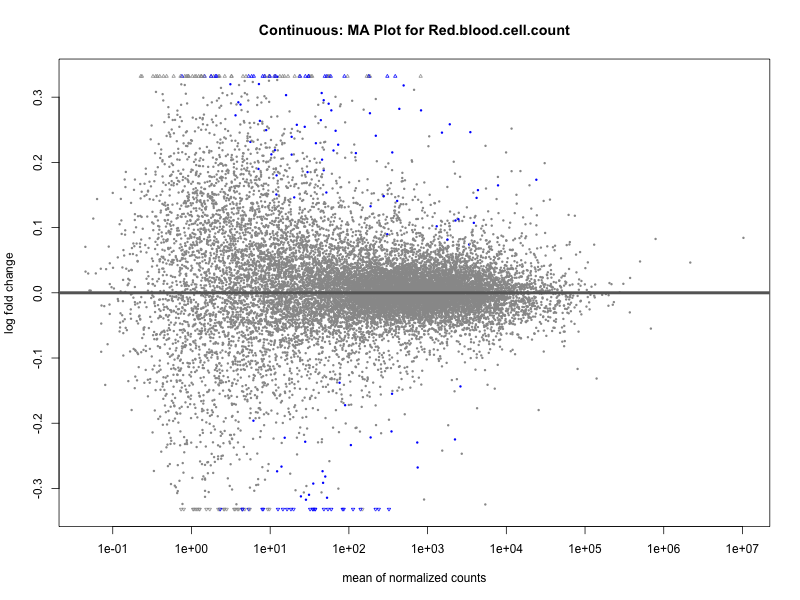
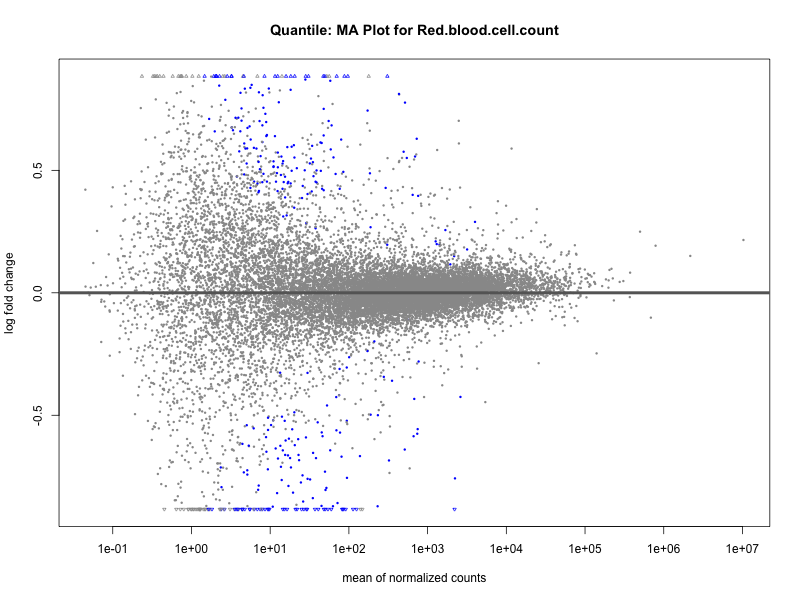
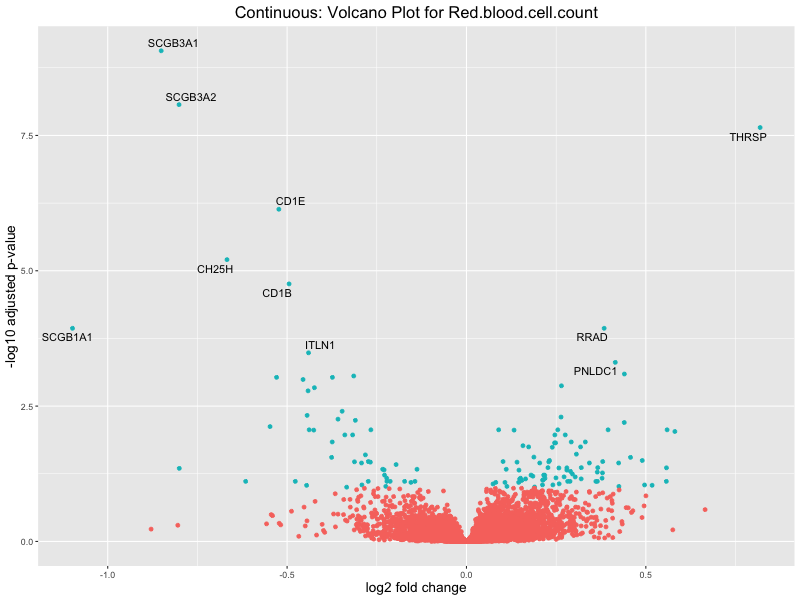
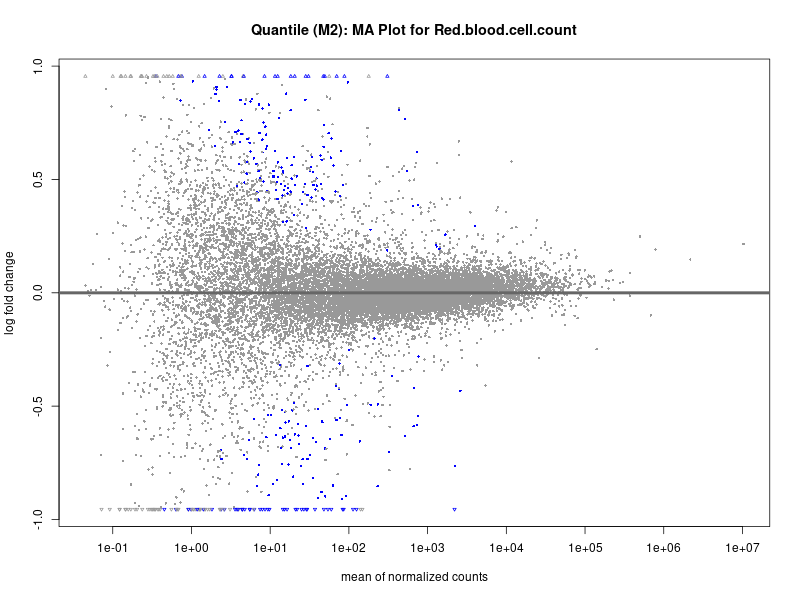
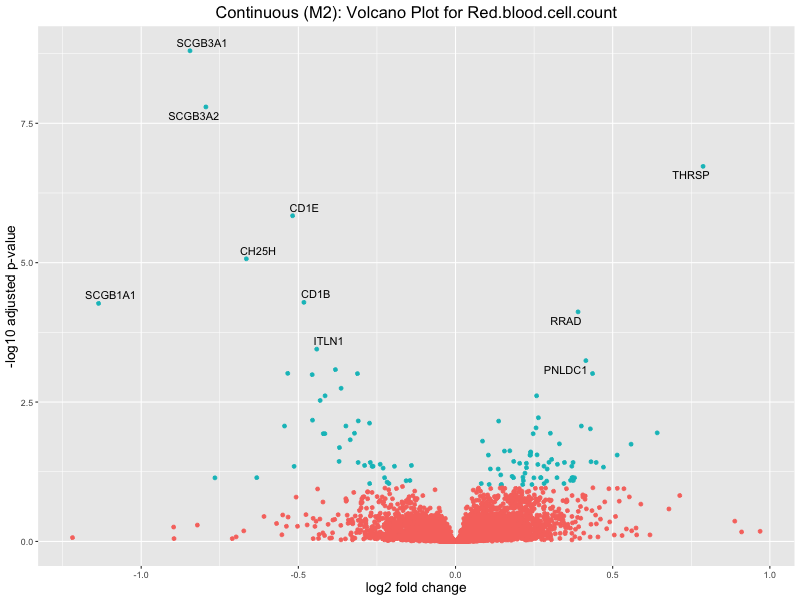
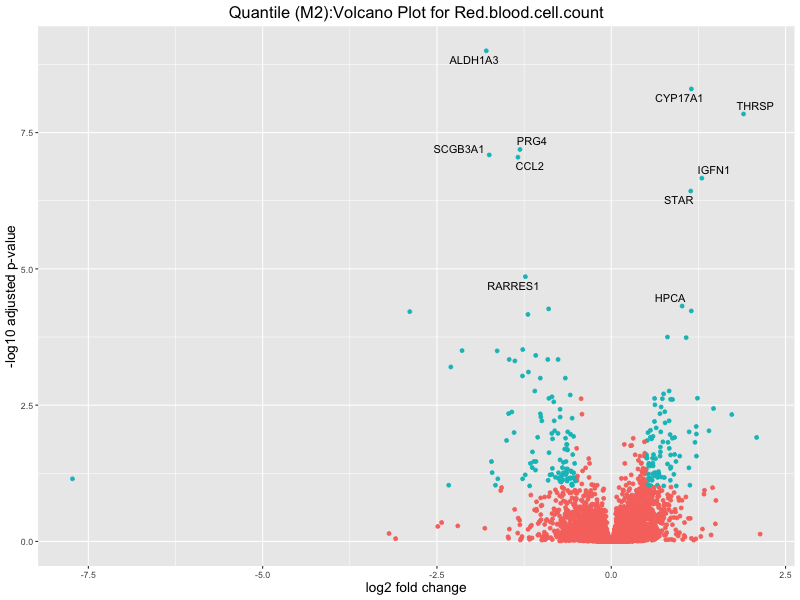
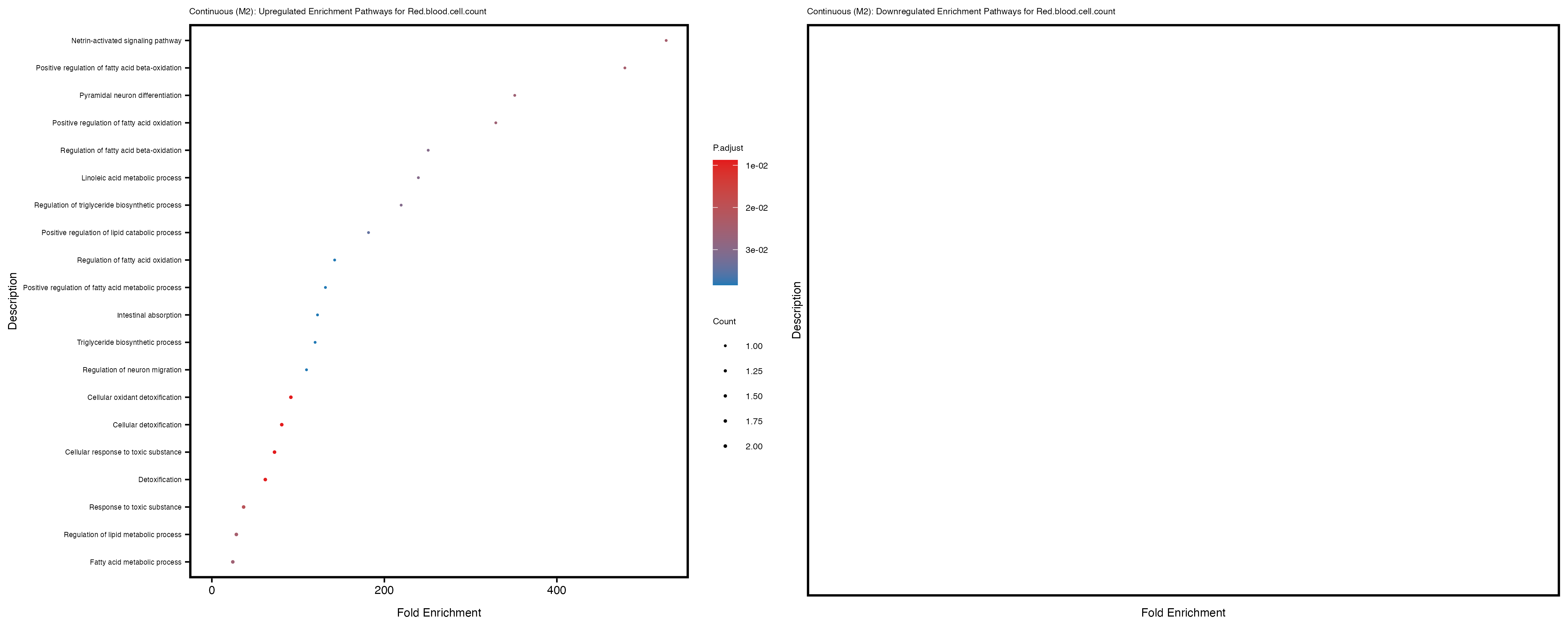
Red.cell.distribution.width
Reticulocyte.count
Reticulocyte.fraction.of.red.cells
White.blood.cell.count
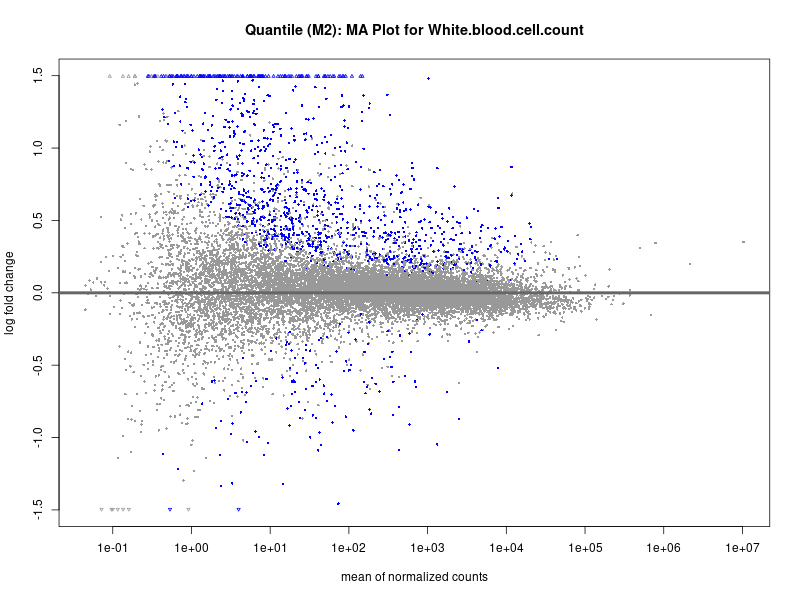
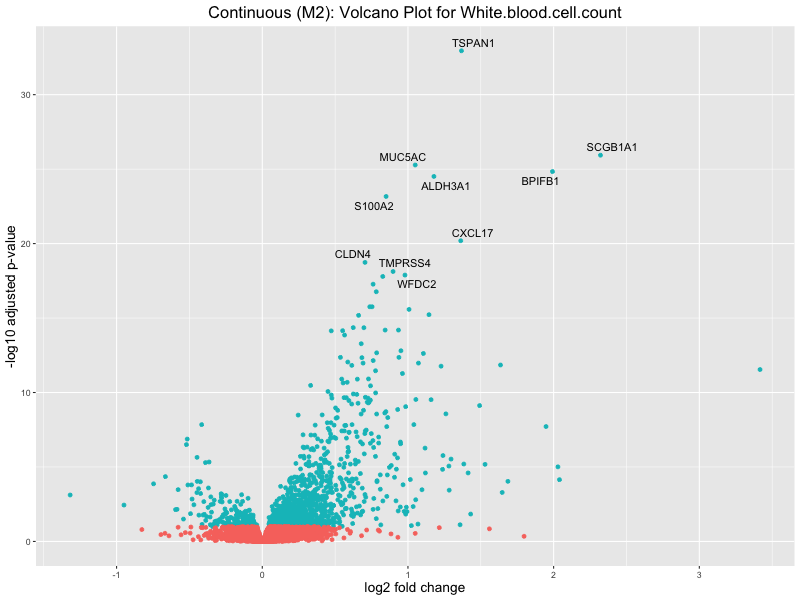
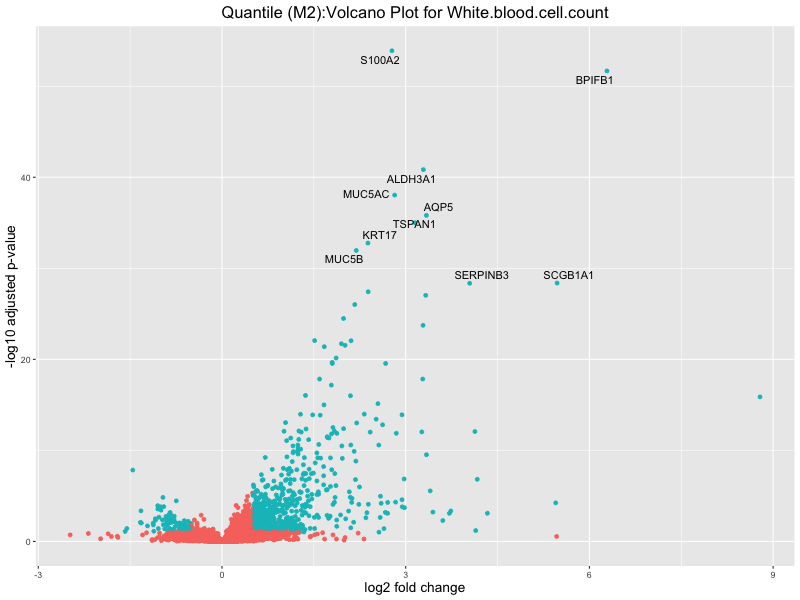
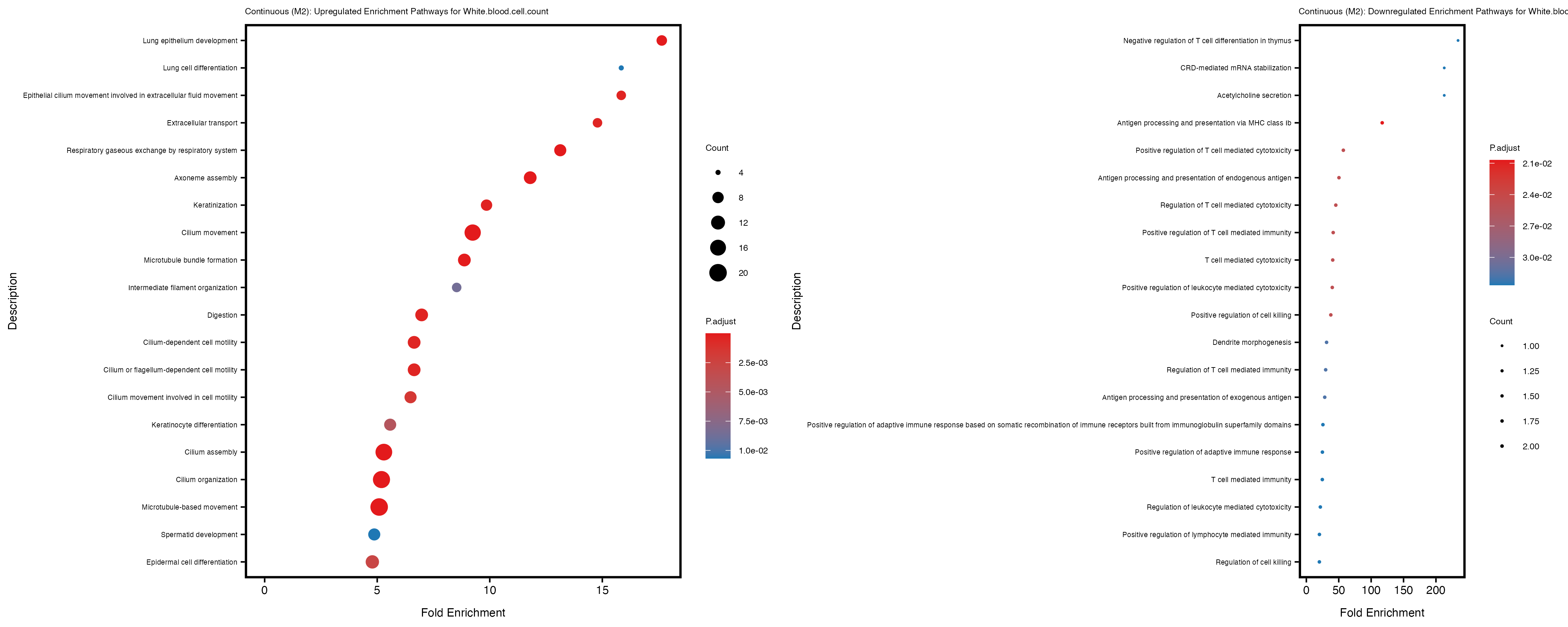
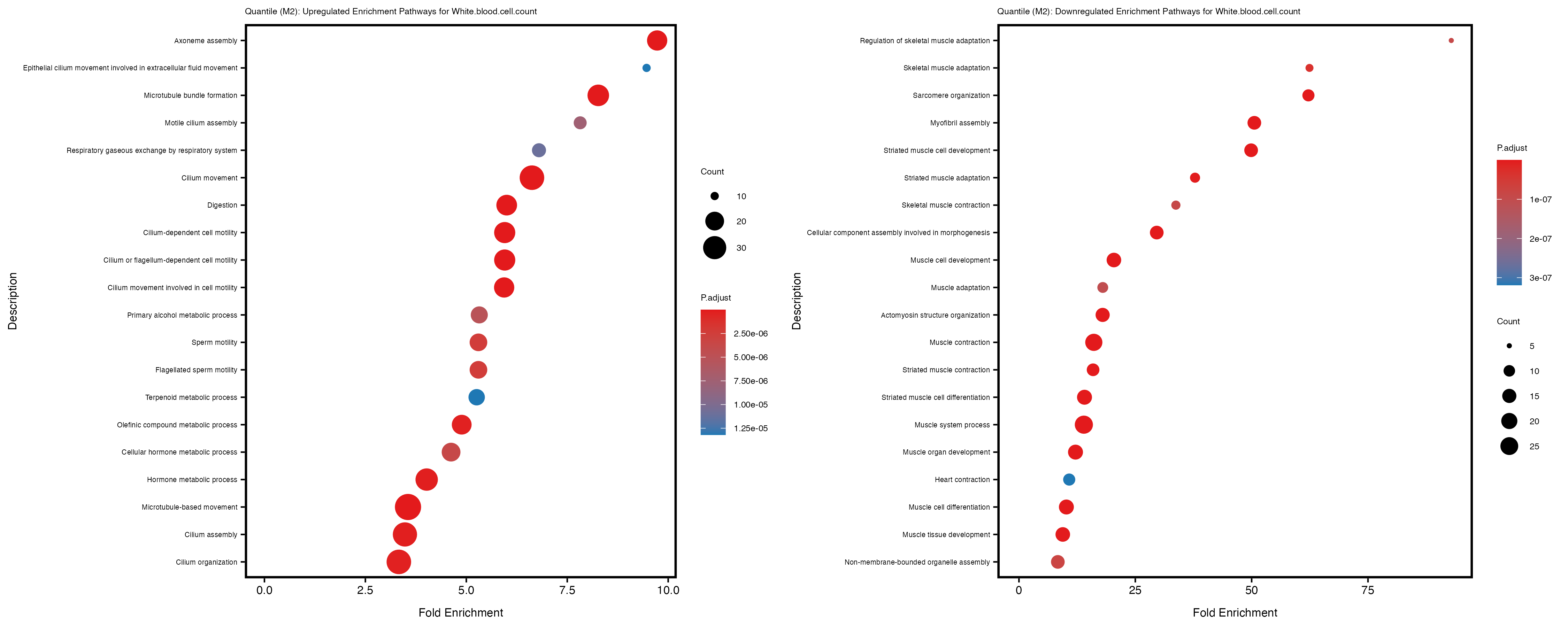
Celiac_GCST90014442
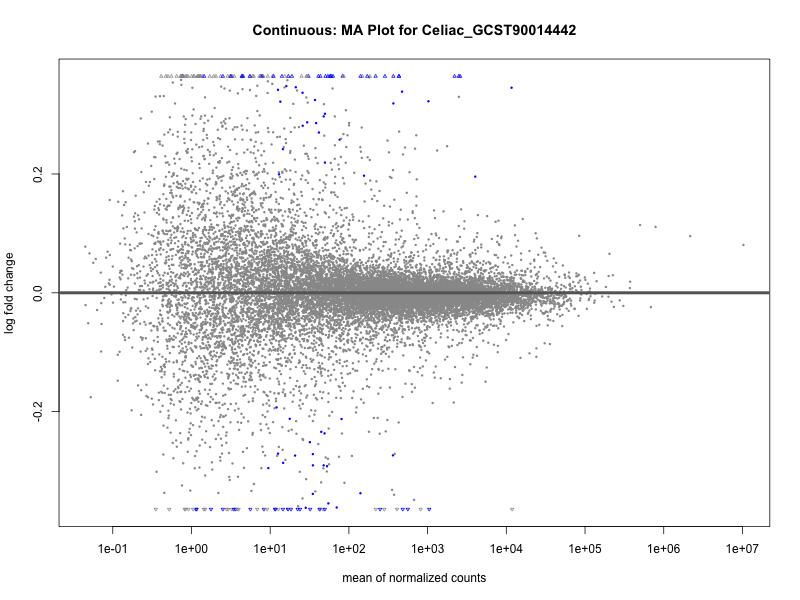
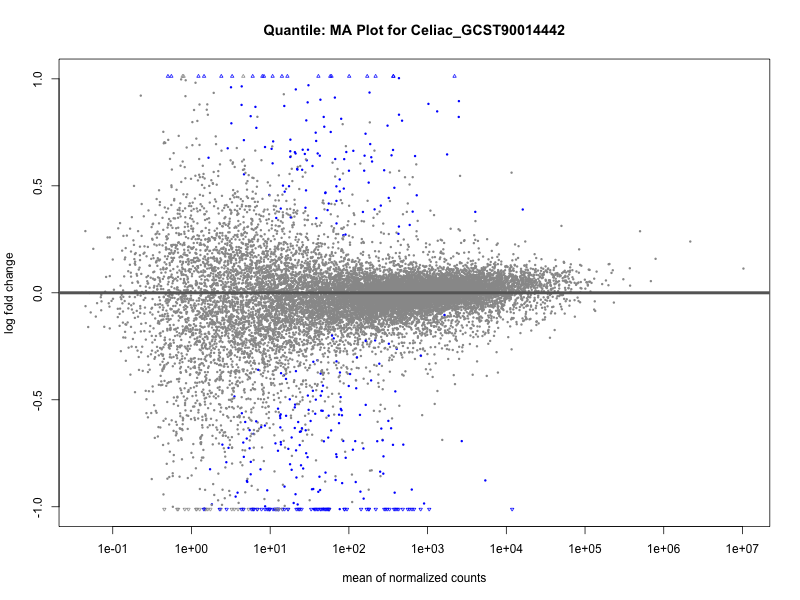
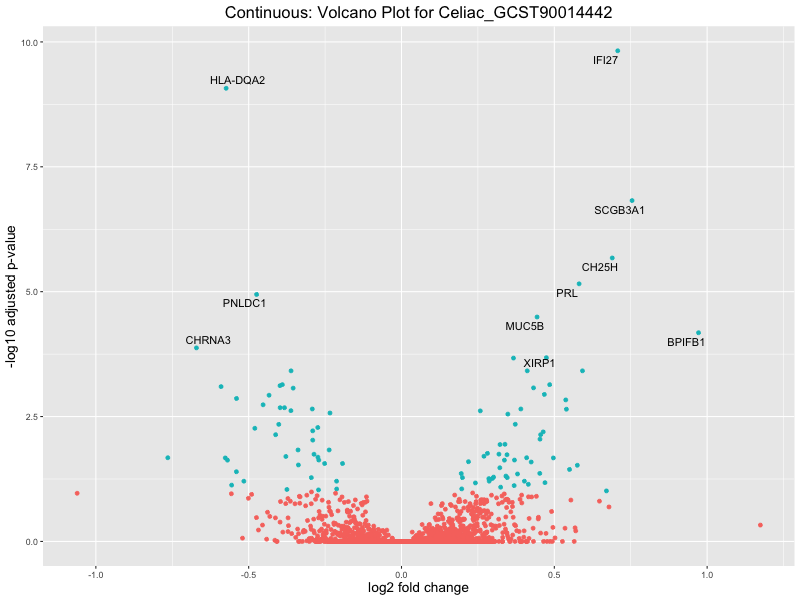
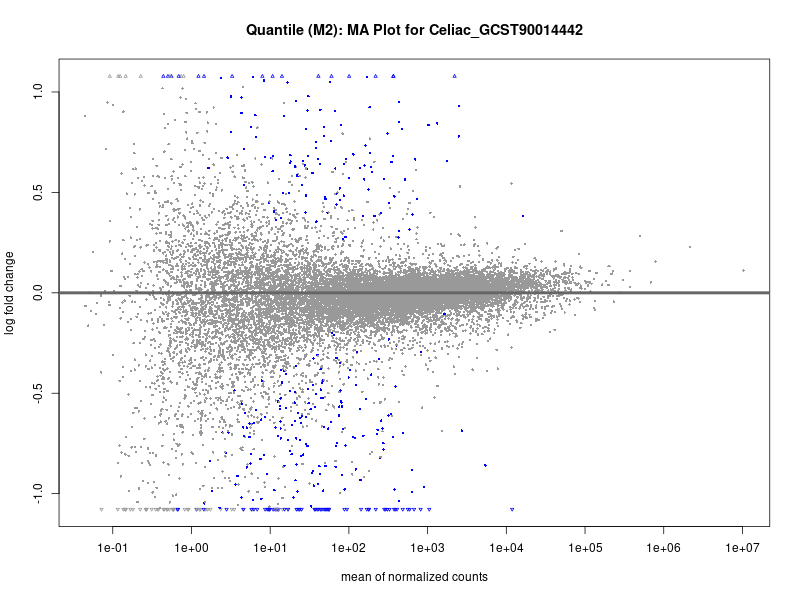
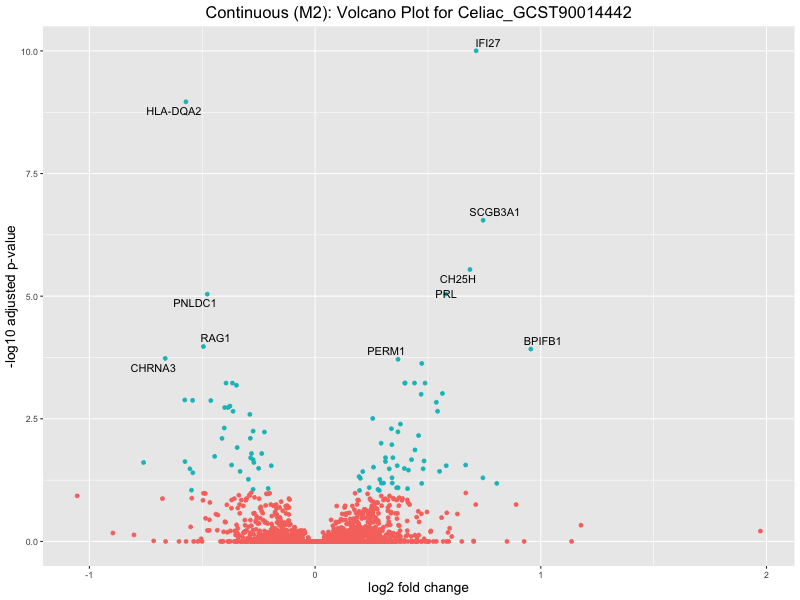
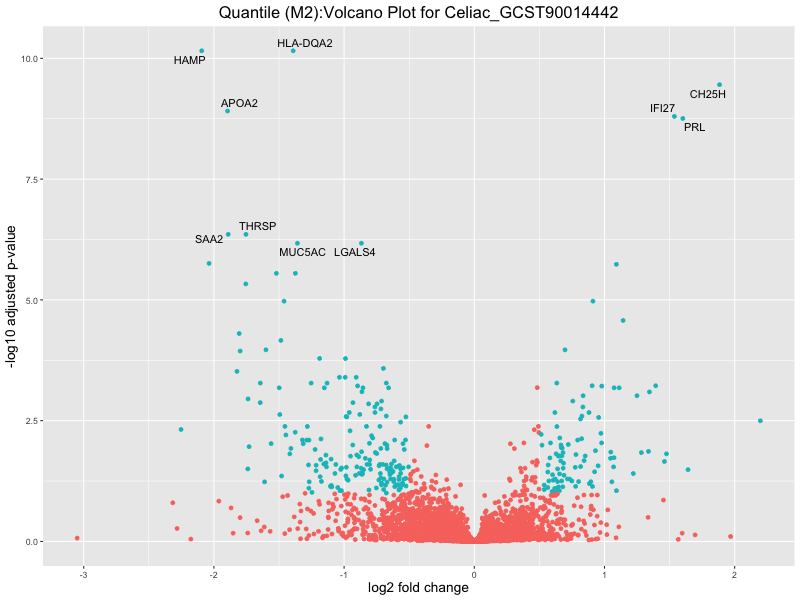
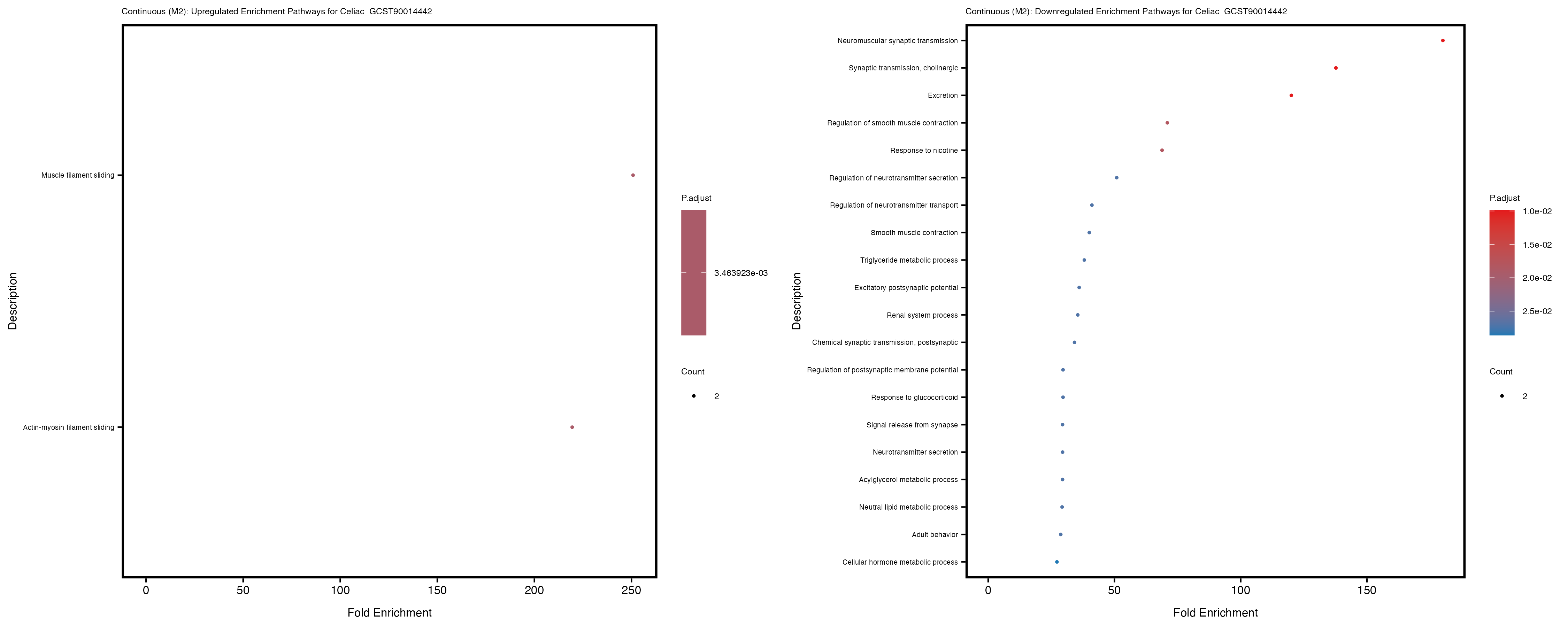
Celiac_GCST90468120
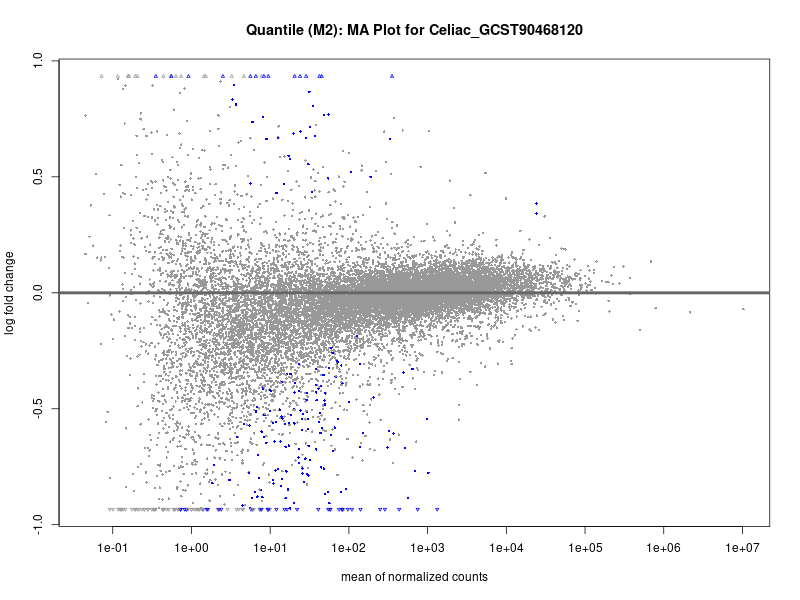
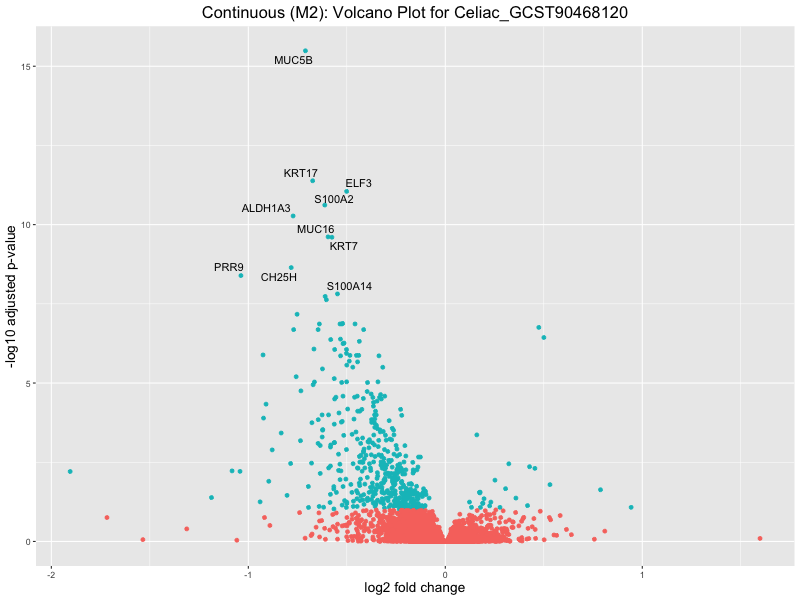
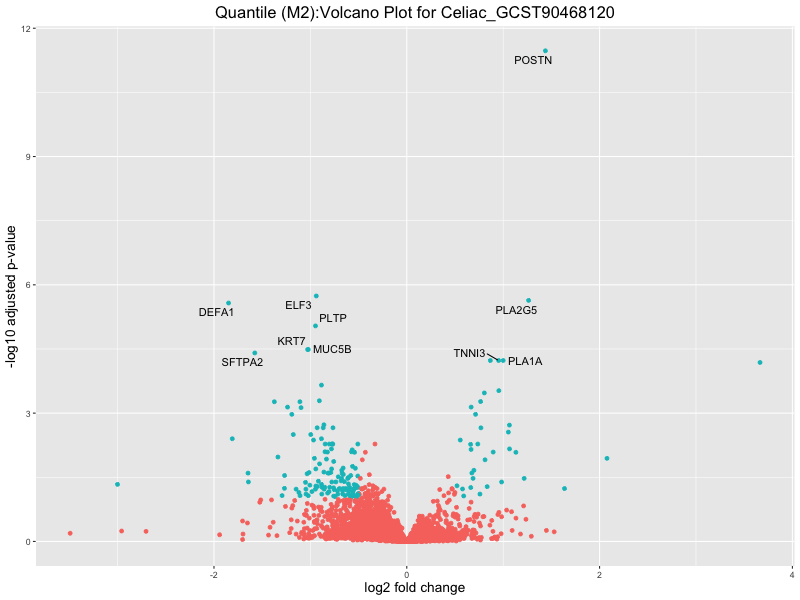
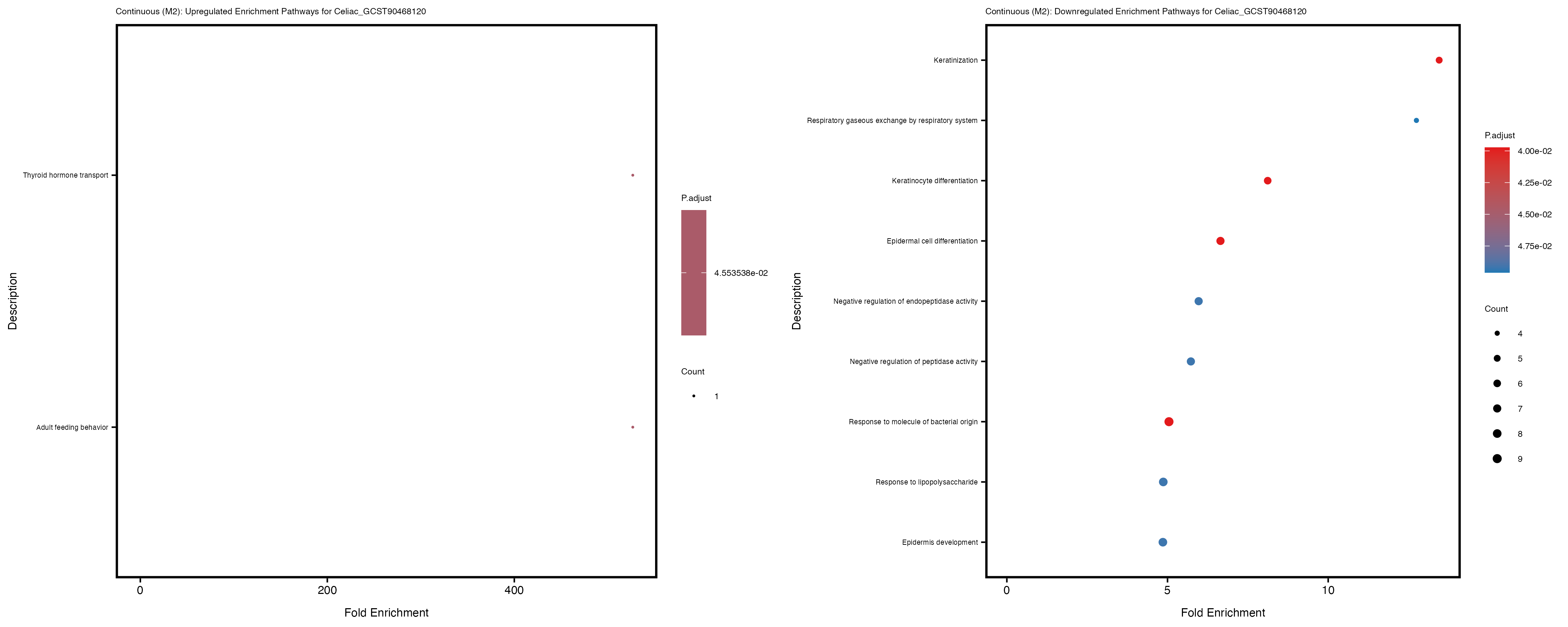
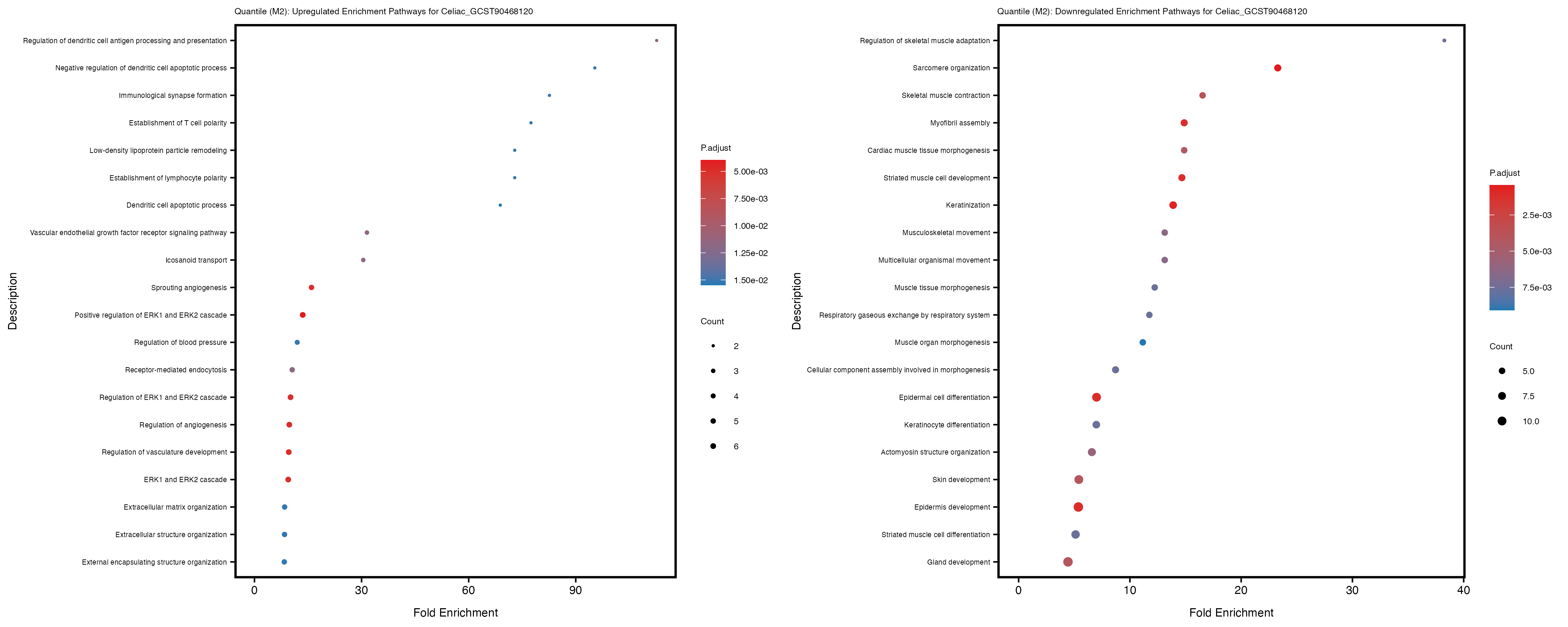
IBD_GCST90013901
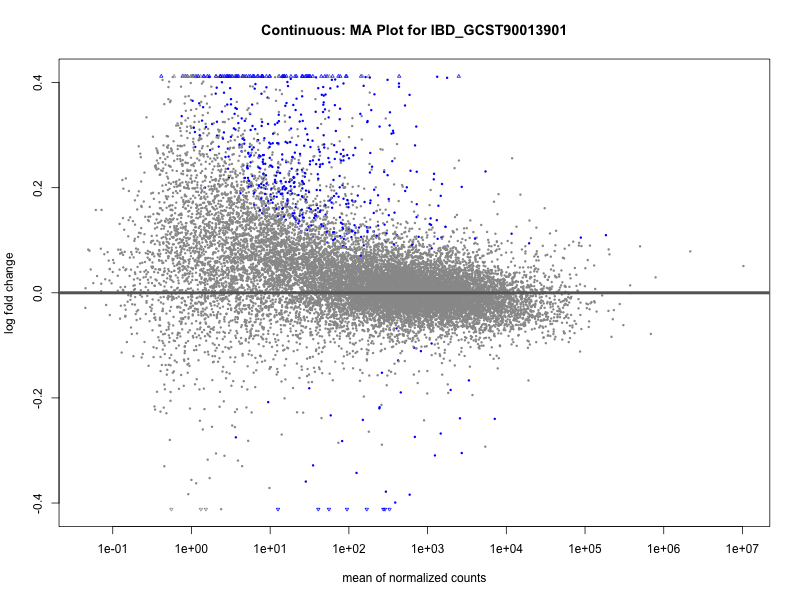
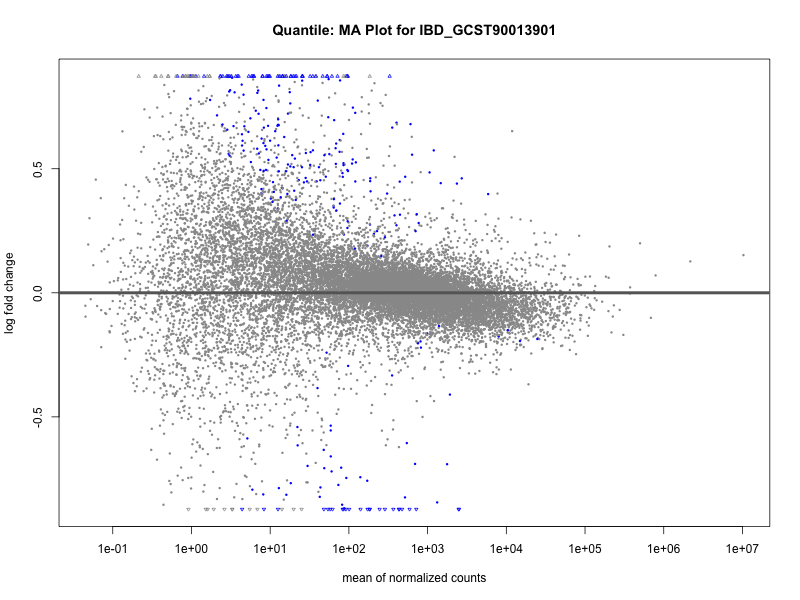
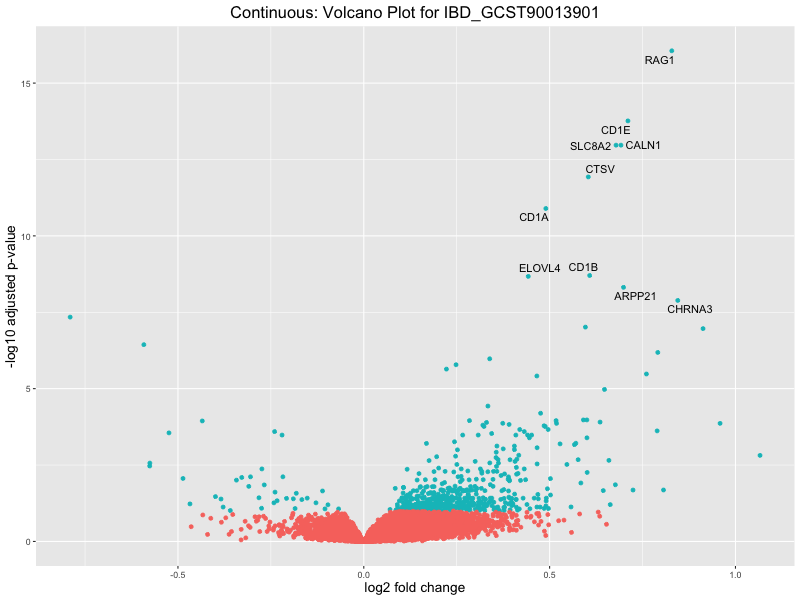
IBD_GCST90013951
LUPUS_GCST003156
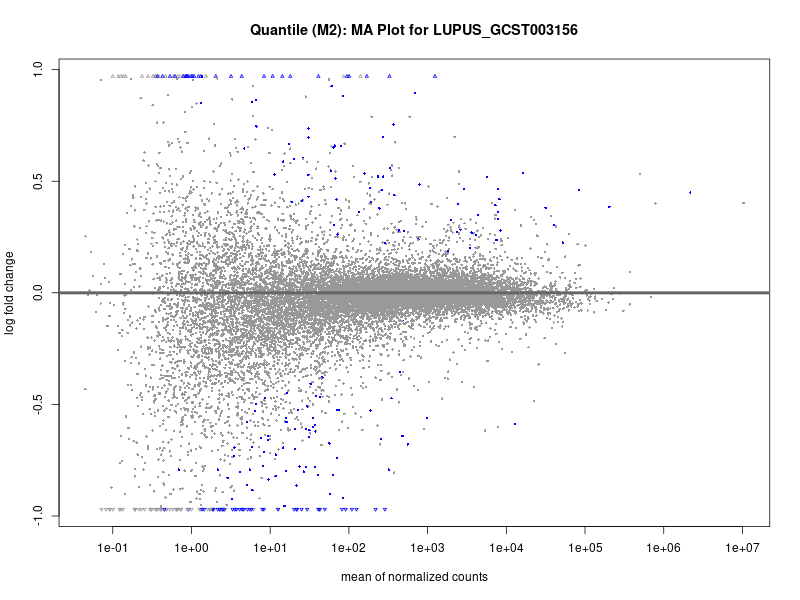
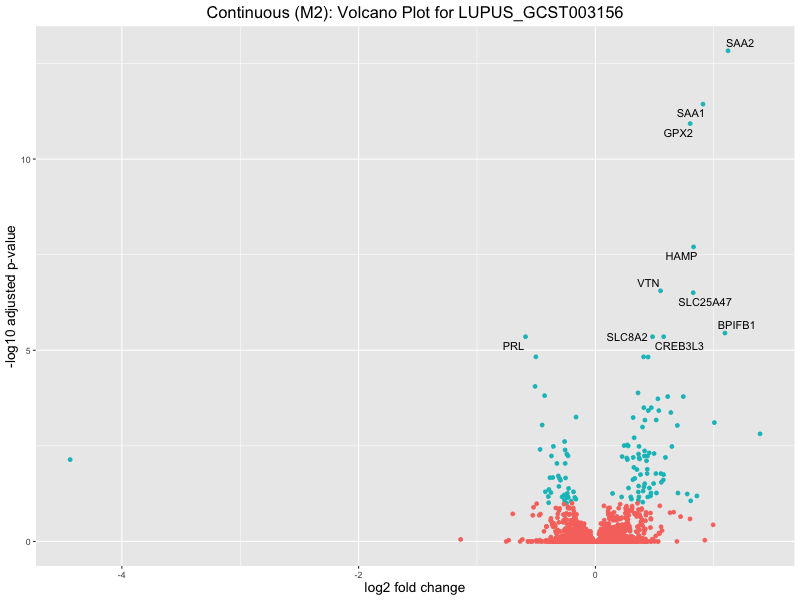
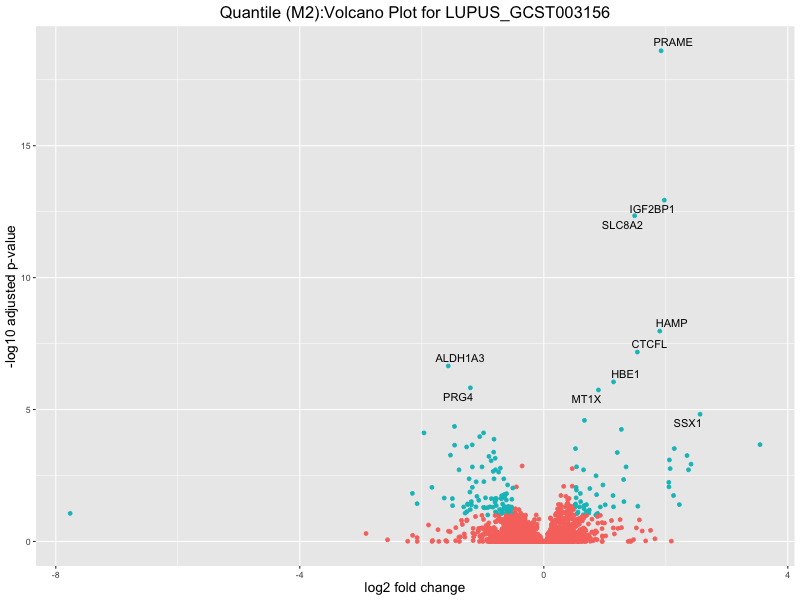
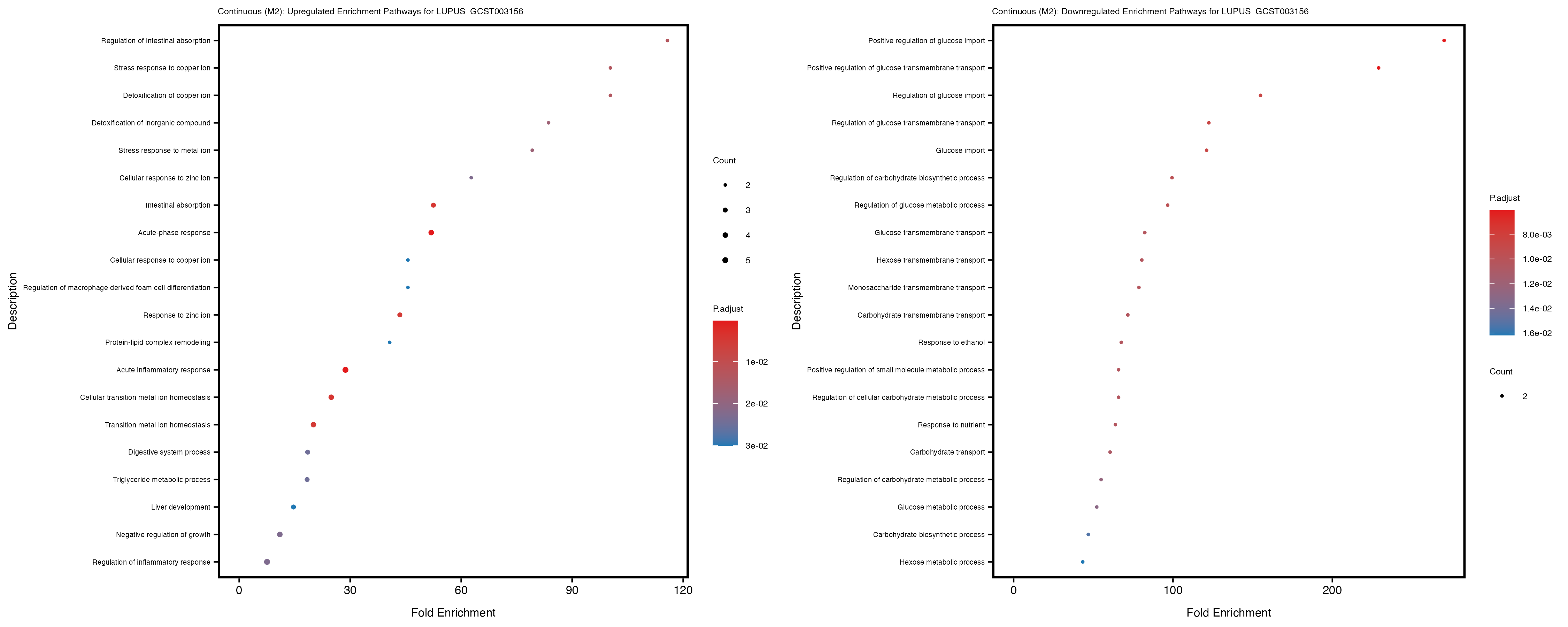
LUPUS_GCST011096
T1D_GCST90000529
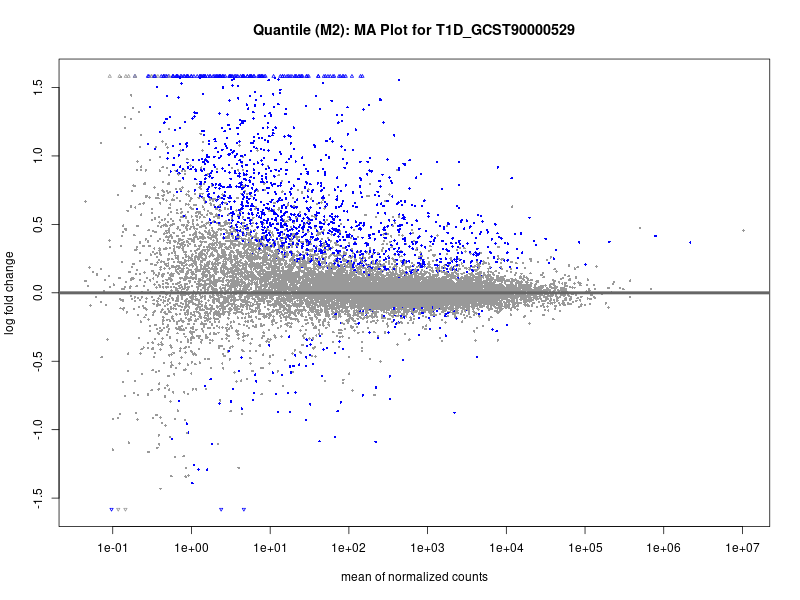
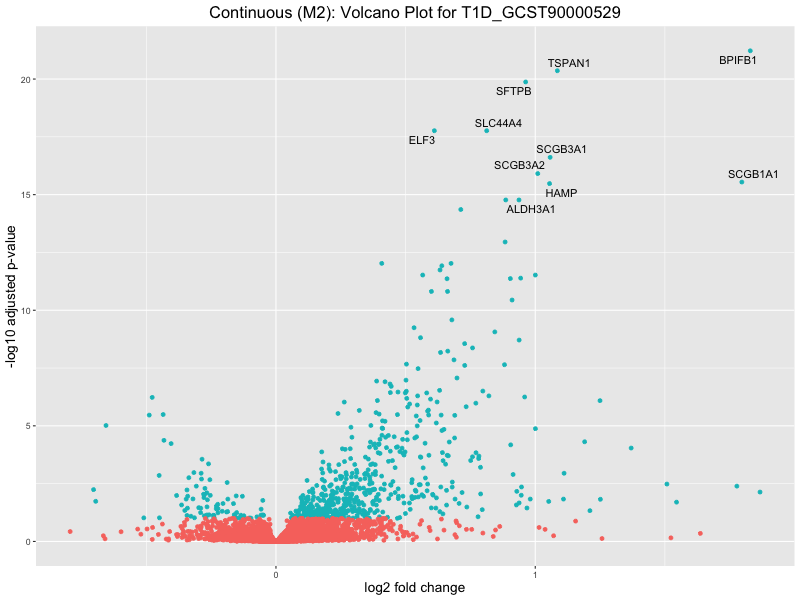
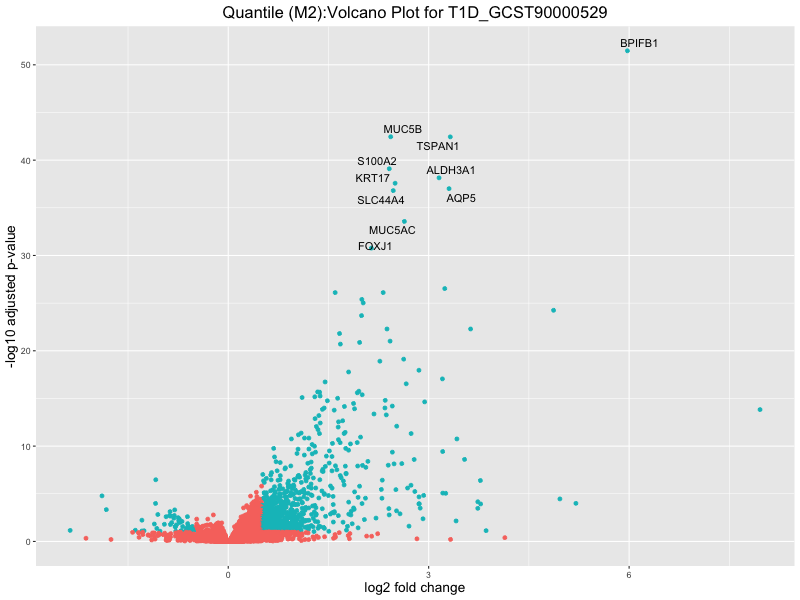
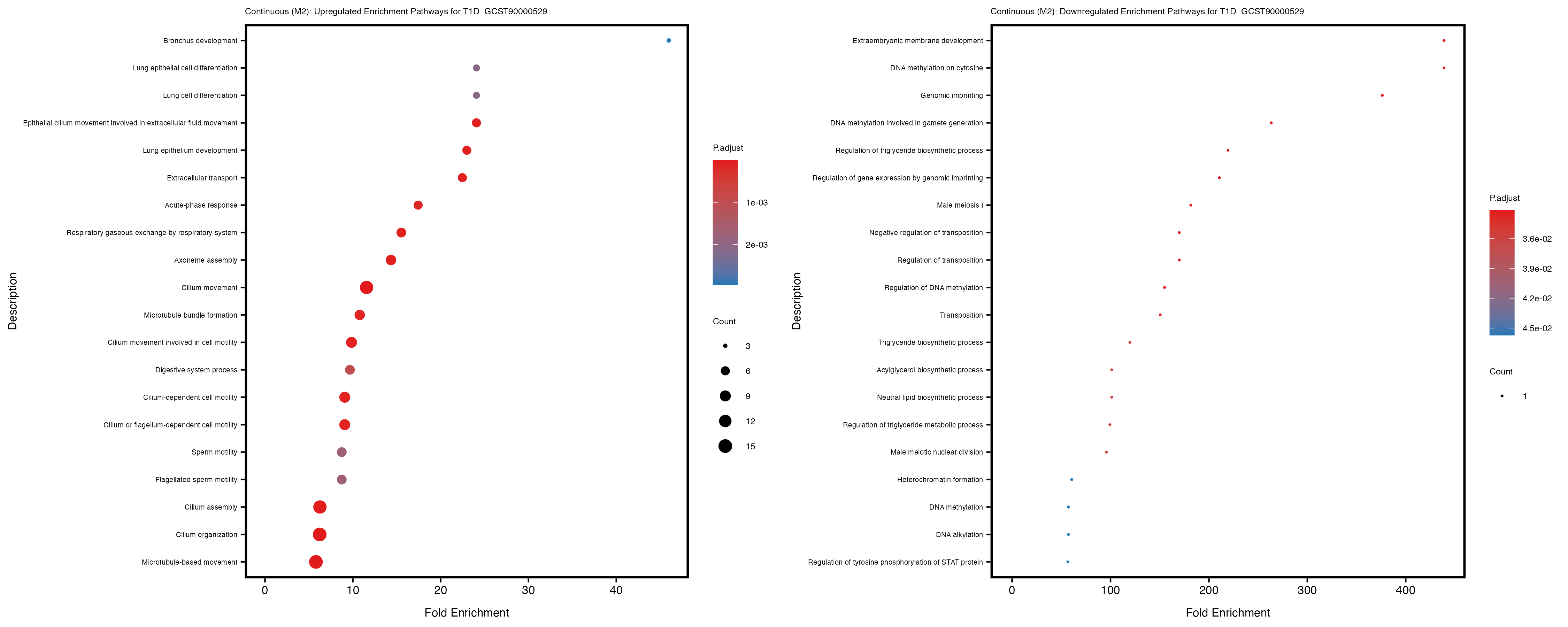
T1D_GCST90014023
sessionInfo()R version 4.2.2 (2022-10-31)
Platform: x86_64-apple-darwin17.0 (64-bit)
Running under: macOS Big Sur ... 10.16
Matrix products: default
BLAS: /Library/Frameworks/R.framework/Versions/4.2/Resources/lib/libRblas.0.dylib
LAPACK: /Library/Frameworks/R.framework/Versions/4.2/Resources/lib/libRlapack.dylib
locale:
[1] en_US.UTF-8/en_US.UTF-8/en_US.UTF-8/C/en_US.UTF-8/en_US.UTF-8
attached base packages:
[1] stats4 stats graphics grDevices utils datasets methods
[8] base
other attached packages:
[1] clusterProfiler_4.6.2 enrichplot_1.18.4
[3] org.Hs.eg.db_3.16.0 AnnotationDbi_1.60.2
[5] genekitr_1.2.8 ggrepel_0.9.6
[7] BiocParallel_1.32.6 DESeq2_1.38.3
[9] SummarizedExperiment_1.28.0 Biobase_2.58.0
[11] MatrixGenerics_1.10.0 matrixStats_1.2.0
[13] GenomicRanges_1.50.2 GenomeInfoDb_1.34.9
[15] IRanges_2.32.0 S4Vectors_0.36.2
[17] BiocGenerics_0.44.0 corrplot_0.95
[19] ggplot2_3.5.1 dplyr_1.1.4
[21] data.table_1.16.4 workflowr_1.7.1
loaded via a namespace (and not attached):
[1] shadowtext_0.1.4 fastmatch_1.1-6 plyr_1.8.9
[4] igraph_1.5.1 lazyeval_0.2.2 splines_4.2.2
[7] usethis_3.1.0 urltools_1.7.3 digest_0.6.37
[10] yulab.utils_0.2.0 htmltools_0.5.8.1 GOSemSim_2.24.0
[13] viridis_0.6.5 GO.db_3.16.0 magrittr_2.0.3
[16] memoise_2.0.1 remotes_2.5.0 openxlsx_4.2.5.2
[19] Biostrings_2.66.0 annotate_1.76.0 graphlayouts_1.0.1
[22] prettyunits_1.2.0 colorspace_2.1-1 blob_1.2.4
[25] xfun_0.50 callr_3.7.6 crayon_1.5.3
[28] RCurl_1.98-1.16 jsonlite_1.8.9 scatterpie_0.2.4
[31] ape_5.7-1 glue_1.8.0 polyclip_1.10-7
[34] gtable_0.3.6 zlibbioc_1.44.0 XVector_0.38.0
[37] DelayedArray_0.24.0 pkgbuild_1.4.6 scales_1.3.0
[40] DOSE_3.24.2 DBI_1.2.3 miniUI_0.1.1.1
[43] Rcpp_1.0.14 viridisLite_0.4.2 xtable_1.8-4
[46] progress_1.2.3 gridGraphics_0.5-1 tidytree_0.4.6
[49] bit_4.5.0.1 europepmc_0.4.3 profvis_0.4.0
[52] htmlwidgets_1.6.4 httr_1.4.7 fgsea_1.24.0
[55] RColorBrewer_1.1-3 ellipsis_0.3.2 urlchecker_1.0.1
[58] pkgconfig_2.0.3 XML_3.99-0.18 farver_2.1.2
[61] sass_0.4.9 locfit_1.5-9.8 labeling_0.4.3
[64] ggplotify_0.1.2 tidyselect_1.2.1 rlang_1.1.5
[67] reshape2_1.4.4 later_1.4.1 munsell_0.5.1
[70] tools_4.2.2 cachem_1.1.0 downloader_0.4
[73] cli_3.6.3 generics_0.1.3 RSQLite_2.3.9
[76] gson_0.1.0 devtools_2.4.5 evaluate_1.0.3
[79] stringr_1.5.1 fastmap_1.2.0 yaml_2.3.10
[82] ggtree_3.6.2 processx_3.8.5 knitr_1.49
[85] bit64_4.6.0-1 fs_1.6.5 tidygraph_1.3.0
[88] zip_2.3.2 purrr_1.0.2 KEGGREST_1.38.0
[91] ggraph_2.1.0 nlme_3.1-160 mime_0.12
[94] whisker_0.4.1 aplot_0.2.4 ggvenn_0.1.10
[97] xml2_1.3.6 compiler_4.2.2 rstudioapi_0.17.1
[100] png_0.1-8 treeio_1.22.0 tibble_3.2.1
[103] tweenr_2.0.3 geneplotter_1.76.0 bslib_0.9.0
[106] stringi_1.8.4 ps_1.8.1 lattice_0.22-6
[109] Matrix_1.6-4 vctrs_0.6.5 pillar_1.10.1
[112] lifecycle_1.0.4 triebeard_0.4.1 jquerylib_0.1.4
[115] cowplot_1.1.3 bitops_1.0-9 httpuv_1.6.15
[118] patchwork_1.3.0 qvalue_2.30.0 R6_2.5.1
[121] promises_1.3.2 gridExtra_2.3 sessioninfo_1.2.2
[124] codetools_0.2-20 pkgload_1.4.0 MASS_7.3-58.1
[127] rprojroot_2.0.4 withr_3.0.2 GenomeInfoDbData_1.2.9
[130] parallel_4.2.2 hms_1.1.3 grid_4.2.2
[133] ggfun_0.1.8 tidyr_1.3.1 HDO.db_0.99.1
[136] rmarkdown_2.29 git2r_0.33.0 getPass_0.2-4
[139] ggforce_0.4.1 shiny_1.10.0 geneset_0.2.7